bitscore colors: <40, 40-50 , 50-80, 80-200, >200
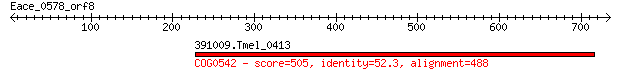
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eace_0578_orf8
Length=735
Score E
Sequences producing significant alignments: (Bits) Value
391009.Tmel_0413 505 2e-141
> 391009.Tmel_0413
Length=789
Score = 505 bits (1301), Expect = 2e-141, Method: Compositional matrix adjust.
Identities = 255/497 (51%), Positives = 346/497 (69%), Gaps = 9/497 (1%)
Query 228 ITQYGEDLTEKAAKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 287
+T+Y DLT+ A +GKL PVIGR++EI+R+ +ILSR P+LIG+PGVGKTA++EGLA
Sbjct 155 LTKYTVDLTKLAKEGKLSPVIGREKEIERVIEILSRKTKNNPVLIGDPGVGKTAIVEGLA 214
Query 288 QRIVEGAVPKGLRGRRIISLDLLNLSAGTSMRGEFEKRMKEIITFLQRRKKDVILFVDEI 347
QRIVEG VP L+ +RI+ LDL + AG+ RGEFE+R+K+++ + + K +VILF+DE+
Sbjct 215 QRIVEGKVPDKLKNKRILMLDLGKMLAGSKYRGEFEERLKKVMEEINKLKNNVILFIDEL 274
Query 348 HTLVGAGKTSGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRRFQNIVVEA 407
HT+VGAG G+ DA+ +LK LARGE+ +GATTLEEY+ HIEKD A RRFQ + VE
Sbjct 275 HTIVGAGAAEGAMDASNMLKPALARGELHAIGATTLEEYRKHIEKDKALARRFQPVFVEE 334
Query 408 PSKERALSILKKVKTNYEKFHSVDLSDEVLEGIVALSDQYVKQRSFPDKALDLLDEACSW 467
PS E + ILK +K YEK H+V ++DE +E LS +Y++ R PDKA+DL+DEA +
Sbjct 335 PSVEETIRILKGLKETYEKHHNVKITDEAIEAAAKLSSKYIQDRFMPDKAIDLIDEAAAK 394
Query 468 KKV-SHNKQIVVLKRKIEEAGKKKDE--------TAAEELQKWKDELAALEELTANGRRL 518
+ S + +I LK KI+ +K DE AAE Q+ EEL
Sbjct 395 VSLKSSDPKISELKEKIKSLSEKIDELTINSKYKEAAEMKQQMFKLQKQYEELEKKYGTD 454
Query 519 ALSLHDVAHILHLWTGIPLGKMTEDEISRVLRLADILSSRVIGQQQAVKAVADALAIQRA 578
+S VA ++ WTGIP KM E E ++L+ +++ R++ Q++AV+ VADA+ RA
Sbjct 455 TVSAETVAKVVESWTGIPATKMMESEKEKLLKFEELVHQRMVDQEEAVRVVADAIRKARA 514
Query 579 GLSPKNKPLGTFMFLGSSGVGKTELAKAVAEEMFDSEKNLIRLDMVEYQEAHSISRLIGP 638
G+ N+P+GTF+FLG SGVGKTELAK +AE +F SE LIR+DM EY E HS++RLIG
Sbjct 515 GIKDPNRPIGTFLFLGPSGVGKTELAKTIAELLFGSENALIRIDMSEYMEKHSVARLIGA 574
Query 639 PPGYMGNDEGGQLTEAVRQKPHSVVLFDEVENAHKNLWSVLLPMLDEGHLSDSKGNRVDF 698
PPGY+G D+GGQLTEAVR+KP+SV+L DEVE AH +++VLL M D+G L+D KGN VDF
Sbjct 575 PPGYVGYDQGGQLTEAVRRKPYSVILLDEVEKAHPEVFNVLLQMFDDGRLTDGKGNTVDF 634
Query 699 TNCLIILTSNIGRQHIL 715
N +II+TSN+ IL
Sbjct 635 RNTIIIMTSNLASDKIL 651
Lambda K H
0.317 0.133 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 346059122915
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40