bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_2119_orf2
Length=145
Score E
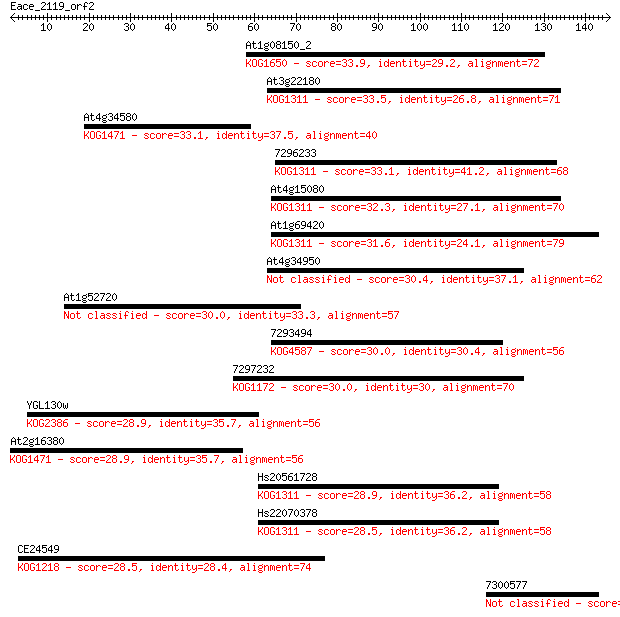
Sequences producing significant alignments: (Bits) Value
At1g08150_2 33.9 0.12
At3g22180 33.5 0.15
At4g34580 33.1 0.18
7296233 33.1 0.20
At4g15080 32.3 0.29
At1g69420 31.6 0.56
At4g34950 30.4 1.3
At1g52720 30.0 1.4
7293494 30.0 1.6
7297232 30.0 1.7
YGL130w 28.9 3.3
At2g16380 28.9 3.8
Hs20561728 28.9 4.0
Hs22070378 28.5 4.1
CE24549 28.5 4.8
7300577 28.1 5.7
> At1g08150_2
Length=2460
Score = 33.9 bits (76), Expect = 0.12, Method: Composition-based stats.
Identities = 21/74 (28%), Positives = 39/74 (52%), Gaps = 8/74 (10%)
Query 58 RPPGEFIRRHGFTRPLQLYQILSWVLFGVDILLFYLVVLPAVS--VPLKAIFGVLFGLTA 115
+ G+ IR G Q+Y+++S++L V+ F + + A + +PL+ F + L+
Sbjct 1162 KESGDLIRMEG-----QIYEVISFILL-VNTTKFVVTTITAYAFKMPLRDSFALALVLSN 1215
Query 116 ASLFGLAYYCTAAD 129
+F LAYY A +
Sbjct 1216 KGIFELAYYTYAVE 1229
> At3g22180
Length=706
Score = 33.5 bits (75), Expect = 0.15, Method: Compositional matrix adjust.
Identities = 19/72 (26%), Positives = 36/72 (50%), Gaps = 1/72 (1%)
Query 63 FIRRHGFTRPLQLYQILSWVLFGVDILLFYLVVLPAVSVPL-KAIFGVLFGLTAASLFGL 121
+R+HG+ P Q+++ +F + ++ FY P V + + + ++ A +F L
Sbjct 1 MVRKHGWQLPAHTLQVIAITVFCLLVVAFYAFFAPFVGGRIWEYVLIGVYSPVAILVFVL 60
Query 122 AYYCTAADPIDP 133
CTA +P DP
Sbjct 61 YVRCTAINPADP 72
> At4g34580
Length=560
Score = 33.1 bits (74), Expect = 0.18, Method: Composition-based stats.
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 4/44 (9%)
Query 19 GACTPEEK----RRPAGGWAEPLYLETAAHKQARCTTIDRTDSR 58
GACT E+K R G W +P L+ A +++A+C+ I + +
Sbjct 298 GACTCEDKGGCMRSDKGPWNDPEVLKIAINREAKCSPISEDEHK 341
> 7296233
Length=968
Score = 33.1 bits (74), Expect = 0.20, Method: Composition-based stats.
