bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_0149_orf2
Length=178
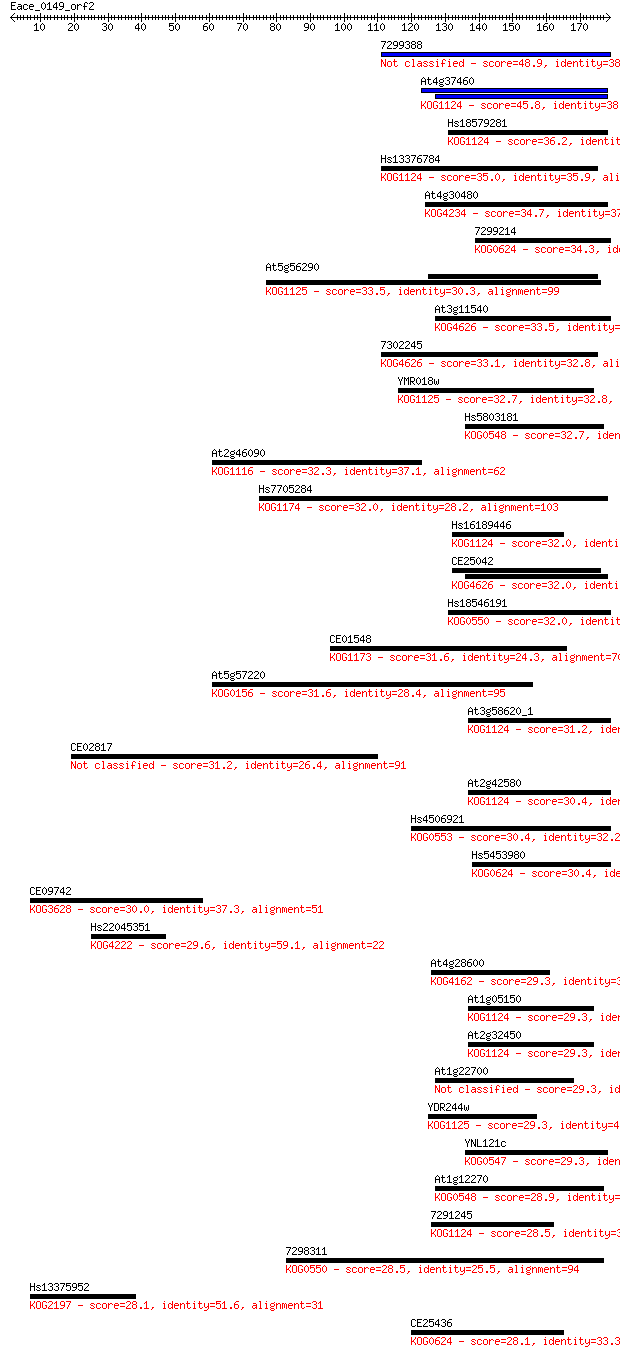
Score E
Sequences producing significant alignments: (Bits) Value
7299388 48.9 5e-06
At4g37460 45.8 4e-05
Hs18579281 36.2 0.038
Hs13376784 35.0 0.088
At4g30480 34.7 0.10
7299214 34.3 0.14
At5g56290 33.5 0.23
At3g11540 33.5 0.26
7302245 33.1 0.33
YMR018w 32.7 0.34
Hs5803181 32.7 0.38
At2g46090 32.3 0.56
Hs7705284 32.0 0.63
Hs16189446 32.0 0.66
CE25042 32.0 0.66
Hs18546191 32.0 0.68
CE01548 31.6 0.86
At5g57220 31.6 0.89
At3g58620_1 31.2 1.0
CE02817 31.2 1.2
At2g42580 30.4 1.7
Hs4506921 30.4 1.8
Hs5453980 30.4 1.9
CE09742 30.0 2.5
Hs22045351 29.6 3.1
At4g28600 29.3 3.8
At1g05150 29.3 4.7
At2g32450 29.3 4.7
At1g22700 29.3 4.8
YDR244w 29.3 4.8
YNL121c 29.3 4.9
At1g12270 28.9 5.6
7291245 28.5 7.4
7298311 28.5 8.1
Hs13375952 28.1 9.0
CE25436 28.1 9.4
> 7299388
Length=868
Score = 48.9 bits (115), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 26/68 (38%), Positives = 38/68 (55%), Gaps = 0/68 (0%)
Query 111 LAQQQGADWAKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDY 170
L Q+Q + WA R + G+ + Q +A Q + AL + P +ALVARGA YAN+ +
Sbjct 293 LRQKQASQWAFRSVADGIEHFKNGQQVEAFQCLNKALNIDPRNVEALVARGALYANRGSF 352
Query 171 EKAVADLE 178
K + D E
Sbjct 353 LKGLQDFE 360
> At4g37460
Length=1013
Score = 45.8 bits (107), Expect = 4e-05, Method: Compositional matrix adjust.
