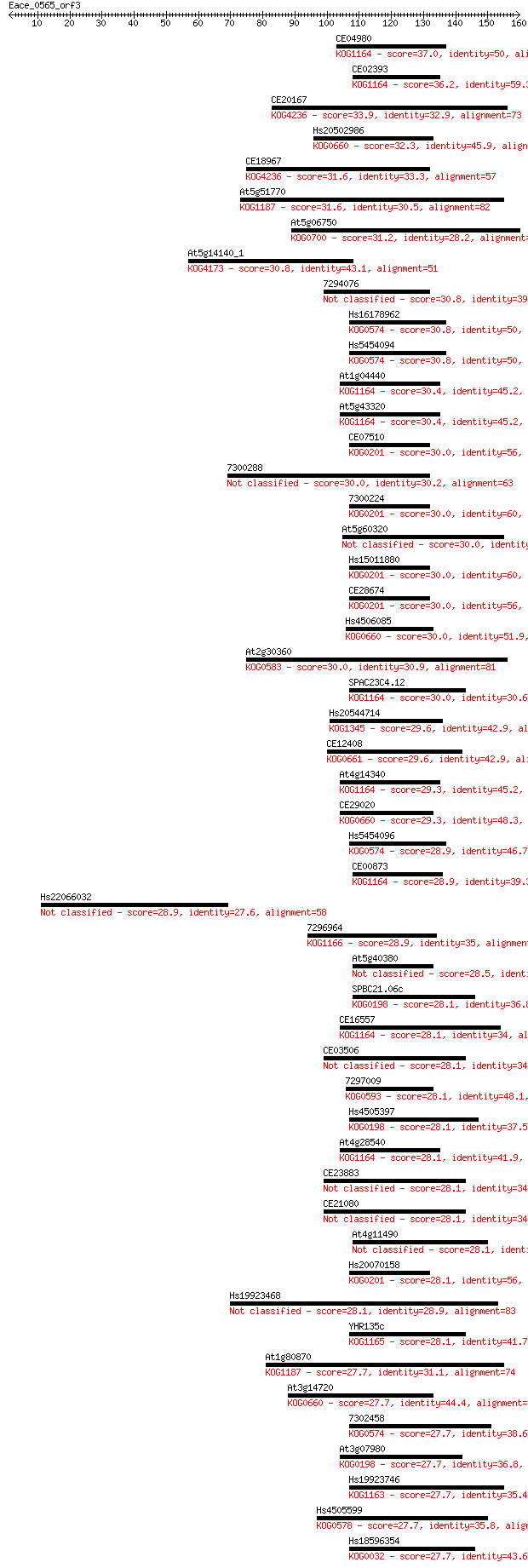
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_0565_orf3
Length=159
Score E
Sequences producing significant alignments: (Bits) Value
CE04980 37.0 0.014
CE02393 36.2 0.027
CE20167 33.9 0.13
Hs20502986 32.3 0.39
CE18967 31.6 0.63
At5g51770 31.6 0.65
At5g06750 31.2 0.93
At5g14140_1 30.8 1.0
7294076 30.8 1.0
Hs16178962 30.8 1.3
Hs5454094 30.8 1.3
At1g04440 30.4 1.3
At5g43320 30.4 1.4
CE07510 30.0 1.7
7300288 30.0 1.8
7300224 30.0 1.9
At5g60320 30.0 1.9
Hs15011880 30.0 2.0
CE28674 30.0 2.0
Hs4506085 30.0 2.0
At2g30360 30.0 2.2
SPAC23C4.12 30.0 2.2
Hs20544714 29.6 2.6
CE12408 29.6 2.7
At4g14340 29.3 3.6
CE29020 29.3 3.7
Hs5454096 28.9 4.2
CE00873 28.9 4.6
Hs22066032 28.9 4.6
7296964 28.9 4.7
At5g40380 28.5 6.2
SPBC21.06c 28.1 6.5
CE16557 28.1 6.5
CE03506 28.1 6.7
7297009 28.1 6.9
Hs4505397 28.1 7.0
At4g28540 28.1 7.5
CE23883 28.1 7.6
CE21080 28.1 7.6
At4g11490 28.1 7.8
Hs20070158 28.1 7.8
Hs19923468 28.1 7.8
YHR135c 28.1 7.9
At1g80870 27.7 8.5
At3g14720 27.7 8.7
7302458 27.7 8.7
At3g07980 27.7 8.8
Hs19923746 27.7 9.0
Hs4505599 27.7 9.2
Hs18596354 27.7 9.3
> CE04980
Length=361
Score = 37.0 bits (84), Expect = 0.014, Method: Compositional matrix adjust.
