bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_1919_orf2
Length=187
Score E
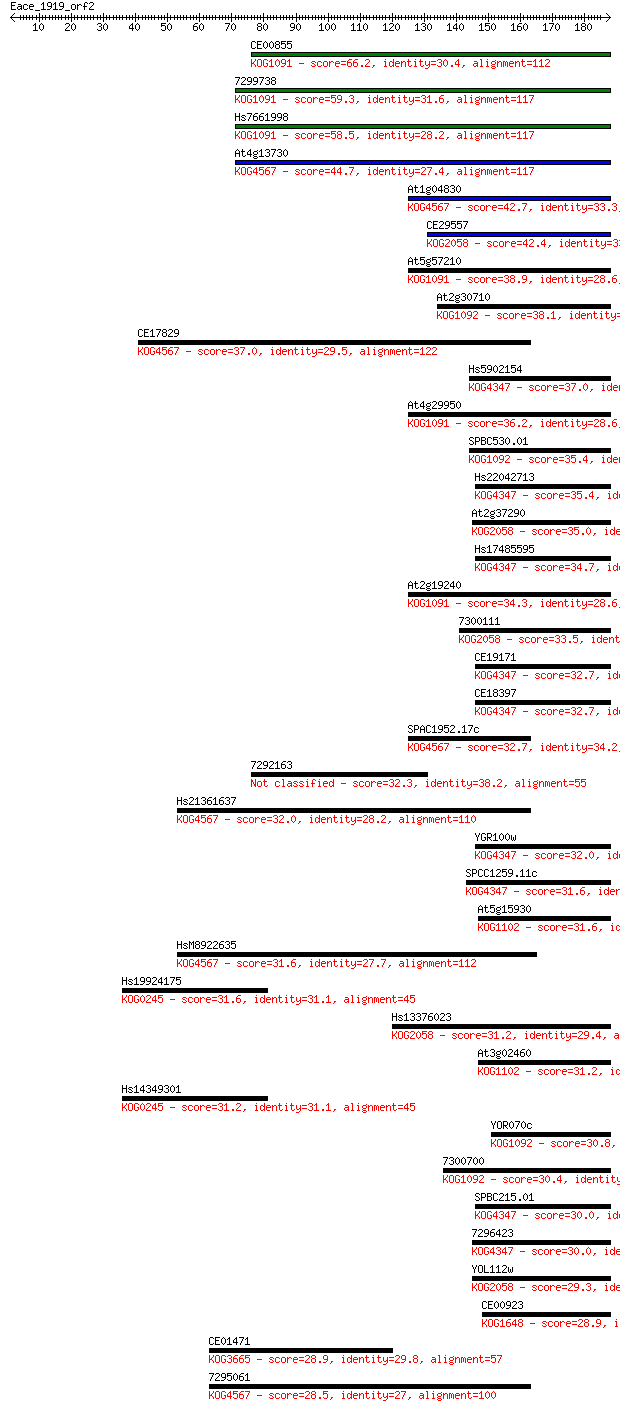
Sequences producing significant alignments: (Bits) Value
CE00855 66.2 4e-11
7299738 59.3 4e-09
Hs7661998 58.5 7e-09
At4g13730 44.7 1e-04
At1g04830 42.7 4e-04
CE29557 42.4 5e-04
At5g57210 38.9 0.007
At2g30710 38.1 0.011
CE17829 37.0 0.022
Hs5902154 37.0 0.026
At4g29950 36.2 0.038
SPBC530.01 35.4 0.065
Hs22042713 35.4 0.070
At2g37290 35.0 0.078
Hs17485595 34.7 0.11
At2g19240 34.3 0.16
7300111 33.5 0.28
CE19171 32.7 0.42
CE18397 32.7 0.43
SPAC1952.17c 32.7 0.47
7292163 32.3 0.57
Hs21361637 32.0 0.71
YGR100w 32.0 0.76
SPCC1259.11c 31.6 0.85
At5g15930 31.6 0.93
HsM8922635 31.6 0.95
Hs19924175 31.6 0.98
Hs13376023 31.2 1.2
At3g02460 31.2 1.3
Hs14349301 31.2 1.4
YOR070c 30.8 1.5
7300700 30.4 1.9
SPBC215.01 30.0 3.0
7296423 30.0 3.1
YOL112w 29.3 4.4
CE00923 28.9 5.9
CE01471 28.9 6.7
7295061 28.5 7.8
> CE00855
Length=585
Score = 66.2 bits (160), Expect = 4e-11, Method: Compositional matrix adjust.
