bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
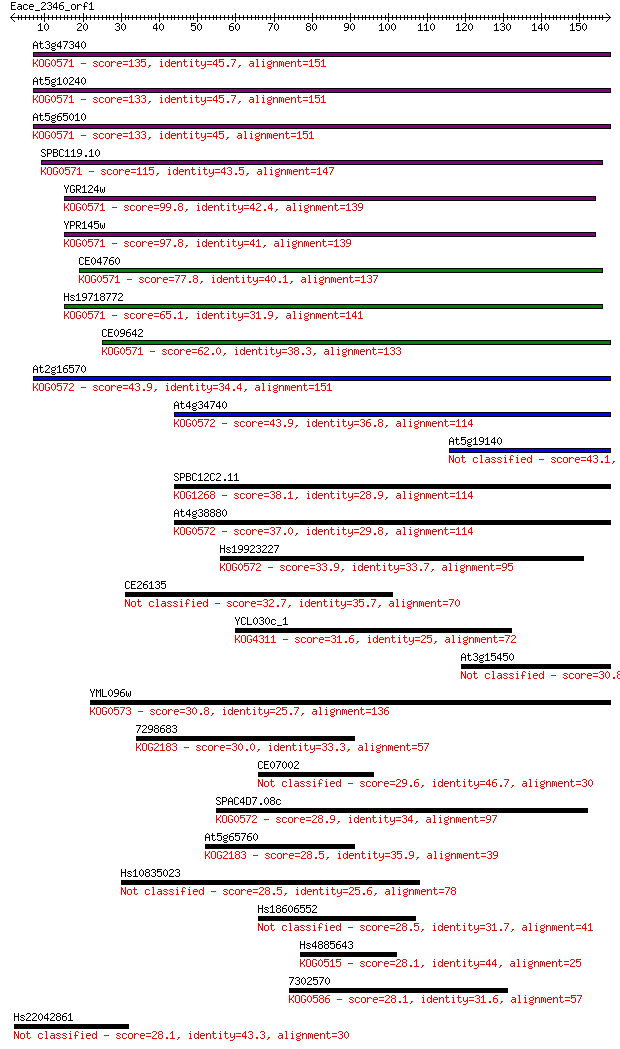
Query= Eace_2346_orf1
Length=157
Score E
Sequences producing significant alignments: (Bits) Value
At3g47340 135 3e-32
At5g10240 133 1e-31
At5g65010 133 1e-31
SPBC119.10 115 3e-26
YGR124w 99.8 2e-21
YPR145w 97.8 7e-21
CE04760 77.8 8e-15
Hs19718772 65.1 5e-11
CE09642 62.0 4e-10
At2g16570 43.9 1e-04
At4g34740 43.9 1e-04
At5g19140 43.1 2e-04
SPBC12C2.11 38.1 0.007
At4g38880 37.0 0.015
Hs19923227 33.9 0.13
CE26135 32.7 0.31
YCL030c_1 31.6 0.64
At3g15450 30.8 1.0
YML096w 30.8 1.2
7298683 30.0 1.8
CE07002 29.6 2.8
SPAC4D7.08c 28.9 4.3
At5g65760 28.5 5.4
Hs10835023 28.5 6.1
Hs18606552 28.5 6.3
Hs4885643 28.1 7.3
7302570 28.1 7.4
Hs22042861 28.1 7.7
> At3g47340
Length=584
Score = 135 bits (340), Expect = 3e-32, Method: Compositional matrix adjust.
