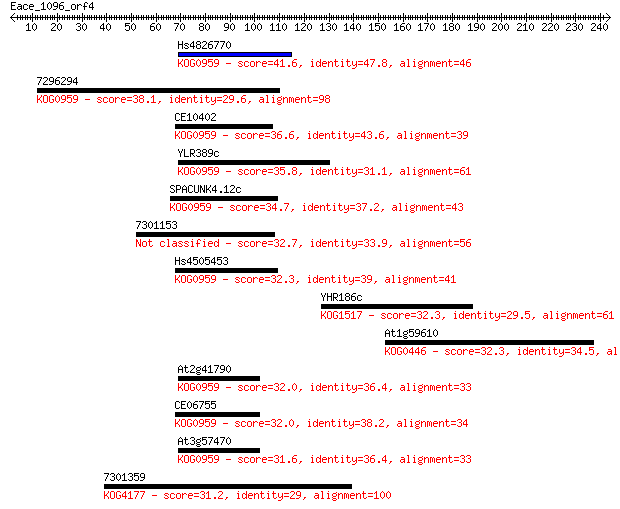
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_1096_orf4
Length=243
Score E
Sequences producing significant alignments: (Bits) Value
Hs4826770 41.6 0.001
7296294 38.1 0.017
CE10402 36.6 0.044
YLR389c 35.8 0.086
SPACUNK4.12c 34.7 0.20
7301153 32.7 0.62
Hs4505453 32.3 0.86
YHR186c 32.3 1.0
At1g59610 32.3 1.0
At2g41790 32.0 1.2
CE06755 32.0 1.3
At3g57470 31.6 1.4
7301359 31.2 2.0
> Hs4826770
Length=1019
Score = 41.6 bits (96), Expect = 0.001, Method: Composition-based stats.
Identities = 22/46 (47%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Query 69 KFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENMAISFRMGR 114
KFGTGN TL RP +EGIDV + + HS Y MA+ +GR
Sbjct 217 KFGTGNKYTLETRPNQEGIDVRQELLKFHSAYYSSNLMAVCV-LGR 261
> 7296294
Length=990
Score = 38.1 bits (87), Expect = 0.017, Method: Composition-based stats.
Identities = 29/102 (28%), Positives = 42/102 (41%), Gaps = 31/102 (30%)
Query 12 VMAVQSEHDKNTPD----LERVALHLALQYLPDSGTEGTGGSGGDSDGDSAGDTCVSPVS 67
+ AV SEH+KN P +++V HLA PD
Sbjct 156 INAVNSEHEKNLPSDLWRIKQVNRHLAK---PDHAYS----------------------- 189
Query 68 GKFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENMAIS 109
KFG+GN TL E P+ + IDV + + H + Y M ++
Sbjct 190 -KFGSGNKTTLSEIPKSKNIDVRDELLKFHKQWYSANIMCLA 230
> CE10402
Length=1067
Score = 36.6 bits (83), Expect = 0.044, Method: Composition-based stats.
Identities = 17/39 (43%), Positives = 25/39 (64%), Gaps = 0/39 (0%)
Query 68 GKFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENM 106
GKFGTGN +TL E R++GI+ ++ + H K Y + M
Sbjct 237 GKFGTGNKQTLLEDARKKGIEPRDALLQFHKKWYSSDIM 275
> YLR389c
Length=988
Score = 35.8 bits (81), Expect = 0.086, Method: Composition-based stats.
Identities = 19/61 (31%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Query 69 KFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENMAISFRMGRIEGDSQPQQQYGLW 128
KF TGN+ETL P+ G++V + + H Y M + +GR + D+ Y L+
Sbjct 227 KFSTGNIETLGTLPKENGLNVRDELLKFHKNFYSANLMKLCI-LGREDLDTLSDWTYDLF 285
Query 129 E 129
+
Sbjct 286 K 286
> SPACUNK4.12c
Length=969
Score = 34.7 bits (78), Expect = 0.20, Method: Composition-based stats.
Identities = 16/43 (37%), Positives = 24/43 (55%), Gaps = 0/43 (0%)
Query 66 VSGKFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENMAI 108
V KF TGN+ETL + P+ G+DV + + + K Y M +
Sbjct 174 VFSKFNTGNIETLGDVPKELGLDVRQELLKFYDKYYSANIMKL 216
> 7301153
Length=1116
Score = 32.7 bits (73), Expect = 0.62, Method: Compositional matrix adjust.