Identities = 17/34 (50%), Positives = 24/34 (70%), Gaps = 0/34 (0%)
Query 103 FLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKE 136
++ +GKGG GE++LA+D G VAIK EPK+
Sbjct 63 LIEGLIGKGGYGEIYLALDVVRGEEVAIKVEPKK 96
> CE02393
Length=366
Score = 36.2 bits (82), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 16/27 (59%), Positives = 20/27 (74%), Gaps = 0/27 (0%)
Query 108 LGKGGNGEVFLAIDRRHGGPVAIKREP 134
+GKGG GE++LAID + VAIK EP
Sbjct 74 IGKGGYGEIYLAIDMKLAEEVAIKAEP 100
> CE20167
Length=722
Score = 33.9 bits (76), Expect = 0.13, Method: Composition-based stats.
Identities = 24/73 (32%), Positives = 33/73 (45%), Gaps = 0/73 (0%)
Query 83 PAESTLFTEYQLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEF 142
P+ + E L ++V D LG G G V+ AI R G VA+K KE +
Sbjct 407 PSRNEDNAEQALEFANLYQVLSDKTLGSGQFGTVYSAIQRHSGKEVAVKVISKERFSKKG 466
Query 143 SLICEIRLEGALL 155
S +R E A+L
Sbjct 467 SGAESMRAEVAIL 479
> Hs20502986
Length=277
Score = 32.3 bits (72), Expect = 0.39, Method: Compositional matrix adjust.
Identities = 17/37 (45%), Positives = 24/37 (64%), Gaps = 2/37 (5%)
Query 96 VFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKR 132
+ RR+ L +LG+G G V+ A+DRR G VAIK+
Sbjct 9 IVRRY--LLRRQLGQGAYGIVWKAVDRRTGEVVAIKK 43
> CE18967
Length=1101
Score = 31.6 bits (70), Expect = 0.63, Method: Composition-based stats.
Identities = 19/57 (33%), Positives = 32/57 (56%), Gaps = 2/57 (3%)
Query 75 VETAGNSVPAESTLFTEYQLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIK 131
V T G + + + TE++ S + +++F + LG G G V+ I RR+G VA+K
Sbjct 745 VPTEGETGHLGAKIQTEHEFS--QLYQIFAEEVLGSGQFGTVYGGIHRRNGQHVAVK 799
> At5g51770
Length=654
Score = 31.6 bits (70), Expect = 0.65, Method: Composition-based stats.
Identities = 25/86 (29%), Positives = 35/86 (40%), Gaps = 4/86 (4%)
Query 73 TPVETAGNSVPAESTLFTEYQLSVFRRFEVFL--DTRLGKGGNGEVFLAI--DRRHGGPV 128
+P A +S P + E+ S R+ + RLG+GG G VF GG V
Sbjct 59 SPAAVASSSTPPQKQPLHEFSYSSLRKATASFSPENRLGQGGFGSVFRGTLSPSSGGGNV 118
Query 129 AIKREPKEMLRHEFSLICEIRLEGAL 154
A+K L+ E E+ G L
Sbjct 119 AVKVMDSGSLQGEREFQNELFFAGKL 144
> At5g06750
Length=386
Score = 31.2 bits (69), Expect = 0.93, Method: Compositional matrix adjust.
