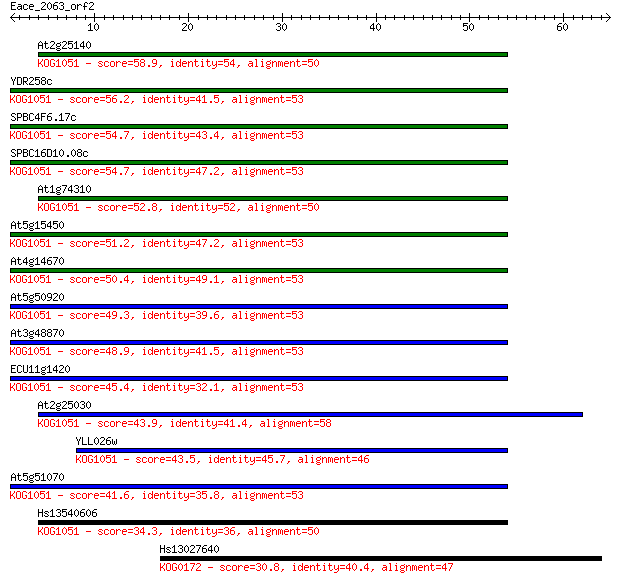
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_2063_orf2
Length=64
Score E
Sequences producing significant alignments: (Bits) Value
At2g25140 58.9 2e-09
YDR258c 56.2 1e-08
SPBC4F6.17c 54.7 4e-08
SPBC16D10.08c 54.7 4e-08
At1g74310 52.8 2e-07
At5g15450 51.2 5e-07
At4g14670 50.4 8e-07
At5g50920 49.3 2e-06
At3g48870 48.9 2e-06
ECU11g1420 45.4 3e-05
At2g25030 43.9 7e-05
YLL026w 43.5 9e-05
At5g51070 41.6 4e-04
Hs13540606 34.3 0.057
Hs13027640 30.8 0.64
> At2g25140
Length=874
Score = 58.9 bits (141), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 27/50 (54%), Positives = 37/50 (74%), Gaps = 0/50 (0%)
Query 4 LADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
L ++L RVIGQ AVK+VADA+ RAGLS N+P+ +FMF+G + VG+
Sbjct 557 LEEVLHHRVIGQDMAVKSVADAIRRSRAGLSDPNRPIASFMFMGPTGVGK 606
> YDR258c
Length=811
Score = 56.2 bits (134), Expect = 1e-08, Method: Composition-based stats.
Identities = 22/53 (41%), Positives = 39/53 (73%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
+L + + L RV+GQ +A+ A++DA+ +QRAGL+ + +P+ +FMFLG + G+
Sbjct 495 LLYMENSLKERVVGQDEAIAAISDAVRLQRAGLTSEKRPIASFMFLGPTGTGK 547
> SPBC4F6.17c
Length=803
Score = 54.7 bits (130), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 23/53 (43%), Positives = 38/53 (71%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
+L + + ++IGQ +A+KA+ADA+ + RAGL N+PL +F+FLG + VG+
Sbjct 493 LLNMEQTIGKKIIGQDEALKAIADAVRLSRAGLQNTNRPLASFLFLGPTGVGK 545
> SPBC16D10.08c
Length=905
Score = 54.7 bits (130), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 25/53 (47%), Positives = 38/53 (71%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
+L + +LS +VIGQ +AV AVA+A+ + RAGLS N+P+ +F+F G S G+
Sbjct 575 LLNMEKVLSKQVIGQNEAVTAVANAIRLSRAGLSDPNQPIASFLFCGPSGTGK 627
> At1g74310
Length=911
Score = 52.8 bits (125), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 26/50 (52%), Positives = 35/50 (70%), Gaps = 0/50 (0%)
Query 4 LADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
LAD L RV+GQ QAV AV++A+ RAGL +P G+F+FLG + VG+
Sbjct 563 LADRLHKRVVGQNQAVNAVSEAILRSRAGLGRPQQPTGSFLFLGPTGVGK 612
> At5g15450
Length=968
Score = 51.2 bits (121), Expect = 5e-07, Method: Compositional matrix adjust.