Identities = 28/74 (37%), Positives = 36/74 (48%), Gaps = 12/74 (16%)
Query 65 RRHGFTRPLQLYQILSW---VLFGVDILLFYLVVLPAVSVPLKAIFGVLFGL-TAASLFG 120
R HG PL QI W +LFGV Y V++PA ++ G L+GL T L
Sbjct 102 RLHGLQLPLHPLQIFGWLVLLLFGV---ASYWVLIPAFHARIQ---GPLYGLITGLYLVH 155
Query 121 LAYYCTA--ADPID 132
+A + TA DP D
Sbjct 156 IASHLTALLTDPAD 169
> At4g15080
Length=736
Score = 32.3 bits (72), Expect = 0.29, Method: Composition-based stats.
Identities = 19/71 (26%), Positives = 36/71 (50%), Gaps = 1/71 (1%)
Query 64 IRRHGFTRPLQLYQILSWVLFGVDILLFYLVVLPAVSVPL-KAIFGVLFGLTAASLFGLA 122
+R+HG+ P +Q+++ +F + + +Y P V + + I ++ A +F L
Sbjct 2 VRKHGWQLPAHKFQVVAITVFCLLSVAYYAFFAPFVGGRIWEYILLGVYSPVALIVFVLY 61
Query 123 YYCTAADPIDP 133
CTA +P DP
Sbjct 62 VRCTAINPADP 72
> At1g69420
Length=519
Score = 31.6 bits (70), Expect = 0.56, Method: Composition-based stats.
Identities = 19/80 (23%), Positives = 36/80 (45%), Gaps = 1/80 (1%)
Query 64 IRRHGFTRPLQLYQILSWVLFGVDILLFYLVVLPAVSVPLKAIFGV-LFGLTAASLFGLA 122
+R+HG+ P Q+++ +F FY+ P V + + ++ + GL
Sbjct 1 MRKHGWQLPYHPLQVVAVAVFLALGFAFYVFFAPFVGKKIHQYIAMGIYTPLITCVVGLY 60
Query 123 YYCTAADPIDPLAFCSGPFV 142
+C A+DP D F S ++
Sbjct 61 IWCAASDPADRGVFRSKKYL 80
> At4g34950
Length=567
Score = 30.4 bits (67), Expect = 1.3, Method: Composition-based stats.
Identities = 23/69 (33%), Positives = 33/69 (47%), Gaps = 10/69 (14%)
Query 63 FIRRHGFTRPLQLYQILSWVLFGVDILLFYLVVLPAVSVPLKAIFGVLFGL-------TA 115
FI++ G RPL + + ++ V LL L LP + GV +G+ TA
Sbjct 409 FIKKAGTPRPL--WNAAAQIIMAVGYLLMALA-LPGSLYIGSMVVGVCYGVRLAITVPTA 465
Query 116 ASLFGLAYY 124
+ LFGL YY
Sbjct 466 SELFGLKYY 474
> At1g52720
Length=117
Score = 30.0 bits (66), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 19/67 (28%), Positives = 32/67 (47%), Gaps = 14/67 (20%)
Query 14 AVGDSGACTPEEKRRPAGGW--------AEPLYLETAAHKQARCTTI--DRTDSRPPGEF 63
A SG+ P++ R+ + W ++P YL ++ C+T+ D+TD G+
Sbjct 15 ASSGSGSLNPDQNRKKSAAWWAPLFGLPSDPDYLNI----ESSCSTVNPDKTDISGSGQK 70
Query 64 IRRHGFT 70
RR FT
Sbjct 71 FRRGCFT 77
> 7293494
Length=872
Score = 30.0 bits (66), Expect = 1.6, Method: Composition-based stats.
Identities = 17/59 (28%), Positives = 32/59 (54%), Gaps = 7/59 (11%)
Query 64 IRRHGFT---RPLQLYQILSWVLFGVDILLFYLVVLPAVSVPLKAIFGVLFGLTAASLF 119
+ R GF +P+Q Y W++ GV +L+ YL ++ + IFG ++G+ + +F
Sbjct 108 VVRSGFNPMVQPMQFYLEFVWLMGGVTLLVLYLY----GTLLSENIFGGIYGVISYLMF 162
> 7297232
Length=1086
Score = 30.0 bits (66), Expect = 1.7, Method: Composition-based stats.