Identities = 69/151 (45%), Positives = 96/151 (63%), Gaps = 8/151 (5%)
Query 7 NPSALRQLCLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPE 66
+ A R L S+ +RHRGPDW+G+ GD LAH+RL+++DP SG QPL + +
Sbjct 13 DSQAKRVRVLELSRRLRHRGPDWSGL---YQNGDNY--LAHQRLAVIDPASGDQPLFNED 67
Query 67 GRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATI 126
+V NGEIYNH +LR RL K +T SDC+VI L++ +G DFV MLDG F+ +
Sbjct 68 KTIVVTVNGEIYNHEELRKRLKNH---KFRTGSDCEVIAHLYEEYGVDFVDMLDGIFSFV 124
Query 127 VVDRHTGDFLVARDPIGISPLYVGHSADGSI 157
++D F+VARD IG++ LY+G DGS+
Sbjct 125 LLDTRDNSFMVARDAIGVTSLYIGWGLDGSV 155
> At5g10240
Length=578
Score = 133 bits (335), Expect = 1e-31, Method: Compositional matrix adjust.
Identities = 69/151 (45%), Positives = 93/151 (61%), Gaps = 8/151 (5%)
Query 7 NPSALRQLCLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPE 66
N A R + S+ +RHRGPDW+G+ LAHERL+IVDPTSG QPL + +
Sbjct 13 NSQAKRSRIIELSRRLRHRGPDWSGLHCYEDC-----YLAHERLAIVDPTSGDQPLYNED 67
Query 67 GRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATI 126
+ NGEIYNH LR L ++ + +T SDC+VI L++ G +FV MLDG FA +
Sbjct 68 KTIAVTVNGEIYNHKALRENL---KSHQFRTGSDCEVIAHLYEEHGEEFVDMLDGMFAFV 124
Query 127 VVDRHTGDFLVARDPIGISPLYVGHSADGSI 157
++D F+ ARD IGI+PLY+G DGS+
Sbjct 125 LLDTRDKSFIAARDAIGITPLYIGWGLDGSV 155
> At5g65010
Length=578
Score = 133 bits (334), Expect = 1e-31, Method: Compositional matrix adjust.
Identities = 68/151 (45%), Positives = 94/151 (62%), Gaps = 8/151 (5%)
Query 7 NPSALRQLCLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPE 66
N A R + S+ +RHRGPDW+G+ LAHERL+I+DPTSG QPL + +
Sbjct 13 NSQAKRSRIIELSRRLRHRGPDWSGLHCYEDC-----YLAHERLAIIDPTSGDQPLYNED 67
Query 67 GRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATI 126
V NGEIYNH LR +L ++ + +T SDC+VI L++ G +F+ MLDG FA +
Sbjct 68 KTVAVTVNGEIYNHKILREKL---KSHQFRTGSDCEVIAHLYEEHGEEFIDMLDGMFAFV 124
Query 127 VVDRHTGDFLVARDPIGISPLYVGHSADGSI 157
++D F+ ARD IGI+PLY+G DGS+
Sbjct 125 LLDTRDKSFIAARDAIGITPLYIGWGLDGSV 155
> SPBC119.10
Length=557
Score = 115 bits (288), Expect = 3e-26, Method: Compositional matrix adjust.
Identities = 64/147 (43%), Positives = 87/147 (59%), Gaps = 7/147 (4%)
Query 9 SALRQLCLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGR 68
A + L SK +RHRGPDW+G + + L HERL+IV SG QPL +G+
Sbjct 15 EAFKPKALHLSKQLRHRGPDWSGKAIRNQT-----ILCHERLAIVGVESGAQPLVSDDGK 69
Query 69 VVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATIVV 128
+V NGEIYNHL+LR L + K +T SDC+VI L++ G +MLDG F+ ++
Sbjct 70 LVLTVNGEIYNHLKLRENLKGN--YKFKTYSDCEVILYLYREHGPACANMLDGMFSWVLY 127
Query 129 DRHTGDFLVARDPIGISPLYVGHSADG 155
D+ + ARDPIGI+ LY G S+D
Sbjct 128 DQDKDKVVAARDPIGITTLYQGFSSDS 154
> YGR124w
Length=572
Score = 99.8 bits (247), Expect = 2e-21, Method: Compositional matrix adjust.
