bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_1784_orf1
Length=158
Score E
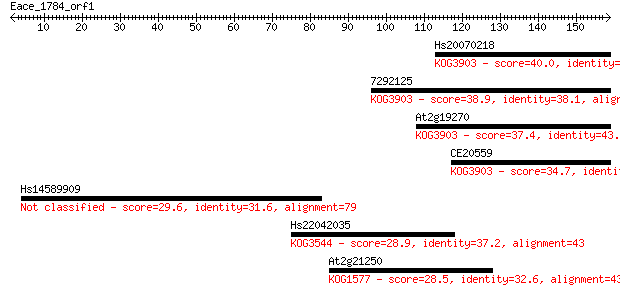
Sequences producing significant alignments: (Bits) Value
Hs20070218 40.0 0.002
7292125 38.9 0.005
At2g19270 37.4 0.012
CE20559 34.7 0.070
Hs14589909 29.6 2.7
Hs22042035 28.9 4.6
At2g21250 28.5 5.3
> Hs20070218
Length=491
Score = 40.0 bits (92), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 18/46 (39%), Positives = 29/46 (63%), Gaps = 0/46 (0%)
Query 113 EPKVMQKRKHQINWLAAEAHDKELEMLEMTARTRQSKYQTAMKYGW 158
+P Q+RKHQI +L +A ++ELE+ + + S+ QT KYG+
Sbjct 446 QPTGQQRRKHQITYLIHQAKERELELKNTWSENKLSRRQTQAKYGF 491
> 7292125
Length=472
Score = 38.9 bits (89), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 24/63 (38%), Positives = 35/63 (55%), Gaps = 1/63 (1%)
Query 96 LATKVWNAEEGVVTETLEPKVMQKRKHQINWLAAEAHDKELEMLEMTARTRQSKYQTAMK 155
LA+ GV+T+ EP +RKHQI +LA +A E E+ M + RQ++ T K
Sbjct 411 LASSTTFQPTGVLTDE-EPVAGTRRKHQITYLAHKAKANEAELQAMWSANRQTRRATQSK 469
Query 156 YGW 158
YG+
Sbjct 470 YGF 472
> At2g19270
Length=359
Score = 37.4 bits (85), Expect = 0.012, Method: Compositional matrix adjust.
Identities = 22/53 (41%), Positives = 30/53 (56%), Gaps = 2/53 (3%)
Query 108 VTETLEPKV--MQKRKHQINWLAAEAHDKELEMLEMTARTRQSKYQTAMKYGW 158
VT + + KV + KRKHQI L + KE E+ E +R +K +T KYGW
Sbjct 307 VTSSSKGKVSKLHKRKHQITALFMDMKHKESELTERRSRGLLTKAETQAKYGW 359
> CE20559
Length=405
Score = 34.7 bits (78), Expect = 0.070, Method: Compositional matrix adjust.
Identities = 17/42 (40%), Positives = 26/42 (61%), Gaps = 0/42 (0%)
Query 117 MQKRKHQINWLAAEAHDKELEMLEMTARTRQSKYQTAMKYGW 158
M +RKHQI +LA+ A +E ++ + A +QSK KYG+
Sbjct 364 MSRRKHQITYLASLAVSREEQLKDQWADQKQSKRMARQKYGF 405
> Hs14589909
Length=846
Score = 29.6 bits (65), Expect = 2.7, Method: Composition-based stats.
Identities = 25/81 (30%), Positives = 39/81 (48%), Gaps = 11/81 (13%)
Query 4 DYGAAEAADYGAAANEAAAAAATAA--GAAWTLKEEEAAIEELGIEKTIISGISQAELRD 61
D+G + AD+G AA A A + G + + E AA+E+ G G +Q L D
Sbjct 150 DHGDVKLADFGVAAKITATIAKRKSFIGTPYWMAPEVAAVEKNG-------GYNQ--LCD 200
Query 62 LRRMGATSVSGATLQNPDWQL 82
+ +G T++ LQ P + L
Sbjct 201 IWAVGITAIELGELQPPMFDL 221
> Hs22042035
Length=1407
Score = 28.9 bits (63), Expect = 4.6, Method: Composition-based stats.
Identities = 16/43 (37%), Positives = 20/43 (46%), Gaps = 4/43 (9%)
Query 75 LQNPDWQLTSQTGPQKKGKLRLATKVWNAEEGVVTETLEPKVM 117
L P W T +GP L + K WN + V LEPKV+
Sbjct 1316 LNTPRWTSTQTSGP----GLPIGFKGWNGQIFKVNTLLEPKVL 1354
> At2g21250
Length=309
Score = 28.5 bits (62), Expect = 5.3, Method: Compositional matrix adjust.
Identities = 14/43 (32%), Positives = 26/43 (60%), Gaps = 3/43 (6%)
Query 85 QTGPQKKGKLRLATKVWNAEEGVVTETLEPKVMQKRKHQINWL 127
+TG K+ L + TK+WN++ G V E + + +K Q+++L
Sbjct 62 KTGLVKREDLFITTKLWNSDHGHVIEACKDSL---KKLQLDYL 101
Lambda K H
0.308 0.120 0.339
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2100092188
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40