Identities = 21/55 (38%), Positives = 31/55 (56%), Gaps = 0/55 (0%)
Query 123 RLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADL 177
RL G+A +AI ++D L+ P Y +AL+ RG AYA + + E A+AD
Sbjct 300 RLSRGIAQVNEGNYTKAISIFDKVLKEEPTYPEALIGRGTAYAFQRELESAIADF 354
Score = 32.0 bits (71), Expect = 0.68, Method: Compositional matrix adjust.
Identities = 21/51 (41%), Positives = 29/51 (56%), Gaps = 0/51 (0%)
Query 127 GVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADL 177
G A A +R+LE AI + A+Q P ++A RG A A +Y +AV DL
Sbjct 338 GTAYAFQRELESAIADFTKAIQSNPAASEAWKRRGQARAALGEYVEAVEDL 388
> Hs18579281
Length=733
Score = 36.2 bits (82), Expect = 0.038, Method: Composition-based stats.
Identities = 20/60 (33%), Positives = 30/60 (50%), Gaps = 13/60 (21%)
Query 131 ARRRQLEQAIQLYDSALQLRPDYADALVARG-------------AAYANKLDYEKAVADL 177
A +LE+A QLY A+ +RPD+ A ++RG AY L+ ++ ADL
Sbjct 359 ANESRLEEADQLYRQAISMRPDFKQAYISRGELLLKMNKPLKAKEAYLKALELDRNNADL 418
> Hs13376784
Length=170
Score = 35.0 bits (79), Expect = 0.088, Method: Compositional matrix adjust.
Identities = 23/64 (35%), Positives = 32/64 (50%), Gaps = 0/64 (0%)
Query 111 LAQQQGADWAKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDY 170
LA +Q A+ G R QL++AI+ Y AL+L+PD+ D + AA D
Sbjct 70 LAIKQNPLLAEAYSNLGNVYKERGQLQEAIEHYRHALRLKPDFIDGYINLAAALVAAGDM 129
Query 171 EKAV 174
E AV
Sbjct 130 EGAV 133
> At4g30480
Length=277
Score = 34.7 bits (78), Expect = 0.10, Method: Compositional matrix adjust.
Identities = 20/54 (37%), Positives = 27/54 (50%), Gaps = 0/54 (0%)
Query 124 LKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADL 177
L GV + + E+ I+ AL+L P Y ALV R A+ +E AV DL
Sbjct 150 LNRGVCFLKLGKCEETIKECTKALELNPTYNKALVRRAEAHEKLEHFEDAVTDL 203
> 7299214
Length=498
Score = 34.3 bits (77), Expect = 0.14, Method: Compositional matrix adjust.
Identities = 13/40 (32%), Positives = 24/40 (60%), Gaps = 0/40 (0%)
Query 139 AIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
A+Q + L+L+PD+ A + RG + +YE+A+ D +
Sbjct 96 AVQDFSRVLELKPDFMAARIQRGVVHMKSGEYEQAIQDFD 135
> At5g56290
Length=728
Score = 33.5 bits (75), Expect = 0.23, Method: Compositional matrix adjust.