Identities = 20/71 (28%), Positives = 36/71 (50%), Gaps = 4/71 (5%)
Query 89 FTEYQLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLICEI 148
F ++ ++V + EV D + GNG VF+ + HGGP A + + H FS + +
Sbjct 47 FGDFSIAVVQANEVIEDHSQVETGNGAVFVGVYDGHGGPEA----SRYISDHLFSHLMRV 102
Query 149 RLEGALLTYKA 159
E + ++ +A
Sbjct 103 SRERSCISEEA 113
> At5g14140_1
Length=320
Score = 30.8 bits (68), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 22/56 (39%), Positives = 32/56 (57%), Gaps = 6/56 (10%)
Query 57 LGVDRPNPIRRCVPPDTPVETAGNS----VPAESTL-FTEYQLSVFRRFEVFLDTR 107
LG NPIRR PPD+P +GN + + TL FTE +++ +F V ++TR
Sbjct 88 LGFPYWNPIRRRFPPDSPFFASGNVERELLAKQVTLDFTEDEINHLHKF-VEIETR 142
> 7294076
Length=1504
Score = 30.8 bits (68), Expect = 1.0, Method: Composition-based stats.
Identities = 13/33 (39%), Positives = 22/33 (66%), Gaps = 0/33 (0%)
Query 99 RFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIK 131
R ++ + +LG G GEV+ A+ +R+G VA+K
Sbjct 368 RTDIMMKHKLGGGQYGEVYEAVWKRYGNTVAVK 400
> Hs16178962
Length=491
Score = 30.8 bits (68), Expect = 1.3, Method: Composition-based stats.
Identities = 15/30 (50%), Positives = 19/30 (63%), Gaps = 0/30 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKE 136
+LG+G G VF AI + G VAIK+ P E
Sbjct 32 KLGEGSYGSVFKAIHKESGQVVAIKQVPVE 61
> Hs5454094
Length=491
Score = 30.8 bits (68), Expect = 1.3, Method: Composition-based stats.
Identities = 15/30 (50%), Positives = 19/30 (63%), Gaps = 0/30 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKE 136
+LG+G G VF AI + G VAIK+ P E
Sbjct 32 KLGEGSYGSVFKAIHKESGQVVAIKQVPVE 61
> At1g04440
Length=468
Score = 30.4 bits (67), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 14/31 (45%), Positives = 20/31 (64%), Gaps = 0/31 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKREP 134
L +LG G GE+FL ++ + G VA+K EP
Sbjct 11 LGRKLGSGSFGEIFLGVNVQTGEEVAVKLEP 41
> At5g43320
Length=480
Score = 30.4 bits (67), Expect = 1.4, Method: Composition-based stats.
Identities = 14/31 (45%), Positives = 20/31 (64%), Gaps = 0/31 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKREP 134
L +LG G GE+FL ++ + G VA+K EP
Sbjct 11 LGRKLGSGSFGELFLGVNVQTGEEVAVKLEP 41
> CE07510
Length=653
Score = 30.0 bits (66), Expect = 1.7, Method: Composition-based stats.
Identities = 14/25 (56%), Positives = 17/25 (68%), Gaps = 0/25 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIK 131
R+G+G GEV+ ID R G VAIK
Sbjct 39 RIGRGSFGEVYKGIDNRTGRVVAIK 63
> 7300288
Length=630
Score = 30.0 bits (66), Expect = 1.8, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 26/63 (41%), Gaps = 0/63 (0%)
Query 69 VPPDTPVETAGNSVPAESTLFTEYQLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPV 128
V P V +VP E + + E +D +LG GG+ VFLA G
Sbjct 304 VEPKELVSIPAVAVPPEQPSHKTSNILKIKNHEYTIDKKLGCGGSSSVFLARRSDSGNEF 363
Query 129 AIK 131
A+K
Sbjct 364 ALK 366
> 7300224
Length=642
Score = 30.0 bits (66), Expect = 1.9, Method: Composition-based stats.