Identities = 34/113 (30%), Positives = 59/113 (52%), Gaps = 2/113 (1%)
Query 76 WARLLRLLSGETLEDWAHQVQEQRQRYRSLRQQQQLSASQLA-TLDPQRFHPLAATAGNP 134
W +LR L ET DW + R YR+ ++ + + DP+ +PLA+ NP
Sbjct 50 WRLVLRCLPYET-SDWEISLSRSRNLYRAHKENHLIDPHDTKFSQDPEFNNPLASIEQNP 108
Query 135 WSQRQADEELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
W+ D +L++ I +D+ RT+PE + F R+ + +L +AK++ V+Y
Sbjct 109 WNTFFEDNDLRDIIGKDVSRTFPEIEFFQNTSIRQMMSDILLVYAKEHPFVNY 161
> 7299738
Length=654
Score = 59.3 bits (142), Expect = 4e-09, Method: Compositional matrix adjust.
Identities = 37/117 (31%), Positives = 62/117 (52%), Gaps = 3/117 (2%)
Query 71 LRRLHWARLLRLLSGETLEDWAHQVQEQRQRYRSLRQQQQLSASQLATLDPQRFHPLAAT 130
R +HWA LLR+L+ E W Q +QR RY R + QLA +D PL+ +
Sbjct 76 FRSVHWALLLRVLTSEH-RSWTSQRLQQRVRYDKFRADYVRNPHQLA-VDCND-DPLSQS 132
Query 131 AGNPWSQRQADEELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
+ W+Q +D++L I +D+ RT+P F K + ++ +LF +A+++ + Y
Sbjct 133 TQSVWNQYFSDQDLFAVIRQDVVRTFPGVDFFRKPLVQNAMVNILFYYAREHPYMCY 189
> Hs7661998
Length=795
Score = 58.5 bits (140), Expect = 7e-09, Method: Compositional matrix adjust.
Identities = 33/118 (27%), Positives = 63/118 (53%), Gaps = 2/118 (1%)
Query 71 LRRLHWARLLRLLSGETLEDWAHQVQEQRQRYRSLRQQQQLSASQLATL-DPQRFHPLAA 129
R + W L +L + W +++E R Y ++++ + ++ D +PL+
Sbjct 86 FRSICWKLFLCVLPQDK-SQWISRIEELRAWYSNIKEIHITNPRKVVGQQDLMINNPLSQ 144
Query 130 TAGNPWSQRQADEELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
G+ W++ D+EL+ I +D++RT+PE Q F +E R+ L VLF +A++N + Y
Sbjct 145 DEGSLWNKFFQDKELRSMIEQDVKRTFPEMQFFQQENVRKILTDVLFCYARENEQLLY 202
> At4g13730
Length=449
Score = 44.7 bits (104), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 32/146 (21%), Positives = 67/146 (45%), Gaps = 30/146 (20%)
Query 71 LRRLHWARLLRLLSGETLEDWAHQVQEQRQRYRSLRQQQQLSASQLA----------TLD 120
+R + W LL LS + W+ ++ ++R +Y+ +++ ++ S++ + D
Sbjct 131 IRSIVWKLLLDYLSPDR-SLWSSELAKKRSQYKQFKEELLMNPSEVTRKMDKSKGGDSND 189
Query 121 PQ--------------RFHPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLFNKE- 165
P+ HPL+ + W+ D E+ E+I RD+ RT+P+ F+ +
Sbjct 190 PKIESPGALSRSEITHEDHPLSLGTTSLWNNFFKDTEVLEQIERDVMRTHPDMHFFSGDS 249
Query 166 ----GTRRSLQRVLFTWAKKNSDVSY 187
+ +L+ +L +AK N + Y
Sbjct 250 AVAKSNQDALKNILTIFAKLNPGIRY 275