Identities = 21/70 (30%), Positives = 37/70 (52%), Gaps = 5/70 (7%)
Query 55 TDSRPPGEFIRRHGFTRPLQLYQILSWVLFGVDILLFYLVVLPAVSVPLKAIFGVLFGLT 114
++ PGE + G R ++ IL ++ GV +LL L+ ++P+ +FGV +
Sbjct 858 SECSAPGEKPQFLG-VREQRVTHILIFLTIGVSVLLTPLLG----NIPMPVLFGVFLYMG 912
Query 115 AASLFGLAYY 124
ASL GL ++
Sbjct 913 VASLKGLQFF 922
> YGL130w
Length=459
Score = 28.9 bits (63), Expect = 3.3, Method: Composition-based stats.
Identities = 20/61 (32%), Positives = 27/61 (44%), Gaps = 8/61 (13%)
Query 5 EAVEDSVGLA-----VGDSGACTPEEKRRPAGGWAEPLYLETAAHKQARCTTIDRTDSRP 59
E++ DSV L VGD C E + AGG PL + + A +T S+P
Sbjct 386 ESINDSVSLEDLEEIVGDIKRCWDERRANMAGGSGRPL---PSQSQNATLSTSKPVHSQP 442
Query 60 P 60
P
Sbjct 443 P 443
> At2g16380
Length=582
Score = 28.9 bits (63), Expect = 3.8, Method: Composition-based stats.
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 5/60 (8%)
Query 1 PQCNEAVEDSVGLAVGDSGACTPEEK----RRPAGGWAEPLYLETAAHKQARCTTIDRTD 56
P+ EA+ D+ L G CT +K R G W +P L+ A + +AR +TI D
Sbjct 279 PKLLEAI-DASELPYFFGGLCTCADKGGCLRSDKGPWNDPELLKIARNPEARFSTISEED 337
> Hs20561728
Length=485
Score = 28.9 bits (63), Expect = 4.0, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 35/68 (51%), Gaps = 10/68 (14%)
Query 61 GEFIRRHGFTRPLQLYQILSWVL--------FGVDILLFYLVVLPAVSVPLKAIFG--VL 110
G+ RR+G++ P QI++W+L FG+ + L +PA + AIF ++
Sbjct 36 GQRSRRNGWSWPPHPLQIVAWLLYLFFAVIGFGILVPLLPHHWVPAGYACMGAIFAGHLV 95
Query 111 FGLTAASL 118
LTA S+
Sbjct 96 VHLTAVSI 103
> Hs22070378
Length=485
Score = 28.5 bits (62), Expect = 4.1, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 35/68 (51%), Gaps = 10/68 (14%)
Query 61 GEFIRRHGFTRPLQLYQILSWVL--------FGVDILLFYLVVLPAVSVPLKAIFG--VL 110
G+ RR+G++ P QI++W+L FG+ + L +PA + AIF ++
Sbjct 36 GQRSRRNGWSWPPHPLQIVAWLLYLFFAVIGFGILVPLLPHHWVPAGYACMGAIFAGHLV 95
Query 111 FGLTAASL 118
LTA S+
Sbjct 96 VHLTAVSI 103
> CE24549
Length=1664
Score = 28.5 bits (62), Expect = 4.8, Method: Composition-based stats.
Identities = 21/82 (25%), Positives = 30/82 (36%), Gaps = 8/82 (9%)
Query 3 CNEAVEDSVGLAVGDSGACTPE--------EKRRPAGGWAEPLYLETAAHKQARCTTIDR 54
C E + G SG C E+ P+G + L+ + QARC +
Sbjct 1425 CKEKCRCANGHCNASSGECKCNLGFTGPSCEQSCPSGKYGLNCTLDCECYGQARCDPVQG 1484
Query 55 TDSRPPGEFIRRHGFTRPLQLY 76
PPG + R F+ P Y
Sbjct 1485 CCDCPPGRYGSRCQFSCPNGFY 1506
> 7300577
Length=863
Score = 28.1 bits (61), Expect = 5.7, Method: Composition-based stats.
Identities = 11/27 (40%), Positives = 19/27 (70%), Gaps = 0/27 (0%)
Query 116 ASLFGLAYYCTAADPIDPLAFCSGPFV 142
A+L+G+ Y TAA+ ++P+ GPF+
Sbjct 57 ATLYGIVVYRTAANLLEPVCSTYGPFL 83
Lambda K H
0.325 0.142 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1712413322
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40