Identities = 59/139 (42%), Positives = 75/139 (53%), Gaps = 8/139 (5%)
Query 15 CLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGRVVCVAN 74
L SK IRHRGPDW+G V++ HERL+IV SG QP+ +G + N
Sbjct 20 ALQLSKKIRHRGPDWSGNAVMNST-----IFVHERLAIVGLDSGAQPITSADGEYMLGVN 74
Query 75 GEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATIVVDRHTGD 134
GEIYNH+QLR + K QT SDC+ I L+ D LDG FA + D
Sbjct 75 GEIYNHIQLREMCSD---YKFQTFSDCEPIIPLYLEHDIDAPKYLDGMFAFCLYDSKKDR 131
Query 135 FLVARDPIGISPLYVGHSA 153
+ ARDPIG+ LY+G S+
Sbjct 132 IVAARDPIGVVTLYMGRSS 150
> YPR145w
Length=572
Score = 97.8 bits (242), Expect = 7e-21, Method: Compositional matrix adjust.
Identities = 57/139 (41%), Positives = 74/139 (53%), Gaps = 8/139 (5%)
Query 15 CLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGRVVCVAN 74
L SK IRHRGPDW+G + + HERL+IV SG QP+ +G + N
Sbjct 20 ALQLSKRIRHRGPDWSGNAIKNST-----IFVHERLAIVGVESGAQPITSSDGEYMLCVN 74
Query 75 GEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATIVVDRHTGD 134
GEIYNH+QLR + E T SDC+ I ++ D LDG FA + D
Sbjct 75 GEIYNHIQLREECADYE---FGTLSDCEPIIPMYLKHDIDAPKYLDGMFAWTLYDAKQDR 131
Query 135 FLVARDPIGISPLYVGHSA 153
+ ARDPIGI+ LY+G S+
Sbjct 132 IVAARDPIGITTLYMGRSS 150
> CE04760
Length=567
Score = 77.8 bits (190), Expect = 8e-15, Method: Compositional matrix adjust.
Identities = 55/138 (39%), Positives = 74/138 (53%), Gaps = 9/138 (6%)
Query 19 SKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGRVVCVANGEIY 78
S+ HRGPD++G + GD L HERL+I+D QPL + NGEIY
Sbjct 27 SRRQHHRGPDFSGCYQDTKTGD---ILVHERLAIMD-LGVVQPLQGTAPNRQVIHNGEIY 82
Query 79 NHLQLRARLPEDEAIKLQTQSDCQVIPSLF-KYFGRDFVSMLDGDFATIVVDRHTGDFLV 137
NH +LR E + L+T D +VI L+ KY ++LDG FA ++ GDFL
Sbjct 83 NHQELRD--TELKKYNLKTHCDSEVIIFLYEKYRDGHICNLLDGVFAFVLC--CDGDFLA 138
Query 138 ARDPIGISPLYVGHSADG 155
ARDP+G+ +Y G DG
Sbjct 139 ARDPLGVKQMYYGIDDDG 156
> Hs19718772
Length=561
Score = 65.1 bits (157), Expect = 5e-11, Method: Composition-based stats.
Identities = 45/143 (31%), Positives = 71/143 (49%), Gaps = 9/143 (6%)
Query 15 CLARSKLIRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGRVVCVA- 73
CL+ K I HRGPD + ++ G RL++VDP G QP+ + + +
Sbjct 18 CLSAMK-IAHRGPDAFRFENVN--GYTNCCFGFHRLAVVDPLFGMQPIRVKKYPYLWLCY 74
Query 74 NGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFG-RDFVSMLDGDFATIVVDRHT 132
NGEIYNH +++ + QT+ D ++I L+ G + MLDG FA +++D
Sbjct 75 NGEIYNHKKMQQHF----EFEYQTKVDGEIILHLYDKGGIEQTICMLDGVFAFVLLDTAN 130
Query 133 GDFLVARDPIGISPLYVGHSADG 155
+ RD G+ PL+ + DG
Sbjct 131 KKVFLGRDTYGVRPLFKAMTEDG 153
> CE09642
Length=551
Score = 62.0 bits (149), Expect = 4e-10, Method: Compositional matrix adjust.