Identities = 15/25 (60%), Positives = 16/25 (64%), Gaps = 0/25 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIK 131
R+GKG GEVF ID R VAIK
Sbjct 18 RIGKGSFGEVFKGIDNRTQQVVAIK 42
> At5g60320
Length=675
Score = 30.0 bits (66), Expect = 1.9, Method: Composition-based stats.
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Query 105 DTRLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLICEIRLEGAL 154
D RLGKGG GEV+ + H G +A+KR + + + E+ G+L
Sbjct 351 DGRLGKGGFGEVYRG-NLPHVGDIAVKRVCHDAKQGMKQFVAEVVTMGSL 399
> Hs15011880
Length=416
Score = 30.0 bits (66), Expect = 2.0, Method: Compositional matrix adjust.
Identities = 15/25 (60%), Positives = 16/25 (64%), Gaps = 0/25 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIK 131
R+GKG GEVF ID R VAIK
Sbjct 29 RIGKGSFGEVFKGIDNRTQQVVAIK 53
> CE28674
Length=651
Score = 30.0 bits (66), Expect = 2.0, Method: Composition-based stats.
Identities = 14/25 (56%), Positives = 17/25 (68%), Gaps = 0/25 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIK 131
R+G+G GEV+ ID R G VAIK
Sbjct 37 RIGRGSFGEVYKGIDNRTGRVVAIK 61
> Hs4506085
Length=365
Score = 30.0 bits (66), Expect = 2.0, Method: Compositional matrix adjust.
Identities = 14/27 (51%), Positives = 17/27 (62%), Gaps = 0/27 (0%)
Query 106 TRLGKGGNGEVFLAIDRRHGGPVAIKR 132
T +G G G V AID+R G VAIK+
Sbjct 29 THVGSGAYGSVCSAIDKRSGEKVAIKK 55
> At2g30360
Length=435
Score = 30.0 bits (66), Expect = 2.2, Method: Compositional matrix adjust.
Identities = 25/81 (30%), Positives = 36/81 (44%), Gaps = 10/81 (12%)
Query 75 VETAGNSVPAESTLFTEYQLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREP 134
+E A S LF +Y+L LG G +VF A DRR G VA+K
Sbjct 4 IEIAAGSGDNNDALFGKYELGKL----------LGCGAFAKVFHARDRRTGQSVAVKILN 53
Query 135 KEMLRHEFSLICEIRLEGALL 155
K+ L +L I+ E +++
Sbjct 54 KKKLLTNPALANNIKREISIM 74
> SPAC23C4.12
Length=400
Score = 30.0 bits (66), Expect = 2.2, Method: Compositional matrix adjust.
Identities = 11/36 (30%), Positives = 22/36 (61%), Gaps = 0/36 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEF 142
++G G G+++L ++ +G VA+K EP + H+
Sbjct 17 KIGSGSFGQIYLGLNTVNGEQVAVKLEPLKARHHQL 52
> Hs20544714
Length=857
Score = 29.6 bits (65), Expect = 2.6, Method: Composition-based stats.
Identities = 15/36 (41%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query 101 EVFLDTR-LGKGGNGEVFLAIDRRHGGPVAIKREPK 135
E++ + R LG+G G V L R+ G P+A+K+ PK
Sbjct 467 ELYEEVRPLGQGCYGRVLLVTHRQKGTPLALKQLPK 502
> CE12408
Length=527
Score = 29.6 bits (65), Expect = 2.7, Method: Composition-based stats.
Identities = 18/42 (42%), Positives = 20/42 (47%), Gaps = 0/42 (0%)
Query 100 FEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHE 141
F + RLG G GEV LA G VAIKR K+ E
Sbjct 43 FRYLMTKRLGDGTFGEVMLAKKIDTGDRVAIKRMKKKFYSWE 84
> At4g14340
Length=457
Score = 29.3 bits (64), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 14/31 (45%), Positives = 20/31 (64%), Gaps = 0/31 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKREP 134
L +LG G GE++L I+ + G VA+K EP
Sbjct 17 LGRKLGSGSFGELYLGINIQTGEEVAVKLEP 47
> CE29020
Length=470
Score = 29.3 bits (64), Expect = 3.7, Method: Compositional matrix adjust.