> At1g04830
Length=451
Score = 42.7 bits (99), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 37/68 (54%), Gaps = 5/68 (7%)
Query 125 HPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLFNKEGT-----RRSLQRVLFTWA 179
HPL+ + W+ D E E+I RD++RT+P+ F+ E + + S++ +L +A
Sbjct 207 HPLSLGKASIWNTYFQDTETIEQIDRDVKRTHPDIPFFSGESSFARSNQESMKNILLVFA 266
Query 180 KKNSDVSY 187
K N + Y
Sbjct 267 KLNQGIRY 274
> CE29557
Length=248
Score = 42.4 bits (98), Expect = 5e-04, Method: Compositional matrix adjust.
Identities = 19/57 (33%), Positives = 32/57 (56%), Gaps = 0/57 (0%)
Query 131 AGNPWSQRQADEELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
A W + + +E+ ++I D+ RT+P+ + EGTR++L R LF A+ V Y
Sbjct 96 ADGVWQRHEVPDEVIKQIKLDLPRTFPDNKFLKTEGTRKTLGRALFAVAEHIPSVGY 152
> At5g57210
Length=737
Score = 38.9 bits (89), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 18/64 (28%), Positives = 34/64 (53%), Gaps = 1/64 (1%)
Query 125 HPLAATAGNPWSQRQADEELQEEIWRDIERTYPER-QLFNKEGTRRSLQRVLFTWAKKNS 183
+PL+ + W + + EL++ + +D+ R YPE F G + L+R+L W K+
Sbjct 94 NPLSQNPDSTWGRFFRNAELEKTLDQDLSRLYPEHGSYFQSSGCQGMLRRILLLWCLKHP 153
Query 184 DVSY 187
++ Y
Sbjct 154 EIGY 157
> At2g30710
Length=343
Score = 38.1 bits (87), Expect = 0.011, Method: Compositional matrix adjust.
Identities = 20/55 (36%), Positives = 32/55 (58%), Gaps = 1/55 (1%)
Query 134 PWSQRQADE-ELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
P S+R DE + +I D RT P+ F +E ++SL+R+L+TWA ++ Y
Sbjct 92 PDSERSDDEINMLRQIAVDCPRTVPDVSFFQQEQVQKSLERILYTWAIRHPASGY 146
> CE17829
Length=484
Score = 37.0 bits (84), Expect = 0.022, Method: Compositional matrix adjust.
Identities = 36/128 (28%), Positives = 52/128 (40%), Gaps = 7/128 (5%)
Query 41 IVRLQRELEAAAPSQAAKEAEDSSSPAVP-FLRRLHWARLLRLLSGETLEDWAHQVQEQR 99
+ +++ L A E S VP LR L W LL L E W + EQR
Sbjct 9 LAKIEEVLLIANKKIDINELRAGCSYGVPESLRPLAWRLLLHYLPLER-HKWQTFLAEQR 67
Query 100 QRYRSLRQQQQL-----SASQLATLDPQRFHPLAATAGNPWSQRQADEELQEEIWRDIER 154
Y + +Q + S Q A + HPL+ + W D ++ +I +D+ R
Sbjct 68 DNYDQMIEQIIVEPGTASLQQSAAQNQDNDHPLSDHPTSDWQAFFQDNKVLSQIDKDVRR 127
Query 155 TYPERQLF 162
YPE Q F
Sbjct 128 LYPEIQFF 135
> Hs5902154
Length=897
Score = 37.0 bits (84), Expect = 0.026, Method: Compositional matrix adjust.