Identities = 51/139 (36%), Positives = 71/139 (51%), Gaps = 18/139 (12%)
Query 25 RGPDWNGIDVISYGGDRIHALAHERLSIVDP--TSGKQPLNDPEGRVVCVANGEIYNHLQ 82
RGPD + V+ +H L RL+IV P T QP+ VVC NGEIYNH
Sbjct 26 RGPD---LTVLEEVQPNVH-LGFHRLAIVMPGDTPSAQPIVGGGLSVVC--NGEIYNHQS 79
Query 83 LRARLPEDEAIKLQTQ-SDCQVIPSLFKYFGRDF---VSMLDGDFATIVVDRHTGDFLVA 138
L+ P L+ SDC I + F +D + LDG FA I+ D + +
Sbjct 80 LKIDCP----FTLKNGGSDCAAIITSFIKHNKDLKEACASLDGVFAFIMAD--DKNVYIG 133
Query 139 RDPIGISPLYVGHSADGSI 157
RDPIG+ PL+ G++++GS+
Sbjct 134 RDPIGVRPLFYGYNSNGSL 152
> At2g16570
Length=566
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 52/191 (27%), Positives = 80/191 (41%), Gaps = 49/191 (25%)
Query 7 NPSALRQLCLARSKLIRHRGPDWNGIDVIS-----------------YGGDRI------H 43
+P A R LC ++HRG + GI +S + ++
Sbjct 99 DPEASR-LCYLALHALQHRGQEGAGIVTVSPEKVLQTITGVGLVSEVFNESKLDQLPGEF 157
Query 44 ALAHERLSIVDPTSGKQPLNDPE--------GRVVCVANGEIYNHLQLRARLPEDEAIKL 95
A+ H R S T+G L + + G + NG + N+ LRA L E+ +I
Sbjct 158 AIGHVRYS----TAGASMLKNVQPFVAGYRFGSIGVAHNGNLVNYKTLRAMLEENGSI-F 212
Query 96 QTQSDCQVIPSLFK------YFGR--DFVSMLDGDFATIVVDRHTGDFLVA-RDPIGISP 146
T SD +V+ L +F R D L G ++ + V T D LVA RDP G P
Sbjct 213 NTSSDTEVVLHLIAISKARPFFMRIIDACEKLQGAYSMVFV---TEDKLVAVRDPYGFRP 269
Query 147 LYVGHSADGSI 157
L +G ++G++
Sbjct 270 LVMGRRSNGAV 280
> At4g34740
Length=561
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 42/131 (32%), Positives = 62/131 (47%), Gaps = 25/131 (19%)
Query 44 ALAHERLSIVDPTSGKQPLNDPE--------GRVVCVANGEIYNHLQLRARLPEDEAIKL 95
A+ H R S T+G L + + G V NG + N+ +LRA L E+ +I
Sbjct 154 AIGHVRYS----TAGSSMLKNVQPFVAGYRFGSVGVAHNGNLVNYTKLRADLEENGSI-F 208
Query 96 QTQSDCQVIPSLFK------YFGR--DFVSMLDGDFATIVVDRHTGDFLVA-RDPIGISP 146
T SD +V+ L +F R D L G ++ + V T D LVA RDP G P
Sbjct 209 NTSSDTEVVLHLIAISKARPFFMRIVDACEKLQGAYSMVFV---TEDKLVAVRDPHGFRP 265
Query 147 LYVGHSADGSI 157
L +G ++G++
Sbjct 266 LVMGRRSNGAV 276
> At5g19140
Length=234
Score = 43.1 bits (100), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 20/42 (47%), Positives = 26/42 (61%), Gaps = 0/42 (0%)
Query 116 VSMLDGDFATIVVDRHTGDFLVARDPIGISPLYVGHSADGSI 157
V+ L GDFA +V D+ T VA D +G PLY G +ADG +
Sbjct 125 VAHLSGDFAFVVFDKSTSTLFVASDQVGKVPLYWGITADGYV 166
> SPBC12C2.11
Length=696