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 0/50 (0%)
Query 125 KAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAV 174
K G A Q AI Y AL L+P+Y A G +YAN+ Y++++
Sbjct 629 KLGATQANSVQSADAISAYQQALDLKPNYVRAWANMGISYANQGMYKESI 678
Score = 30.4 bits (67), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 27/101 (26%), Positives = 44/101 (43%), Gaps = 2/101 (1%)
Query 77 LSFLSTTSPSHAIRVAIAKAETDAAVRGGGVGRLL--AQQQGADWAKRRLKAGVACARRR 134
L +L +H AIA E ++ + RL A Q + A + GV R
Sbjct 545 LKYLYGWLRNHPKYGAIAPPELADSLYHADIARLFNEASQLNPEDADVHIVLGVLYNLSR 604
Query 135 QLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVA 175
+ ++AI + +ALQL+P+ GA AN + A++
Sbjct 605 EFDRAITSFQTALQLKPNDYSLWNKLGATQANSVQSADAIS 645
> At3g11540
Length=914
Score = 33.5 bits (75), Expect = 0.26, Method: Composition-based stats.
Identities = 18/52 (34%), Positives = 25/52 (48%), Gaps = 0/52 (0%)
Query 127 GVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
GV + Q + A+ Y+ A RP YA+A G Y N+ D E A+ E
Sbjct 193 GVVYSEMMQYDNALSCYEKAALERPMYAEAYCNMGVIYKNRGDLEMAITCYE 244
> 7302245
Length=1059
Score = 33.1 bits (74), Expect = 0.33, Method: Compositional matrix adjust.
Identities = 21/64 (32%), Positives = 31/64 (48%), Gaps = 0/64 (0%)
Query 111 LAQQQGADWAKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDY 170
LA +Q A+ G R QL++A+ Y A++L+PD+ D + AA D
Sbjct 109 LAIKQNPVLAEAYSNLGNVFKERGQLQEALDNYRRAVRLKPDFIDGYINLAAALVAARDM 168
Query 171 EKAV 174
E AV
Sbjct 169 ESAV 172
> YMR018w
Length=514
Score = 32.7 bits (73), Expect = 0.34, Method: Compositional matrix adjust.
Identities = 19/58 (32%), Positives = 24/58 (41%), Gaps = 4/58 (6%)
Query 116 GADWAKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKA 173
G W + G A + AI Y+ QLRP++ AY NK DY KA
Sbjct 394 GTIWNR----YGAILANTKSYHSAINAYNKCKQLRPNFTRVRYNLAIAYMNKGDYVKA 447
> Hs5803181
Length=543
Score = 32.7 bits (73), Expect = 0.38, Method: Composition-based stats.
Identities = 17/41 (41%), Positives = 23/41 (56%), Gaps = 0/41 (0%)
Query 136 LEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVAD 176
++ A+Q Y A++L P R AAYA K DY+KA D
Sbjct 20 IDDALQCYSEAIKLDPHNHVLYSNRSAAYAKKGDYQKAYED 60
> At2g46090
Length=364
Score = 32.3 bits (72), Expect = 0.56, Method: Compositional matrix adjust.
Identities = 23/68 (33%), Positives = 31/68 (45%), Gaps = 6/68 (8%)
Query 61 AEAWEALL----QRLSDGCFLSFLSTTSPSHAIRVA--IAKAETDAAVRGGGVGRLLAQQ 114
A+ W+ LL RL C +S L T+ PSHAI + + DA + GG G L
Sbjct 68 AKEWKKLLPHLRSRLGKDCNVSELLTSGPSHAIDITREAIRDGADAVIAVGGDGTLHEVV 127
Query 115 QGADWAKR 122
G W +
Sbjct 128 NGFFWEGK 135
> Hs7705284
Length=565
Score = 32.0 bits (71), Expect = 0.63, Method: Compositional matrix adjust.
Identities = 29/103 (28%), Positives = 42/103 (40%), Gaps = 12/103 (11%)
Query 75 CFLSFLSTTSPSHAIRVAIAKAETDAAVRGGGVGRLLAQQQGADWAKRRLKAGVACARRR 134
C+ + S++IR +A V V + L GA+ L A V
Sbjct 375 CYEGLIECYLASNSIR--------EAMVMANNVYKTL----GANAQTLTLLATVCLEDPV 422
Query 135 QLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADL 177
E+A L D AL RPDY A+V + + + YE +A L
Sbjct 423 TQEKAKTLLDKALTQRPDYIKAVVKKAELLSREQKYEDGIALL 465
> Hs16189446
Length=303
Score = 32.0 bits (71), Expect = 0.66, Method: Compositional matrix adjust.