Identities = 17/44 (38%), Positives = 25/44 (56%), Gaps = 0/44 (0%)
Query 144 LQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
+ EEI RD+ R+ PE F E +L+RVL +A +N + Y
Sbjct 303 VTEEIERDLHRSLPEHPAFQNETGIAALRRVLTAYAHRNPKIGY 346
> At4g29950
Length=828
Score = 36.2 bits (82), Expect = 0.038, Method: Compositional matrix adjust.
Identities = 18/64 (28%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Query 125 HPLAATAGNPWSQRQADEELQEEIWRDIERTYPER-QLFNKEGTRRSLQRVLFTWAKKNS 183
+PL+ + W + + EL++ + +D+ R YPE F G + L+R+L W K+
Sbjct 80 NPLSQNPDSTWGRFFRNAELEKTLDQDLSRLYPEHWSYFQAPGCQGMLRRILLLWCLKHP 139
Query 184 DVSY 187
+ Y
Sbjct 140 EYGY 143
> SPBC530.01
Length=514
Score = 35.4 bits (80), Expect = 0.065, Method: Compositional matrix adjust.
Identities = 17/48 (35%), Positives = 26/48 (54%), Gaps = 4/48 (8%)
Query 144 LQEEIWR----DIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
L + IWR D+ RT P L+ T+R L+R+L+ WA ++ Y
Sbjct 268 LDQTIWRQIVLDVPRTNPSILLYQNPLTQRMLERILYVWASRHPASGY 315
> Hs22042713
Length=1266
Score = 35.4 bits (80), Expect = 0.070, Method: Compositional matrix adjust.
Identities = 17/42 (40%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 146 EEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
EEI RD+ R+ PE F E +L+RVL +A +N ++ Y
Sbjct 558 EEIERDLHRSLPEHPAFQNEMGIAALRRVLTAYAFRNPNIGY 599
> At2g37290
Length=544
Score = 35.0 bits (79), Expect = 0.078, Method: Compositional matrix adjust.
Identities = 17/43 (39%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query 145 QEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
+++I +DI RT+P N+ G R SL+R+L +A N V Y
Sbjct 234 KKQIEKDIPRTFPGHPALNENG-RDSLRRILLAYACHNPSVGY 275
> Hs17485595
Length=796
Score = 34.7 bits (78), Expect = 0.11, Method: Composition-based stats.
Identities = 16/42 (38%), Positives = 24/42 (57%), Gaps = 0/42 (0%)
Query 146 EEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
EEI RD+ R+ PE F + +L+RVL +A +N + Y
Sbjct 530 EEIERDLRRSLPEHPAFQSDTGISALRRVLTAYAYRNPKIGY 571
> At2g19240
Length=840
Score = 34.3 bits (77), Expect = 0.16, Method: Compositional matrix adjust.
Identities = 18/64 (28%), Positives = 32/64 (50%), Gaps = 1/64 (1%)
Query 125 HPLAATAGNPWSQRQADEELQEEIWRDIERTYPER-QLFNKEGTRRSLQRVLFTWAKKNS 183
+PL+ + W Q + EL++ + +D+ R YPE F + L+R+L W K+
Sbjct 84 NPLSQNPNSTWGQFFRNAELEKTLDQDLSRLYPEHWCYFQTPRYQGMLRRILLLWCLKHP 143
Query 184 DVSY 187
+ Y
Sbjct 144 EYGY 147
> 7300111
Length=299
Score = 33.5 bits (75), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 16/47 (34%), Positives = 26/47 (55%), Gaps = 2/47 (4%)
Query 141 DEELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
D+E+ + I D+ RT+P+ F+ + R L +L +A N DV Y
Sbjct 76 DKEISDSISIDLPRTFPDNIHFDMKKQR--LYNILIAYAHHNRDVGY 120
> CE19171
Length=1270
Score = 32.7 bits (73), Expect = 0.42, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 146 EEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
EEI RD+ R+ PE F + +L+R+L +A +N ++ Y
Sbjct 555 EEIERDLHRSLPEHPAFQQGPGIDALRRILTAYAFRNPNIGY 596
> CE18397
Length=1268
Score = 32.7 bits (73), Expect = 0.43, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 146 EEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
EEI RD+ R+ PE F + +L+R+L +A +N ++ Y
Sbjct 555 EEIERDLHRSLPEHPAFQQGPGIDALRRILTAYAFRNPNIGY 596
> SPAC1952.17c
Length=541
Score = 32.7 bits (73), Expect = 0.47, Method: Compositional matrix adjust.