Score = 38.1 bits (87), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 33/130 (25%), Positives = 60/130 (46%), Gaps = 20/130 (15%)
Query 44 ALAHERLS---IVDPTSGKQPLNDPEGRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSD 100
A++H R + I P + +DP + V NG + N+ +LR L E + ++++D
Sbjct 82 AISHTRWATHGIPSPINCHPQRSDPHSEFIVVHNGILTNYRELRTVL-ESRGMVFESETD 140
Query 101 CQVIPSLFKYF-----GRDFVSM-------LDGDFATIVVDRHT-GDFLVARDPIGISPL 147
+ + L K+ G DF S+ L+G +A ++ H G+ + R SPL
Sbjct 141 TECVAKLAKFIYDTTPGVDFTSLAKLVFRELEGAYALLIKSSHYPGEVVATRRG---SPL 197
Query 148 YVGHSADGSI 157
VG ++ +
Sbjct 198 IVGVKSEQKL 207
> At4g38880
Length=532
Score = 37.0 bits (84), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 34/130 (26%), Positives = 57/130 (43%), Gaps = 23/130 (17%)
Query 44 ALAHERLSIVDPTSGKQPLNDPE--------GRVVCVANGEIYNHLQLRARLPEDEAIKL 95
A+ H R S TSG L + + G + NG N+ QL+ +L E +I +
Sbjct 143 AIGHVRYS----TSGASMLKNVQPFIASCKLGSLAVAHNGNFVNYKQLKTKLEEMGSIFI 198
Query 96 QTQSDCQVIPSLF-KYFGRDFV-------SMLDGDFATIVVDRHTGDFLVARDPIGISPL 147
T SD +++ L K + F+ L G ++ + V + RDP G PL
Sbjct 199 -TSSDTELVLHLIAKSKAKTFLLRVIDACEKLRGAYSMVFV--FEDKLIAVRDPFGFRPL 255
Query 148 YVGHSADGSI 157
+G ++G++
Sbjct 256 VMGRRSNGAV 265
> Hs19923227
Length=517
Score = 33.9 bits (76), Expect = 0.13, Method: Compositional matrix adjust.
Identities = 32/115 (27%), Positives = 48/115 (41%), Gaps = 21/115 (18%)
Query 56 TSGKQPLNDPE--------GRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSL 107
T+GK L + + G++ NGE+ N +LR +L I L T SD ++I L
Sbjct 96 TTGKCELENCQPFVVETLHGKIAVAHNGELVNAARLRKKLLR-HGIGLSTSSDSEMITQL 154
Query 108 FKYF-------GRDFVS-----MLDGDFATIVVDRHTGDFLVARDPIGISPLYVG 150
Y D+V+ M + A ++ H RDP G PL +G
Sbjct 155 LAYTPPQEQDDTPDWVARIKNLMKEAPTAYSLLIMHRDVIYAVRDPYGNRPLCIG 209
> CE26135
Length=438
Score = 32.7 bits (73), Expect = 0.31, Method: Compositional matrix adjust.
Identities = 25/70 (35%), Positives = 33/70 (47%), Gaps = 6/70 (8%)
Query 31 GIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGRVVCVANGEIYNHLQLRARLPED 90
GID I GG A H+ SIV+P G P N G++ N ++ L L L ED
Sbjct 355 GIDEIDNGGGTAIACGHDLTSIVNPRCGNIPFN---GKITFFNNA--FHRLYLNITLDED 409
Query 91 EAIKLQTQSD 100
I +T+ D
Sbjct 410 IQI-FKTKLD 418
> YCL030c_1
Length=371
Score = 31.6 bits (70), Expect = 0.64, Method: Compositional matrix adjust.