Identities = 12/33 (36%), Positives = 20/33 (60%), Gaps = 0/33 (0%)
Query 132 RRRQLEQAIQLYDSALQLRPDYADALVARGAAY 164
R + E+A+ + AL++ P + DA V RG +Y
Sbjct 5 RINEFEEAVNFFTWALKINPCFLDAYVGRGNSY 37
> CE25042
Length=1151
Score = 32.0 bits (71), Expect = 0.66, Method: Compositional matrix adjust.
Identities = 15/44 (34%), Positives = 27/44 (61%), Gaps = 0/44 (0%)
Query 132 RRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVA 175
+ QL+ A++ Y A++L+P++ DA + AA + D E+AV
Sbjct 205 EKGQLQDALENYKLAVKLKPEFIDAYINLAAALVSGGDLEQAVT 248
Score = 29.3 bits (64), Expect = 4.5, Method: Compositional matrix adjust.
Identities = 17/47 (36%), Positives = 28/47 (59%), Gaps = 7/47 (14%)
Query 136 LEQAIQLYDSALQLRPDYADALVARGAAYANKL-----DYEKAVADL 177
+ +AIQ Y +AL+L+PD+ DA A+ +++ DY+K V L
Sbjct 549 MAEAIQSYSTALKLKPDFPDAYC--NLAHCHQIICDWNDYDKRVRKL 593
> Hs18546191
Length=237
Score = 32.0 bits (71), Expect = 0.68, Method: Compositional matrix adjust.
Identities = 16/48 (33%), Positives = 26/48 (54%), Gaps = 0/48 (0%)
Query 131 ARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
++ R+L+ AIQ + ++L Y A + R Y N YE+AV + E
Sbjct 97 SKLRKLDNAIQDCTNTVKLNDTYLKACLRRAQCYMNTKQYEEAVHNYE 144
> CE01548
Length=655
Score = 31.6 bits (70), Expect = 0.86, Method: Compositional matrix adjust.
Identities = 17/70 (24%), Positives = 31/70 (44%), Gaps = 3/70 (4%)
Query 96 AETDAAVRGGGVGRLLAQQQGADWAKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYAD 155
ETD + + +L ++ W G R+ +L +AI Y A+++ P + D
Sbjct 474 TETDEII---PIEEVLKKKIDDFWHPMLNNIGHIARRQGRLNEAIMFYQKAIRMEPKFVD 530
Query 156 ALVARGAAYA 165
A+ + YA
Sbjct 531 AIASTALCYA 540
> At5g57220
Length=491
Score = 31.6 bits (70), Expect = 0.89, Method: Compositional matrix adjust.
Identities = 27/101 (26%), Positives = 46/101 (45%), Gaps = 6/101 (5%)
Query 61 AEAWEALLQRLSDGCFLSFLSTTSPSHAIRVAI--AKAETDAAVRGGGVGRLLAQQQGA- 117
EA +A LQRL D C ++ S T SH + + + K +D ++G + +LA A
Sbjct 242 GEAMDAFLQRLLDECRINGESNTMVSHLLSLQLDQPKYYSDVIIKGLMLSMMLAGTDTAA 301
Query 118 ---DWAKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYAD 155
+WA L ++ + E ++ + L PD A+
Sbjct 302 VTLEWAMANLLKKPEVLKKAKAEIDEKIGEERLVDEPDIAN 342
> At3g58620_1
Length=578
Score = 31.2 bits (69), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 137 EQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
E+++ + AL+++P Y AL+ R A+Y +E AV D E
Sbjct 500 EKSVDDCNQALRIQPSYTKALLRRAASYGKLGRWEDAVRDYE 541
> CE02817
Length=1177
Score = 31.2 bits (69), Expect = 1.2, Method: Composition-based stats.
