bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_2445_orf1
Length=110
Score E
Sequences producing significant alignments: (Bits) Value
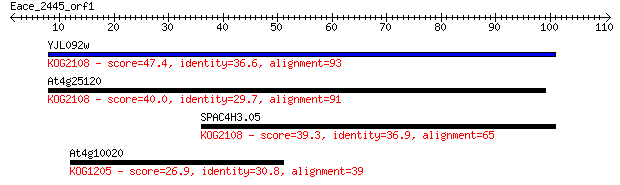
YJL092w 47.4 6e-06
At4g25120 40.0 0.001
SPAC4H3.05 39.3 0.002
At4g10020 26.9 8.6
> YJL092w
Length=1174
Score = 47.4 bits (111), Expect = 6e-06, Method: Composition-based stats.
Identities = 34/93 (36%), Positives = 45/93 (48%), Gaps = 23/93 (24%)
Query 8 LDFTDLTVKALKLLEPPATRAELGAQWPYLVVDEFQDTNSNQFKFICLLALQRPPNIQTL 67
LDF DL + +LL TR + + +++VDEFQDTN Q + L A
Sbjct 217 LDFDDLLMYTFRLL----TRVRVLSNIKHVLVDEFQDTNGIQLDLMFLFA---------- 262
Query 68 HTAVGNPTLPNSTPGRGGVTVVGDDDQSIYKWR 100
GN L G+T+VGD DQSIY +R
Sbjct 263 ---KGNHHLSR------GMTIVGDPDQSIYAFR 286
> At4g25120
Length=938
Score = 40.0 bits (92), Expect = 0.001, Method: Composition-based stats.
Identities = 27/92 (29%), Positives = 39/92 (42%), Gaps = 25/92 (27%)
Query 8 LDFTDLTVKALKLLEP-PATRAELGAQWPYLVVDEFQDTNSNQFKFICLLALQRPPNIQT 66
LD+ DL ++ LL P E W ++VDEFQDT++ Q+K + +L
Sbjct 417 LDYHDLISCSVTLLSDFPEVFKECQDTWKAIIVDEFQDTSTMQYKLLRMLG--------- 467
Query 67 LHTAVGNPTLPNSTPGRGGVTVVGDDDQSIYK 98
+T+VGDDDQ K
Sbjct 468 ---------------SHNHITIVGDDDQITVK 484
> SPAC4H3.05
Length=887
Score = 39.3 bits (90), Expect = 0.002, Method: Composition-based stats.
Identities = 24/65 (36%), Positives = 34/65 (52%), Gaps = 23/65 (35%)
Query 36 YLVVDEFQDTNSNQFKFICLLALQRPPNIQTLHTAVGNPTLPNSTPGRGGVTVVGDDDQS 95
++++DEFQDT+ Q+ + LLALQ NS +T+VGD DQS
Sbjct 234 HILIDEFQDTSKIQYFLVKLLALQ------------------NSD-----ITIVGDPDQS 270
Query 96 IYKWR 100
IY +R
Sbjct 271 IYGFR 275
> At4g10020
Length=389
Score = 26.9 bits (58), Expect = 8.6, Method: Composition-based stats.
Identities = 12/39 (30%), Positives = 17/39 (43%), Gaps = 0/39 (0%)
Query 12 DLTVKALKLLEPPATRAELGAQWPYLVVDEFQDTNSNQF 50
DL + L+ PPAT + WP L F + N +
Sbjct 3 DLLNSVMNLVAPPATMVVMAFAWPLLSFISFSERAYNSY 41
Lambda K H
0.319 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1195973986
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40