Identities = 18/72 (25%), Positives = 34/72 (47%), Gaps = 3/72 (4%)
Query 60 QPLNDPEGRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSML 119
+ LN P+ RVV NG N ++ + +D+ + ++ S + + G
Sbjct 92 EQLNVPKERVVVEENGVFSNQFMVKQKFSQDKIVSIKKLSKDMLTKEV---LGEVRTDRP 148
Query 120 DGDFATIVVDRH 131
DG + T+VVD++
Sbjct 149 DGLYTTLVVDQY 160
> At3g15450
Length=253
Score = 30.8 bits (68), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 16/39 (41%), Positives = 19/39 (48%), Gaps = 0/39 (0%)
Query 119 LDGDFATIVVDRHTGDFLVARDPIGISPLYVGHSADGSI 157
L+G FA +V D T A G LY G S DGS+
Sbjct 130 LEGSFAFVVYDTQTSSVFSALSSDGGESLYWGISGDGSV 168
> YML096w
Length=525
Score = 30.8 bits (68), Expect = 1.2, Method: Composition-based stats.
Identities = 35/141 (24%), Positives = 59/141 (41%), Gaps = 22/141 (15%)
Query 22 IRHRGPDWNGIDVISYGGDRIHALAHERLSIVDPTSGKQPLNDPEGRVVCVANGEIYNHL 81
I RGP+++ + + RI + LS+ P + KQ +N + R NGE+YN
Sbjct 47 IAARGPNYSSLRAVK--AYRISWFS-SVLSLRQPFT-KQSIN-VDDRYFLQFNGELYN-- 99
Query 82 QLRARLPEDEAIKLQTQSDCQVIPSLFKYFGR-----DFVSMLDGDFATIVVDRHTGDFL 136
+ I Q +D I S+ + D + L+G++A + D ++
Sbjct 100 ---------KEIS-QGDNDSLYIASMLQNLKEGMGVIDVIKSLEGEYAYTIYDVNSSKLY 149
Query 137 VARDPIGISPLYVGHSADGSI 157
RDPIG L + D +
Sbjct 150 FGRDPIGRRSLSYSVTPDNEL 170
> 7298683
Length=475
Score = 30.0 bits (66), Expect = 1.8, Method: Composition-based stats.
Identities = 19/59 (32%), Positives = 28/59 (47%), Gaps = 2/59 (3%)
Query 34 VISYGGDRIHALAHERLS--IVDPTSGKQPLNDPEGRVVCVANGEIYNHLQLRARLPED 90
++ YGG + A + S ++DP SG L P +V + E +HL LR P D
Sbjct 395 ILRYGGRNLEAATNIIFSNGLLDPWSGGGVLQAPNDKVFVIILPEGAHHLDLRHSDPAD 453
> CE07002
Length=131
Score = 29.6 bits (65), Expect = 2.8, Method: Compositional matrix adjust.
Identities = 14/30 (46%), Positives = 18/30 (60%), Gaps = 0/30 (0%)
Query 66 EGRVVCVANGEIYNHLQLRARLPEDEAIKL 95
EGR V + NG +Y QL AR+P I+L
Sbjct 102 EGRTVTMVNGALYFDYQLVARIPPHYDIRL 131
> SPAC4D7.08c
Length=533
Score = 28.9 bits (63), Expect = 4.3, Method: Compositional matrix adjust.