Identities = 24/91 (26%), Positives = 42/91 (46%), Gaps = 4/91 (4%)
Query 19 QMLGMAPSPEPHSSSGSSSSNSSSSARKKASNDNNRALICSEAEAWEALLQRLSDGCFLS 78
+ML PHS++ SSS + S ++ + + +L+ S A ++ + S G +
Sbjct 1051 EMLVSYSQSAPHSNANSSSPQMNGSMQRVKAYNYPDSLLYSPAGTTSSIAETNSSGNGAT 1110
Query 79 FLSTTSPSHAIRVAIAKAETDAAVRGGGVGR 109
SP++ A A E+D +R GG R
Sbjct 1111 RAQCDSPNN----APASDESDQGIRCGGSSR 1137
> At2g42580
Length=618
Score = 30.4 bits (67), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 137 EQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
E++++ + AL+ +P Y AL+ R A+Y +E AV D E
Sbjct 509 EKSVEDCNHALKSQPSYIKALLRRAASYGKLGRWEDAVKDYE 550
> Hs4506921
Length=313
Score = 30.4 bits (67), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 19/59 (32%), Positives = 26/59 (44%), Gaps = 0/59 (0%)
Query 120 AKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
A+R G + E A+ Y A++L P A R AAY+ +Y AV D E
Sbjct 91 AERLKTEGNEQMKVENFEAAVHFYGKAIELNPANAVYFCNRAAAYSKLGNYAGAVQDCE 149
> Hs5453980
Length=504
Score = 30.4 bits (67), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 15/41 (36%), Positives = 24/41 (58%), Gaps = 0/41 (0%)
Query 138 QAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADLE 178
+AI++ LQ+ PD +AL R AY + Y++A+ D E
Sbjct 324 EAIRVCSEVLQMEPDNVNALKDRAEAYLIEEMYDEAIQDYE 364
> CE09742
Length=203
Score = 30.0 bits (66), Expect = 2.5, Method: Compositional matrix adjust.
Identities = 19/51 (37%), Positives = 30/51 (58%), Gaps = 1/51 (1%)
Query 7 AAHRRLHAFSALQMLGMAPSPEPHSSSGSSSSNSSSSARKKASNDNNRALI 57
A H+ L +S + LG P +PH +G+S S SSS+ + K SN + R ++
Sbjct 136 AVHQALAEYSDGRKLGPQPV-KPHRRNGTSVSRSSSAQKSKRSNHSGRRML 185
> Hs22045351
Length=1190
Score = 29.6 bits (65), Expect = 3.1, Method: Composition-based stats.
Identities = 13/22 (59%), Positives = 16/22 (72%), Gaps = 0/22 (0%)
Query 25 PSPEPHSSSGSSSSNSSSSARK 46
PSP HSSSG++SS S+ RK
Sbjct 1136 PSPGGHSSSGTASSKGSTGPRK 1157
> At4g28600
Length=710
Score = 29.3 bits (64), Expect = 3.8, Method: Compositional matrix adjust.
Identities = 13/35 (37%), Positives = 23/35 (65%), Gaps = 0/35 (0%)
Query 126 AGVACARRRQLEQAIQLYDSALQLRPDYADALVAR 160
AGV RR QLE+A++ + +AL + P + +L ++
Sbjct 603 AGVLYNRRGQLEEAMEAFTTALDIDPMHVPSLTSK 637
> At1g05150
Length=808
Score = 29.3 bits (64), Expect = 4.7, Method: Compositional matrix adjust.
Identities = 12/37 (32%), Positives = 23/37 (62%), Gaps = 0/37 (0%)
Query 137 EQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKA 173
E+AI+++ A+ L+P + DAL G Y + +++A
Sbjct 395 ERAIEVFQRAIDLKPGHVDALYNLGGLYMDLGRFQRA 431
> At2g32450
Length=802
Score = 29.3 bits (64), Expect = 4.7, Method: Compositional matrix adjust.
Identities = 12/37 (32%), Positives = 23/37 (62%), Gaps = 0/37 (0%)
Query 137 EQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKA 173
E+AI+++ A+ L+P + DAL G Y + +++A
Sbjct 390 ERAIEVFQRAIDLKPGHVDALYNLGGLYMDLGRFQRA 426
> At1g22700
Length=299
Score = 29.3 bits (64), Expect = 4.8, Method: Compositional matrix adjust.