Identities = 19/56 (33%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Query 52 DSDGDSAGDTCVSPVSGKFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENMA 107
D +G +AGDT + SG GTGN +R G+ L R + C++P N++
Sbjct 520 DGNGSTAGDTSIGKDSGSLGTGN----SSGGKRYGLPPLNGPRNACAICHIPYNLS 571
> Hs4505453
Length=1219
Score = 32.3 bits (72), Expect = 0.86, Method: Composition-based stats.
Identities = 16/41 (39%), Positives = 20/41 (48%), Gaps = 0/41 (0%)
Query 68 GKFGTGNLETLCERPRREGIDVLASMRRLHSKCYVPENMAI 108
GKF GN ETL PR+ ID A +R + Y M +
Sbjct 409 GKFFWGNAETLKHEPRKNNIDTHARLREFWMRYYSSHYMTL 449
> YHR186c
Length=1557
Score = 32.3 bits (72), Expect = 1.0, Method: Composition-based stats.
Identities = 18/61 (29%), Positives = 30/61 (49%), Gaps = 0/61 (0%)
Query 127 LWEVIELISAAVRERASSASPEAPGASLLQRQTVSQKEETHKEGPSEARTSTSTPAGAHQ 186
+W+ +L V + AP A+ L+ Q + Q++ET + G S + T AG+ Q
Sbjct 421 MWKSWDLAMDEVLTKIVIDLKNAPPATALESQMILQQQETLQNGGSSKSNAQDTKAGSIQ 480
Query 187 T 187
T
Sbjct 481 T 481
> At1g59610
Length=920
Score = 32.3 bits (72), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 29/90 (32%), Positives = 43/90 (47%), Gaps = 18/90 (20%)
Query 153 SLLQRQTVSQKEETHKEGPSEARTSTSTPAGAHQTLREGDNPQAASRDPTRDTAEDDGGE 212
SLL R T Q + GPS + G+ ++LR+ PQ +D ++T E G +
Sbjct 526 SLLNRATSPQPD-----GPS-------STGGSLKSLRDKLMPQDKDKDKEKETPEVSGLK 573
Query 213 KELPEGIILS---LKRD---HGWRRRLFVF 236
PEG I + +K+ +GW RR FV
Sbjct 574 TAGPEGEITAGYLMKKSAKTNGWSRRWFVL 603
> At2g41790
Length=970
Score = 32.0 bits (71), Expect = 1.2, Method: Composition-based stats.
Identities = 12/33 (36%), Positives = 22/33 (66%), Gaps = 0/33 (0%)
Query 69 KFGTGNLETLCERPRREGIDVLASMRRLHSKCY 101
KF TGN++TL RP+ +G+D + + + + + Y
Sbjct 178 KFSTGNMDTLHVRPQAKGVDTRSELIKFYEEHY 210
> CE06755
Length=980
Score = 32.0 bits (71), Expect = 1.3, Method: Composition-based stats.
Identities = 13/34 (38%), Positives = 22/34 (64%), Gaps = 0/34 (0%)
Query 68 GKFGTGNLETLCERPRREGIDVLASMRRLHSKCY 101
KFGTGN +TL E R++G++ ++ + + K Y
Sbjct 178 AKFGTGNKKTLLEEARKKGVEPRDALLQFYKKWY 211
> At3g57470
Length=989
Score = 31.6 bits (70), Expect = 1.4, Method: Composition-based stats.
Identities = 12/33 (36%), Positives = 20/33 (60%), Gaps = 0/33 (0%)
Query 69 KFGTGNLETLCERPRREGIDVLASMRRLHSKCY 101
KF TGN++TL RP G+D + + + + + Y
Sbjct 180 KFSTGNMDTLHVRPEENGVDTRSELIKFYDEHY 212
> 7301359
Length=1181
Score = 31.2 bits (69), Expect = 2.0, Method: Compositional matrix adjust.
Identities = 29/102 (28%), Positives = 40/102 (39%), Gaps = 4/102 (3%)
Query 39 PDSGTEGTGGSGGDSDGDSAGDTCVSPVSGKFGTGNLETLCERPRREGI--DVLASMRRL 96
P G E G G SD ++ VSG + L L E RE I D+LA M
Sbjct 867 PAQGAEANGAEGSSSDDLLPDADTITNVSGFLSSQQLHHLIELFEREQITLDILAEMG-- 924
Query 97 HSKCYVPENMAISFRMGRIEGDSQPQQQYGLWEVIELISAAV 138
H A FR ++G +Q + G+ + L + V
Sbjct 925 HDDLKQVGVSAYGFRHKILKGIAQLRSTTGIGNNVNLCTLLV 966
Lambda K H
0.311 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4867742342
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40