Identities = 33/122 (27%), Positives = 51/122 (41%), Gaps = 27/122 (22%)
Query 55 PTSGK------QPL--NDPEGRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPS 106
PT+G QP N P G +V NG + N +LR L + + T SD +++ +
Sbjct 79 PTAGSCAHSEAQPFYVNSPYG-LVLGHNGNLINGPELRRFLDTEAHRHVNTGSDSELLLN 137
Query 107 LFKY-----------------FGRDFVSMLDGDFATIVVDRHTGDFLVARDPIGISPLYV 149
+F Y R+ ++G +A + + G L RDP GI PL +
Sbjct 138 IFAYELQRLDKFRINENDIFEALRNVYDRVNGGYACVAMIAGLG-VLGFRDPNGIRPLVI 196
Query 150 GH 151
G
Sbjct 197 GE 198
> At5g65760
Length=529
Score = 28.5 bits (62), Expect = 5.4, Method: Compositional matrix adjust.
Identities = 14/39 (35%), Positives = 20/39 (51%), Gaps = 0/39 (0%)
Query 52 IVDPTSGKQPLNDPEGRVVCVANGEIYNHLQLRARLPED 90
++DP SG L + +V + E +HL LR PED
Sbjct 454 LLDPWSGGSVLKNLSDTIVALVTKEGAHHLDLRPSTPED 492
> Hs10835023
Length=2695
Score = 28.5 bits (62), Expect = 6.1, Method: Composition-based stats.
Identities = 20/84 (23%), Positives = 38/84 (45%), Gaps = 8/84 (9%)
Query 30 NGIDVISYGGDR------IHALAHERLSIVDPTSGKQPLNDPEGRVVCVANGEIYNHLQL 83
+G DV+ + DR I + ER + + + ++ E VC +Y ++
Sbjct 1323 SGEDVLVFYNDRASFQTLIQMMRSERDRMDENSPLMYHIHLVELLAVCTEGKNVYTEIKC 1382
Query 84 RARLPEDEAIKLQTQSDCQVIPSL 107
+ LP D+ +++ T DC IP +
Sbjct 1383 NSLLPLDDIVRVVTHEDC--IPEV 1404
> Hs18606552
Length=636
Score = 28.5 bits (62), Expect = 6.3, Method: Composition-based stats.
Identities = 13/41 (31%), Positives = 20/41 (48%), Gaps = 0/41 (0%)
Query 66 EGRVVCVANGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPS 106
+ R ++G+ HLQ R P KL+T + Q +PS
Sbjct 100 QARAPVPSHGQAMKHLQGNVRTPAGSMTKLETHNTSQPLPS 140
> Hs4885643
Length=1005
Score = 28.1 bits (61), Expect = 7.3, Method: Composition-based stats.
Identities = 11/25 (44%), Positives = 17/25 (68%), Gaps = 0/25 (0%)
Query 77 IYNHLQLRARLPEDEAIKLQTQSDC 101
+Y LQLR +L +++ KLQ Q +C
Sbjct 143 LYKELQLRNKLNQEQNAKLQQQREC 167
> 7302570
Length=702
Score = 28.1 bits (61), Expect = 7.4, Method: Compositional matrix adjust.
Identities = 18/57 (31%), Positives = 28/57 (49%), Gaps = 5/57 (8%)
Query 74 NGEIYNHLQLRARLPEDEAIKLQTQSDCQVIPSLFKYFGRDFVSMLDGDFATIVVDR 130
NGEI++HL R+ E EA ++ TQ + S Y R V D +++D+
Sbjct 122 NGEIFDHLVANGRMKEPEAARVFTQ-----LVSAVHYCHRRGVVHRDLKAENVLLDK 173
> Hs22042861
Length=811
Score = 28.1 bits (61), Expect = 7.7, Method: Composition-based stats.
Identities = 13/30 (43%), Positives = 16/30 (53%), Gaps = 0/30 (0%)
Query 2 TGPPLNPSALRQLCLARSKLIRHRGPDWNG 31
T PL+ A QLC K + HRGP+ G
Sbjct 770 TRSPLHSHATLQLCHGDGKTVTHRGPNVGG 799
Lambda K H
0.321 0.142 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2106641548
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40