Identities = 13/38 (34%), Positives = 21/38 (55%), Gaps = 0/38 (0%)
Query 125 HPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLF 162
HPL + + W + D ++ E+I +DI RT P+ F
Sbjct 18 HPLNTSDDSKWKEYFDDNQILEQIDKDIRRTLPDLSFF 55
> 7292163
Length=307
Score = 32.3 bits (72), Expect = 0.57, Method: Compositional matrix adjust.
Identities = 21/62 (33%), Positives = 30/62 (48%), Gaps = 9/62 (14%)
Query 76 WARLLRLLSG------ETLEDWAHQVQEQRQR-YRSLRQQQQLSASQLATLDPQRFHPLA 128
WA+L +L+ + +EDW R R + R +LS S+ T DP+ F PL
Sbjct 208 WAKLAHMLNSVPQGAVKNVEDWKQTFDAWRYRIFMYTRYNSKLSMSE--TTDPKNFKPLT 265
Query 129 AT 130
AT
Sbjct 266 AT 267
> Hs21361637
Length=400
Score = 32.0 bits (71), Expect = 0.71, Method: Compositional matrix adjust.
Identities = 31/120 (25%), Positives = 52/120 (43%), Gaps = 11/120 (9%)
Query 53 PSQAAKEAEDSSSPAVPF---LRRLHWARLLRLLSGETLEDWAHQVQEQRQRY----RSL 105
PS A ++ + S +P LR L W LL L E W + +QR+ Y R +
Sbjct 19 PSIALEKLRELSFSGIPCEGGLRCLCWKILLNYLPLER-ASWTSILAKQRELYAQFLREM 77
Query 106 RQQQQLSASQLA-TLDPQRF--HPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLF 162
Q ++ + + + + F HPL + W+ D E+ +I +D+ R P+ F
Sbjct 78 IIQPGIAKANMGVSREDVTFEDHPLNPNPDSRWNTYFKDNEVLLQIDKDVRRLCPDISFF 137
> YGR100w
Length=950
Score = 32.0 bits (71), Expect = 0.76, Method: Composition-based stats.
Identities = 15/42 (35%), Positives = 25/42 (59%), Gaps = 0/42 (0%)
Query 146 EEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
+EI +D++R+ PE + E + L+ VL ++ KN DV Y
Sbjct 287 DEIEKDLKRSLPEYSAYQTEEGIQRLRNVLTAYSWKNPDVGY 328
> SPCC1259.11c
Length=720
Score = 31.6 bits (70), Expect = 0.85, Method: Composition-based stats.
Identities = 13/45 (28%), Positives = 25/45 (55%), Gaps = 0/45 (0%)
Query 143 ELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
E EEI +D+ R+ P+ + +L+R+L +++ N +V Y
Sbjct 256 EYSEEIEKDLTRSLPDYPAYQSPTGINTLRRILLFYSETNKEVGY 300
> At5g15930
Length=356
Score = 31.6 bits (70), Expect = 0.93, Method: Compositional matrix adjust.