Identities = 14/29 (48%), Positives = 18/29 (62%), Gaps = 0/29 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKR 132
L RLGKG G V+ A D+R VA+K+
Sbjct 15 LQKRLGKGAYGIVWKAYDKRSRETVALKK 43
> Hs5454096
Length=487
Score = 28.9 bits (63), Expect = 4.2, Method: Composition-based stats.
Identities = 14/30 (46%), Positives = 19/30 (63%), Gaps = 0/30 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKE 136
+LG+G G V+ AI + G VAIK+ P E
Sbjct 35 KLGEGSYGSVYKAIHKETGQIVAIKQVPVE 64
> CE00873
Length=363
Score = 28.9 bits (63), Expect = 4.6, Method: Compositional matrix adjust.
Identities = 11/28 (39%), Positives = 18/28 (64%), Gaps = 0/28 (0%)
Query 108 LGKGGNGEVFLAIDRRHGGPVAIKREPK 135
+G GG G++F+ +D + A+K EPK
Sbjct 78 IGNGGYGQIFMVMDVKKNDERAMKIEPK 105
> Hs22066032
Length=167
Score = 28.9 bits (63), Expect = 4.6, Method: Compositional matrix adjust.
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 2/59 (3%)
Query 11 FEVFTHLDAAEDCKFRMIPKSGINVNGKPFGADGNAACSFSFRRILLGVD-RPNPIRRC 68
E +D+A++CKF++ +G + P + S +R L G P P+R+C
Sbjct 86 LESRDDIDSAKECKFKLFTAAGPSQRKAPMSPE-PVVLSLGGQRPLAGPPLDPTPVRKC 143
> 7296964
Length=1099
Score = 28.9 bits (63), Expect = 4.7, Method: Composition-based stats.
Identities = 14/40 (35%), Positives = 21/40 (52%), Gaps = 0/40 (0%)
Query 94 LSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKRE 133
L+V +D +G+G G V+ A D R G VA+K +
Sbjct 806 LNVLEGVTFSIDKEVGRGSYGSVYKATDSRTGNVVALKYQ 845
> At5g40380
Length=629
Score = 28.5 bits (62), Expect = 6.2, Method: Compositional matrix adjust.
Identities = 15/25 (60%), Positives = 18/25 (72%), Gaps = 1/25 (4%)
Query 108 LGKGGNGEVFLAIDRRHGGPVAIKR 132
LG+GGNG VFL I +G VA+KR
Sbjct 299 LGQGGNGTVFLGI-LPNGKNVAVKR 322
> SPBC21.06c
Length=1062
Score = 28.1 bits (61), Expect = 6.5, Method: Composition-based stats.
Identities = 14/39 (35%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Query 108 LGKGGNGEVFLAIDRRHGGPVAIKR-EPKEMLRHEFSLI 145
LGKG G V+ ++ ++G VA+K+ + +ML+ + S+I
Sbjct 15 LGKGAFGAVYRGLNIKNGETVAVKKVKLSKMLKSDLSVI 53
> CE16557
Length=318
Score = 28.1 bits (61), Expect = 6.5, Method: Compositional matrix adjust.
Identities = 17/50 (34%), Positives = 26/50 (52%), Gaps = 0/50 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLICEIRLEGA 153
++ +G GG G++F A+D VA+K EPK + L I +E A
Sbjct 40 VEKMVGGGGFGQIFRAVDMETKLVVAVKVEPKSVESGRIVLELHILVELA 89
> CE03506
Length=1181
Score = 28.1 bits (61), Expect = 6.7, Method: Composition-based stats.
Identities = 15/45 (33%), Positives = 25/45 (55%), Gaps = 1/45 (2%)
Query 99 RFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKE-MLRHEF 142
R E+ + +LG G G+V+ +RH +A+K ++ M HEF
Sbjct 293 RSEIIMHNKLGGGQYGDVYEGYWKRHDCTIAVKALKEDAMPLHEF 337
> 7297009
Length=392
Score = 28.1 bits (61), Expect = 6.9, Method: Compositional matrix adjust.
