bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_1946_orf1
Length=84
Score E
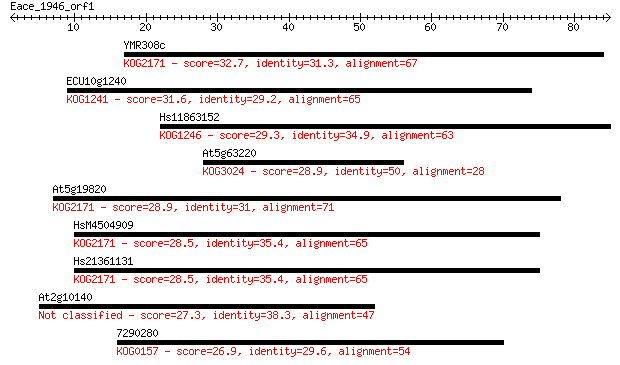
Sequences producing significant alignments: (Bits) Value
YMR308c 32.7 0.17
ECU10g1240 31.6 0.36
Hs11863152 29.3 1.9
At5g63220 28.9 2.5
At5g19820 28.9 2.6
HsM4504909 28.5 3.3
Hs21361131 28.5 3.6
At2g10140 27.3 6.2
7290280 26.9 8.2
> YMR308c
Length=1089
Score = 32.7 bits (73), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 21/69 (30%), Positives = 35/69 (50%), Gaps = 2/69 (2%)
Query 17 LLEGTLSDDCGVRRQAEASLSE--LKETAADMFICYLLETLSMHPEEQLRLQAAVLLRSL 74
+++ S D +R AE +LSE + E + + +L E + + + +AVL R L
Sbjct 15 IVQAFASPDNQIRSVAEKALSEEWITENNIEYLLTFLAEQAAFSQDTTVAALSAVLFRKL 74
Query 75 AFRGPLSSK 83
A + P SSK
Sbjct 75 ALKAPPSSK 83
> ECU10g1240
Length=854
Score = 31.6 bits (70), Expect = 0.36, Method: Composition-based stats.
Identities = 19/66 (28%), Positives = 34/66 (51%), Gaps = 1/66 (1%)
Query 9 MDASKFCSLLEGTLSDDCGVRRQAEASLSELKETAADMFICYLLETL-SMHPEEQLRLQA 67
MD + L L D G ++AEA + EL+ + FI L+E + +QLR+ +
Sbjct 27 MDRRGLEACLYEVLKSDPGAGKRAEARILELQSGDFERFISMLVEVFCDLKSNDQLRMVS 86
Query 68 AVLLRS 73
++L++
Sbjct 87 GIILKN 92
> Hs11863152
Length=1246
Score = 29.3 bits (64), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 22/66 (33%), Positives = 33/66 (50%), Gaps = 7/66 (10%)
Query 22 LSDDCGVRRQAEASLSELKETAADMFICYLLETLSMHPEEQLRLQAAVLLRSLAF---RG 78
L+D + R A+ L++L+E + YLL S+ PEE RL+ VL+ +G
Sbjct 680 LADMLRIPRTAQDRLAKLQEA----YCQYLLSYDSLSPEEHRRLEKEVLMEKEILEKRKG 735
Query 79 PLSSKT 84
PL T
Sbjct 736 PLEGHT 741
> At5g63220
Length=316
Score = 28.9 bits (63), Expect = 2.5, Method: Compositional matrix adjust.
Identities = 14/28 (50%), Positives = 18/28 (64%), Gaps = 0/28 (0%)
Query 28 VRRQAEASLSELKETAADMFICYLLETL 55
+++QAE EL E+ FI YLLETL
Sbjct 233 IKKQAETKNPELSESDLIQFISYLLETL 260
> At5g19820
Length=1116
Score = 28.9 bits (63), Expect = 2.6, Method: Compositional matrix adjust.
Identities = 22/71 (30%), Positives = 33/71 (46%), Gaps = 0/71 (0%)
Query 7 LKMDASKFCSLLEGTLSDDCGVRRQAEASLSELKETAADMFICYLLETLSMHPEEQLRLQ 66
L D++ F +L+ +S R AE+ + K++ D L L + P + R
Sbjct 17 LGSDSAPFETLISHLMSSSNEQRSSAESLFNLAKQSNPDTLSLKLAHLLQLSPHPEGRAM 76
Query 67 AAVLLRSLAFR 77
AAVLLR L R
Sbjct 77 AAVLLRKLLTR 87
> HsM4504909
Length=1097
Score = 28.5 bits (62), Expect = 3.3, Method: Compositional matrix adjust.
Identities = 23/66 (34%), Positives = 34/66 (51%), Gaps = 4/66 (6%)
Query 10 DASKFCSLLEGTLSDDCGVRRQAEASLSELKETAADMFICYLLETL-SMHPEEQLRLQAA 68
+ +F LL LS D VR+QAE + + + I +LL+ + + E+ R AA
Sbjct 7 EQQQFYLLLGNLLSPDNVVRKQAEETYENIPGQSK---ITFLLQAIRNTTAAEEARQMAA 63
Query 69 VLLRSL 74
VLLR L
Sbjct 64 VLLRRL 69
> Hs21361131
Length=1097
Score = 28.5 bits (62), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 23/66 (34%), Positives = 34/66 (51%), Gaps = 4/66 (6%)
Query 10 DASKFCSLLEGTLSDDCGVRRQAEASLSELKETAADMFICYLLETL-SMHPEEQLRLQAA 68
+ +F LL LS D VR+QAE + + + I +LL+ + + E+ R AA
Sbjct 7 EQQQFYLLLGNLLSPDNVVRKQAEETYENIPGQSK---ITFLLQAIRNTTAAEEARQMAA 63
Query 69 VLLRSL 74
VLLR L
Sbjct 64 VLLRRL 69
> At2g10140
Length=663
Score = 27.3 bits (59), Expect = 6.2, Method: Composition-based stats.
Identities = 18/53 (33%), Positives = 27/53 (50%), Gaps = 9/53 (16%)
Query 5 KPLKMDASKFCSLLEG---TLSDDCGVRRQAEASL---SELKETAADMFICYL 51
+P++ ++KF LLEG TL D C R+ ++ L S L A +CY
Sbjct 157 EPVRNHSNKFYDLLEGANNTLYDYC---REGQSQLSLASRLMHNKAGTIVCYF 206
> 7290280
Length=501
Score = 26.9 bits (58), Expect = 8.2, Method: Composition-based stats.
Identities = 16/54 (29%), Positives = 26/54 (48%), Gaps = 1/54 (1%)
Query 16 SLLEGTLSDDCGVRRQAEASLSELKETAADMFICYLLETLSMHPEEQLRLQAAV 69
S ++G D +R + + + E +T I + L +S HPE Q RLQ +
Sbjct 289 STIDGAPLSDEDIREEVDTFMFEGHDTTTSA-ISFCLYEISRHPEVQQRLQQEI 341
Lambda K H
0.318 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1197406694
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40