Identities = 17/43 (39%), Positives = 23/43 (53%), Gaps = 2/43 (4%)
Query 147 EIWRDIERTYPERQLFNKE--GTRRSLQRVLFTWAKKNSDVSY 187
+I RDI RT+P F K +RSL VL ++ + DV Y
Sbjct 126 DIIRDISRTFPSHVFFQKRHGPGQRSLYNVLKAYSVYDRDVGY 168
> HsM8922635
Length=275
Score = 31.6 bits (70), Expect = 0.95, Method: Compositional matrix adjust.
Identities = 31/122 (25%), Positives = 53/122 (43%), Gaps = 11/122 (9%)
Query 53 PSQAAKEAEDSSSPAVPF---LRRLHWARLLRLLSGETLEDWAHQVQEQRQRY----RSL 105
PS A ++ + S +P LR L W LL L E W + +QR+ Y R +
Sbjct 19 PSIALEKLRELSFSGIPCEGGLRCLCWKILLNYLPLER-ASWTSILAKQRELYAQFLREM 77
Query 106 RQQQQLSASQLA-TLDPQRF--HPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLF 162
Q ++ + + + + F HPL + W+ D E+ +I +D+ R P+ F
Sbjct 78 IIQPGIAKANMGVSREDVTFEDHPLNPNPDSRWNTYFKDNEVLLQIDKDVRRLCPDISFF 137
Query 163 NK 164
+
Sbjct 138 QR 139
> Hs19924175
Length=1690
Score = 31.6 bits (70), Expect = 0.98, Method: Compositional matrix adjust.
Identities = 14/45 (31%), Positives = 24/45 (53%), Gaps = 0/45 (0%)
Query 36 RQQLDIVRLQRELEAAAPSQAAKEAEDSSSPAVPFLRRLHWARLL 80
RQ+LD++R + A PS ++ ++ + PF R W RL+
Sbjct 806 RQRLDLMREMYDRAAEVPSSVIEDCDNVVTGGDPFYDRFPWFRLV 850
> Hs13376023
Length=257
Score = 31.2 bits (69), Expect = 1.2, Method: Compositional matrix adjust.
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 11/71 (15%)
Query 120 DPQRFHPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLFNKE---GTRRSLQRVLF 176
+P +H L NP L++ I D+ RT+P+ F K +R+L VL
Sbjct 4 NPGYYHQLLQGERNP--------RLEDAIRTDLNRTFPDNVKFRKTTDPCLQRTLYNVLL 55
Query 177 TWAKKNSDVSY 187
+ N V Y
Sbjct 56 AYGHHNQGVGY 66
> At3g02460
Length=304
Score = 31.2 bits (69), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 17/43 (39%), Positives = 23/43 (53%), Gaps = 2/43 (4%)
Query 147 EIWRDIERTYPERQLFNKE--GTRRSLQRVLFTWAKKNSDVSY 187
+I RDI RT+P F K +RSL VL ++ + DV Y
Sbjct 129 DIIRDISRTFPSHVFFQKRHGPGQRSLYNVLKAYSVYDRDVGY 171
> Hs14349301
Length=1770
Score = 31.2 bits (69), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 14/45 (31%), Positives = 25/45 (55%), Gaps = 0/45 (0%)
Query 36 RQQLDIVRLQRELEAAAPSQAAKEAEDSSSPAVPFLRRLHWARLL 80
+Q+LD++R + S A E+E + + + PF R HW +L+
Sbjct 800 KQRLDLMREMYDRAGEMASSAQDESETTVTGSDPFYDRFHWFKLV 844
> YOR070c
Length=637
Score = 30.8 bits (68), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 14/37 (37%), Positives = 21/37 (56%), Gaps = 0/37 (0%)
Query 151 DIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
DI RT P L+ + + SLQR+L+ WA ++ Y
Sbjct 340 DIPRTNPHIPLYQFKSVQNSLQRILYLWAIRHPASGY 376
> 7300700
Length=545
Score = 30.4 bits (67), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 14/52 (26%), Positives = 27/52 (51%), Gaps = 0/52 (0%)
Query 136 SQRQADEELQEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
SQ + ++ +I D+ R P+ LF ++ + +R+LF WA ++ Y
Sbjct 292 SQDETQQDTYRQIHIDVPRMNPQIPLFQQQLVQEMFERILFIWAIRHPASGY 343
> SPBC215.01
Length=827
Score = 30.0 bits (66), Expect = 3.0, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 24/42 (57%), Gaps = 0/42 (0%)
Query 146 EEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
EEI +D+ R+ PE + E +L+ VL ++ KN +V Y
Sbjct 263 EEIEKDLGRSLPEYPAYQNEEGINALRNVLVAFSWKNQEVGY 304
> 7296423
Length=1291
Score = 30.0 bits (66), Expect = 3.1, Method: Compositional matrix adjust.