Identities = 13/27 (48%), Positives = 17/27 (62%), Gaps = 0/27 (0%)
Query 106 TRLGKGGNGEVFLAIDRRHGGPVAIKR 132
+RLG+G G V+ DR G VA+KR
Sbjct 8 SRLGEGSYGVVYKCRDRETGALVAVKR 34
> Hs4505397
Length=947
Score = 28.1 bits (61), Expect = 7.0, Method: Composition-based stats.
Identities = 15/40 (37%), Positives = 22/40 (55%), Gaps = 0/40 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLIC 146
RLG+G GEV D++ G A+K+ E+ R E + C
Sbjct 405 RLGRGSFGEVHRMEDKQTGFQCAVKKVRLEVFRAEELMAC 444
> At4g28540
Length=321
Score = 28.1 bits (61), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 13/31 (41%), Positives = 19/31 (61%), Gaps = 0/31 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKREP 134
L ++G G GE+FLA+ + G A+K EP
Sbjct 11 LGRKIGGGSFGELFLAVSLQTGEEAAVKLEP 41
> CE23883
Length=1186
Score = 28.1 bits (61), Expect = 7.6, Method: Composition-based stats.
Identities = 15/45 (33%), Positives = 25/45 (55%), Gaps = 1/45 (2%)
Query 99 RFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKE-MLRHEF 142
R E+ + +LG G G+V+ +RH +A+K ++ M HEF
Sbjct 298 RSEIIMHNKLGGGQYGDVYEGYWKRHDCTIAVKALKEDAMPLHEF 342
> CE21080
Length=1196
Score = 28.1 bits (61), Expect = 7.6, Method: Composition-based stats.
Identities = 15/45 (33%), Positives = 25/45 (55%), Gaps = 1/45 (2%)
Query 99 RFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKE-MLRHEF 142
R E+ + +LG G G+V+ +RH +A+K ++ M HEF
Sbjct 308 RSEIIMHNKLGGGQYGDVYEGYWKRHDCTIAVKALKEDAMPLHEF 352
> At4g11490
Length=636
Score = 28.1 bits (61), Expect = 7.8, Method: Composition-based stats.
Identities = 17/48 (35%), Positives = 28/48 (58%), Gaps = 7/48 (14%)
Query 108 LGKGGNGEVFLAIDRRHGGPVAIKREPKEM------LRHEFSLICEIR 149
LG+GG GEVF + + G +A+KR KE ++E SL+ +++
Sbjct 327 LGQGGFGEVFKGV-LQDGSEIAVKRLSKESAQGVQEFQNETSLVAKLQ 373
> Hs20070158
Length=443
Score = 28.1 bits (61), Expect = 7.8, Method: Compositional matrix adjust.
Identities = 14/25 (56%), Positives = 16/25 (64%), Gaps = 0/25 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIK 131
++GKG GEVF ID R VAIK
Sbjct 41 KIGKGSFGEVFKGIDNRTQKVVAIK 65
> Hs19923468
Length=878
Score = 28.1 bits (61), Expect = 7.8, Method: Composition-based stats.
Identities = 24/91 (26%), Positives = 41/91 (45%), Gaps = 9/91 (9%)
Query 70 PPDTPVETAGNSVPAESTLFTEYQLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVA 129
P P A S+ ++ E + + +++F D LG G G V+ R+ G VA
Sbjct 520 PGHAPHRQASLSISVSNSQIQE-NVDIATVYQIFPDEVLGSGQFGVVYGGKHRKTGRDVA 578
Query 130 IK-----REP---KEMLRHEFSLICEIRLEG 152
+K R P + LR+E +++ +R G
Sbjct 579 VKVIDKLRFPTKQESQLRNEVAILQSLRHPG 609
> YHR135c
Length=538
Score = 28.1 bits (61), Expect = 7.9, Method: Composition-based stats.