Identities = 14/41 (34%), Positives = 23/41 (56%), Gaps = 0/41 (0%)
Query 127 GVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANK 167
GV+ R +L++ I ++ A++L+P Y A G AY K
Sbjct 211 GVSYVREDKLDKGIAQFEMAVKLQPGYVTAWNNLGDAYEKK 251
> YDR244w
Length=612
Score = 29.3 bits (64), Expect = 4.8, Method: Composition-based stats.
Identities = 13/32 (40%), Positives = 19/32 (59%), Gaps = 0/32 (0%)
Query 125 KAGVACARRRQLEQAIQLYDSALQLRPDYADA 156
+ G + A + E+AIQ Y ALQL+P + A
Sbjct 496 RLGASLANSNRSEEAIQAYHRALQLKPSFVRA 527
> YNL121c
Length=617
Score = 29.3 bits (64), Expect = 4.9, Method: Composition-based stats.
Identities = 13/42 (30%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 136 LEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVADL 177
L++ +++ AL+L+PDY+ L+ R +A + A+ DL
Sbjct 148 LKKVVEMSTKALELKPDYSKVLLRRASANEGLGKFADAMFDL 189
> At1g12270
Length=572
Score = 28.9 bits (63), Expect = 5.6, Method: Compositional matrix adjust.
Identities = 14/50 (28%), Positives = 24/50 (48%), Gaps = 0/50 (0%)
Query 127 GVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAYANKLDYEKAVAD 176
G A +++ E AIQ Y +A+++ + L R A Y Y + + D
Sbjct 251 GNAAYKKKDFETAIQHYSTAIEIDDEDISYLTNRAAVYLEMGKYNECIED 300
> 7291245
Length=870
Score = 28.5 bits (62), Expect = 7.4, Method: Compositional matrix adjust.
Identities = 13/36 (36%), Positives = 20/36 (55%), Gaps = 0/36 (0%)
Query 126 AGVACARRRQLEQAIQLYDSALQLRPDYADALVARG 161
A + + +LE+A LY A+ +R DY A + RG
Sbjct 547 ANLIAKNQTRLEEADHLYRQAISMRSDYVQAYINRG 582
> 7298311
Length=508
Score = 28.5 bits (62), Expect = 8.1, Method: Compositional matrix adjust.
Identities = 24/94 (25%), Positives = 40/94 (42%), Gaps = 4/94 (4%)
Query 83 TSPSHAIRVAIAKAETDAAVRGGGVGRLLAQQQGADWAKRRLKAGVACARRRQLEQAIQL 142
TS + + V I + V+ + + A A+ + K G + + + A++L
Sbjct 16 TSSNSEMDVEITTEQPTIDVKAEQI----VPKDAATIAEEKKKLGNDQYKAQNYQNALKL 71
Query 143 YDSALQLRPDYADALVARGAAYANKLDYEKAVAD 176
Y A+ L PD A R A Y L+Y A+ D
Sbjct 72 YTDAISLCPDSAAYYGNRAACYMMLLNYNSALTD 105
> Hs13375952
Length=648
Score = 28.1 bits (61), Expect = 9.0, Method: Compositional matrix adjust.
Identities = 16/36 (44%), Positives = 22/36 (61%), Gaps = 5/36 (13%)
Query 7 AAHRRLHAFSAL-----QMLGMAPSPEPHSSSGSSS 37
A HR+L A L ++LG+AP EPHS S ++S
Sbjct 573 ALHRQLTACPELPHNVQEILGLAPPAEPHSPSPTAS 608
> CE25436
Length=494
Score = 28.1 bits (61), Expect = 9.4, Method: Compositional matrix adjust.
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 0/45 (0%)
Query 120 AKRRLKAGVACARRRQLEQAIQLYDSALQLRPDYADALVARGAAY 164
A+R +AG A RQ A+ Y A++L P A+ R Y
Sbjct 24 AQREYEAGNALFVNRQYSDALTHYHKAIELNPTMYQAIFRRATTY 68
Lambda K H
0.315 0.125 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2779358582
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40