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 0/43 (0%)
Query 145 QEEIWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
+EI RD+ R+ PE F +L+RVL +A +N V Y
Sbjct 505 HDEIDRDLPRSLPEHPAFQSTDGIGALRRVLQAYALRNPQVGY 547
> YOL112w
Length=492
Score = 29.3 bits (64), Expect = 4.4, Method: Compositional matrix adjust.
Identities = 17/50 (34%), Positives = 27/50 (54%), Gaps = 7/50 (14%)
Query 145 QEEIWRDIERTYPERQLFNKEGTR-------RSLQRVLFTWAKKNSDVSY 187
E I RD+ RT+P+ F+KE + RSL+RVL ++ + + Y
Sbjct 191 MEAIERDLYRTFPDNIHFHKESFQNGEPAIIRSLRRVLMAFSVYDKTIGY 240
> CE00923
Length=807
Score = 28.9 bits (63), Expect = 5.9, Method: Compositional matrix adjust.
Identities = 12/40 (30%), Positives = 23/40 (57%), Gaps = 0/40 (0%)
Query 148 IWRDIERTYPERQLFNKEGTRRSLQRVLFTWAKKNSDVSY 187
I +D+ER F+ + SL+RV++T+ ++N + Y
Sbjct 595 IDKDVERCDRNLMFFSNKDNLESLRRVMYTYVRRNLEEGY 634
> CE01471
Length=799
Score = 28.9 bits (63), Expect = 6.7, Method: Compositional matrix adjust.
Identities = 17/59 (28%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Query 63 SSSPAVPFLRRLHWARLLRLLSGETLEDWAHQV--QEQRQRYRSLRQQQQLSASQLATL 119
S+ A+P L RL R+ L+ + L ++ H+V + LRQ+++L+ L T+
Sbjct 10 CSTSAIPRLARLATERIAELIENDALPEFDHKVIASCSNDIFEVLRQKKKLNHQTLQTM 68
> 7295061
Length=403
Score = 28.5 bits (62), Expect = 7.8, Method: Compositional matrix adjust.
Identities = 27/115 (23%), Positives = 45/115 (39%), Gaps = 16/115 (13%)
Query 63 SSSPAVPFLRRLHWARLLRLLSGETLEDWAHQVQEQRQRYRSLRQQQQLS---ASQLATL 119
+ P V R L W LL L G W + ++R Y+ ++ L +S A++
Sbjct 33 NGVPDVQSFRALSWKLLLGYL-GPRRSSWTTTLAQKRALYKQFIEELVLPPGHSSNRASV 91
Query 120 DPQ------------RFHPLAATAGNPWSQRQADEELQEEIWRDIERTYPERQLF 162
D + HPL+ + W+ D E +I +D+ R P+ F
Sbjct 92 DGGDGDKVDSGGVGLQDHPLSEGPESAWNTFFNDNEFLLQIDKDVRRLCPDISFF 146
Lambda K H
0.316 0.126 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3058524910
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40