Identities = 25/53 (47%), Positives = 36/53 (67%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
+L L + L RV+GQ AV AVA+A+ RAGLS +P+ +FMF+G + VG+
Sbjct 639 LLHLEEELHKRVVGQNPAVTAVAEAIQRSRAGLSDPGRPIASFMFMGPTGVGK 691
> At4g14670
Length=623
Score = 50.4 bits (119), Expect = 8e-07, Method: Compositional matrix adjust.
Identities = 26/53 (49%), Positives = 36/53 (67%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
++ LAD L RV+GQ +AVKAVA A+ R GL +P G+F+FLG + VG+
Sbjct 525 LISLADKLHERVVGQDEAVKAVAAAILRSRVGLGRPQQPSGSFLFLGPTGVGK 577
> At5g50920
Length=929
Score = 49.3 bits (116), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 21/53 (39%), Positives = 36/53 (67%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
+L++ + L R+IGQ +AVKA++ A+ R GL N+P+ +F+F G + VG+
Sbjct 599 LLKMEETLHKRIIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTGVGK 651
> At3g48870
Length=952
Score = 48.9 bits (115), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 22/53 (41%), Positives = 36/53 (67%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
+L++ L +RVIGQ +AVKA++ A+ R GL N+P+ +F+F G + VG+
Sbjct 620 LLQMEQTLHTRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTGVGK 672
> ECU11g1420
Length=851
Score = 45.4 bits (106), Expect = 3e-05, Method: Composition-based stats.
Identities = 17/53 (32%), Positives = 33/53 (62%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
++ ++ + R+ GQ AV A+ D++ R GL ++P+G+F+ LG + VG+
Sbjct 543 LMEMSSRIKKRIFGQDHAVDAIVDSILQSRVGLDDDDRPVGSFLLLGPTGVGK 595
> At2g25030
Length=265
Score = 43.9 bits (102), Expect = 7e-05, Method: Composition-based stats.
Identities = 24/61 (39%), Positives = 35/61 (57%), Gaps = 3/61 (4%)
Query 4 LADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR---GPSVGFE 60
L IL R+I Q V++VADA+ +AG+S N+ + +FMF+G V G S G+
Sbjct 2 LEQILHERIIAQDLDVESVADAIRCSKAGISDPNRLIASFMFMGQPSVVSQLVGASPGYV 61
Query 61 G 61
G
Sbjct 62 G 62
> YLL026w
Length=908
Score = 43.5 bits (101), Expect = 9e-05, Method: Composition-based stats.
Identities = 21/46 (45%), Positives = 33/46 (71%), Gaps = 1/46 (2%)
Query 8 LSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
LSS V+GQ A+KAV++A+ + R+GL+ +P +F+FLG S G+
Sbjct 576 LSSEVVGQMDAIKAVSNAVRLSRSGLANPRQP-ASFLFLGLSGSGK 620
> At5g51070
Length=945
Score = 41.6 bits (96), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 19/53 (35%), Positives = 32/53 (60%), Gaps = 0/53 (0%)
Query 1 VLRLADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
++ L D L RV+GQ +AV A++ A+ R GL ++P+ +F G + VG+
Sbjct 618 LMSLEDQLRGRVVGQDEAVAAISRAVKRSRVGLKDPDRPIAAMLFCGPTGVGK 670
> Hs13540606
Length=707
Score = 34.3 bits (77), Expect = 0.057, Method: Compositional matrix adjust.
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Query 4 LADILSSRVIGQQQAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGR 53
L L +IGQ+ A+ V A+ + G + PL F+FLGSS +G+
Sbjct 339 LEQRLKEHIIGQESAIATVGAAIRRKENGWYDEEHPL-VFLFLGSSGIGK 387
> Hs13027640
Length=926
Score = 30.8 bits (68), Expect = 0.64, Method: Composition-based stats.
Identities = 19/47 (40%), Positives = 25/47 (53%), Gaps = 0/47 (0%)
Query 17 QAVKAVADALAIQRAGLSPKNKPLGTFMFLGSSDVGRGPSVGFEGEP 63
QAV+AV DA GL PK+ TF+F G+ +V +G F P
Sbjct 193 QAVQAVRDAGYEISLGLMPKSIGPLTFVFTGTGNVSKGAQAIFNELP 239
Lambda K H
0.317 0.137 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1169706042
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40