Identities = 15/40 (37%), Positives = 24/40 (60%), Gaps = 4/40 (10%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKE----MLRHEF 142
++G+G G +F + +G PVAIK EP++ LR E+
Sbjct 74 KIGEGSFGVLFEGTNMINGVPVAIKFEPRKTEAPQLRDEY 113
> At1g80870
Length=692
Score = 27.7 bits (60), Expect = 8.5, Method: Composition-based stats.
Identities = 23/81 (28%), Positives = 38/81 (46%), Gaps = 8/81 (9%)
Query 81 SVPAESTLFTEYQLSVFRRFEVFLDTR-------LGKGGNGEVFLAIDRRHGGPVAIKRE 133
++P + + +L +F E+ L T +GKGG+G VF I R G A+KR
Sbjct 53 TIPFDVAAASPLKLQLFTYKELKLATNDFDESNVIGKGGSGTVFRGIT-RDGKLFAVKRL 111
Query 134 PKEMLRHEFSLICEIRLEGAL 154
++ E E+++ G L
Sbjct 112 DNLSIQTETEFQNELQILGGL 132
> At3g14720
Length=586
Score = 27.7 bits (60), Expect = 8.7, Method: Composition-based stats.
Identities = 20/46 (43%), Positives = 25/46 (54%), Gaps = 6/46 (13%)
Query 88 LFTEY-QLSVFRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIKR 132
FTEY + +R EV +GKG G V AID + G VAIK+
Sbjct 3 FFTEYGDANRYRILEV-----IGKGSYGVVCAAIDTQTGEKVAIKK 43
> 7302458
Length=596
Score = 27.7 bits (60), Expect = 8.7, Method: Composition-based stats.
Identities = 17/44 (38%), Positives = 24/44 (54%), Gaps = 2/44 (4%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLICEIRL 150
+LG+G G V+ A+ + VAIK P E HE +I EI +
Sbjct 47 KLGEGSYGSVYKAVHKESSSIVAIKLVPVESDLHE--IIKEISI 88
> At3g07980
Length=1367
Score = 27.7 bits (60), Expect = 8.8, Method: Composition-based stats.
Identities = 14/38 (36%), Positives = 21/38 (55%), Gaps = 0/38 (0%)
Query 104 LDTRLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHE 141
L +GKG G V++ +D +G VAIK+ E + E
Sbjct 22 LGDEIGKGAYGRVYIGLDLENGDFVAIKQVSLENIGQE 59
> Hs19923746
Length=337
Score = 27.7 bits (60), Expect = 9.0, Method: Compositional matrix adjust.
Identities = 17/48 (35%), Positives = 29/48 (60%), Gaps = 2/48 (4%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLICEIRLEGAL 154
++G G G+++LAI+ +G VA+K E ++ RH L+ E +L L
Sbjct 22 KIGSGSFGDIYLAINITNGEEVAVKLESQKA-RHP-QLLYESKLYKIL 67
> Hs4505599
Length=525
Score = 27.7 bits (60), Expect = 9.2, Method: Composition-based stats.
Identities = 19/59 (32%), Positives = 34/59 (57%), Gaps = 11/59 (18%)
Query 97 FRRFEVFLDTRLGKGGNGEVFLAIDRRHGGPVAIK-----REP-KEMLRHEFSLICEIR 149
+ R+E ++G+G +G VF A D G VAIK ++P KE++ +E ++ E++
Sbjct 249 YTRYE-----KIGQGASGTVFTATDVALGQEVAIKQINLQKQPKKELIINEILVMKELK 302
> Hs18596354
Length=304
Score = 27.7 bits (60), Expect = 9.3, Method: Compositional matrix adjust.
Identities = 17/39 (43%), Positives = 23/39 (58%), Gaps = 0/39 (0%)
Query 107 RLGKGGNGEVFLAIDRRHGGPVAIKREPKEMLRHEFSLI 145
RLG G EV LA +R VA+K PK+ LR + +L+
Sbjct 103 RLGSGAFSEVVLAQERGSAHLVALKCIPKKALRGKEALV 141
Lambda K H
0.324 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2136300674
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40