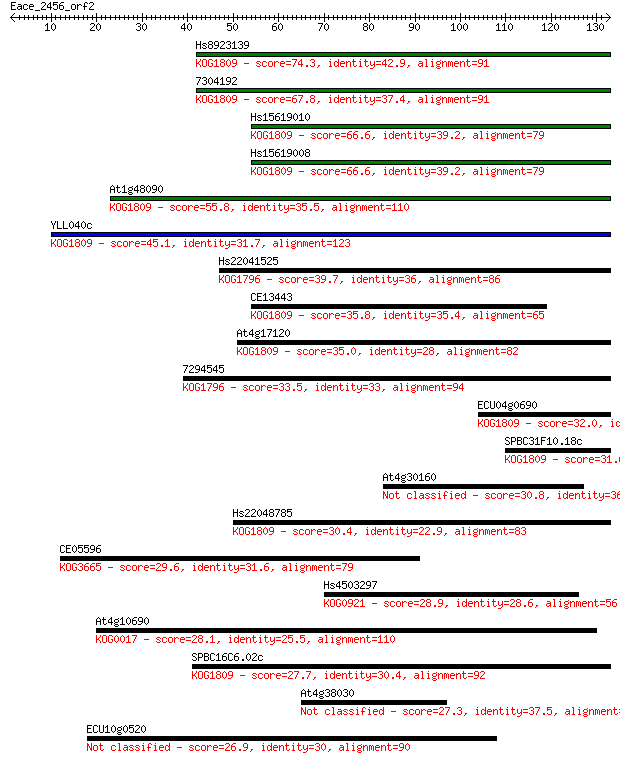
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_2456_orf2
Length=132
Score E
Sequences producing significant alignments: (Bits) Value
Hs8923139 74.3 6e-14
7304192 67.8 5e-12
Hs15619010 66.6 1e-11
Hs15619008 66.6 1e-11
At1g48090 55.8 2e-08
YLL040c 45.1 3e-05
Hs22041525 39.7 0.002
CE13443 35.8 0.021
At4g17120 35.0 0.041
7294545 33.5 0.12
ECU04g0690 32.0 0.36
SPBC31F10.18c 31.6 0.36
At4g30160 30.8 0.65
Hs22048785 30.4 0.96
CE05596 29.6 1.8
Hs4503297 28.9 2.5
At4g10690 28.1 4.7
SPBC16C6.02c 27.7 5.4
At4g38030 27.3 7.6
ECU10g0520 26.9 9.4
> Hs8923139
Length=442
Score = 74.3 bits (181), Expect = 6e-14, Method: Compositional matrix adjust.
Identities = 39/92 (42%), Positives = 53/92 (57%), Gaps = 1/92 (1%)
Query 42 NTIGLAANAVDSVSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIGEGV 101
+T+G AA V ++ +G L+ T D EY +R+ E R D GK GV
Sbjct 158 HTVGGAAGVVSRITGSVGKGLAAITMDKEYQQKRREELSRQPRDFGDSLARGGKGFLRGV 217
Query 102 WS-LTNIVTKPIEGAQREGVGGFFKGIGKGIV 132
+T I+TKP+EGA++EG GFFKGIGKG+V
Sbjct 218 VGGVTGIITKPVEGAKKEGAAGFFKGIGKGLV 249
> 7304192
Length=3242
Score = 67.8 bits (164), Expect = 5e-12, Method: Composition-based stats.
Identities = 34/92 (36%), Positives = 54/92 (58%), Gaps = 1/92 (1%)
Query 42 NTIGLAANAVDSVSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIGEG- 100
+T+G AA AV ++ +G L+ TFD +Y +R++ + +G + K + G
Sbjct 2964 HTVGGAAGAVSKITGAMGKGLAALTFDEDYQKKRRQGIQNKPKNFHEGLARSSKGLVMGF 3023
Query 101 VWSLTNIVTKPIEGAQREGVGGFFKGIGKGIV 132
V +T +VTKP+ GA+ GV GFFKG+GKG +
Sbjct 3024 VDGVTGVVTKPVTGARDNGVEGFFKGLGKGAI 3055
> Hs15619010
Length=3174
Score = 66.6 bits (161), Expect = 1e-11, Method: Composition-based stats.
Identities = 31/80 (38%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Query 54 VSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIGEG-VWSLTNIVTKPI 112
++ + ++ T D +Y +R+ + A R+G GK + G V +T IVTKPI
Sbjct 2924 ITGAMAKGVAAMTMDEDYQQKRREAMNKQPAGFREGITRGGKGLVSGFVSGITGIVTKPI 2983
Query 113 EGAQREGVGGFFKGIGKGIV 132
+GAQ+ G GFFKG+GKG+V
Sbjct 2984 KGAQKGGAAGFFKGVGKGLV 3003
> Hs15619008
Length=3095
Score = 66.6 bits (161), Expect = 1e-11, Method: Composition-based stats.
Identities = 31/80 (38%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Query 54 VSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIGEG-VWSLTNIVTKPI 112
++ + ++ T D +Y +R+ + A R+G GK + G V +T IVTKPI
Sbjct 2924 ITGAMAKGVAAMTMDEDYQQKRREAMNKQPAGFREGITRGGKGLVSGFVSGITGIVTKPI 2983
Query 113 EGAQREGVGGFFKGIGKGIV 132
+GAQ+ G GFFKG+GKG+V
Sbjct 2984 KGAQKGGAAGFFKGVGKGLV 3003
> At1g48090
Length=4099
Score = 55.8 bits (133), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 39/122 (31%), Positives = 66/122 (54%), Gaps = 16/122 (13%)
Query 23 SLGQSSLLN---------LPRMPFEL--GKNTIGLAANAVDSVSAGLGSLLSTFTFDSEY 71
S+ QS+++N L P +L G + +G A++A+ +S G+ +L + D ++
Sbjct 3742 SMRQSTMINNAIRNVKKDLLGQPLQLLSGVDILGNASSALGHMSQGIAAL----SMDKKF 3797
Query 72 INRRQRERVRNTASMRDGFLSAGKNIGEGVWS-LTNIVTKPIEGAQREGVGGFFKGIGKG 130
I RQR+ + D G + +G++ +T I+TKP+EGA+ GV GF G GKG
Sbjct 3798 IQSRQRQENKGVEDFGDIIREGGGALAKGLFRGVTGILTKPLEGAKSSGVEGFVSGFGKG 3857
Query 131 IV 132
I+
Sbjct 3858 II 3859
> YLL040c
Length=3144
Score = 45.1 bits (105), Expect = 3e-05, Method: Composition-based stats.
Identities = 39/129 (30%), Positives = 63/129 (48%), Gaps = 17/129 (13%)
Query 10 ERYTYHLLNSVFSSLGQSSLLNLPRMPFELGKNTI-GLAANAVDSVSAGLGSLLS--TFT 66
E Y +++N +G ++L + K T+ GL+ DS+S GS+ + T
Sbjct 2861 EPYQGYMMNDRPQEIG----IHLAKGGLSFAKKTVFGLS----DSMSKFTGSMAKGLSVT 2912
Query 67 FDSEYIN-RRQRERVR--NTASMRDGFLSAGKNIGEGVWSLTNIVTKPIEGAQREGVGGF 123
D E+ RR ++R+ N ++ + S +G G L+ I P + Q+EG GF
Sbjct 2913 QDLEFQRVRRLQQRINKNNRNALANSAQSFASTLGSG---LSGIALDPYKAMQKEGAAGF 2969
Query 124 FKGIGKGIV 132
KG+GKGIV
Sbjct 2970 LKGLGKGIV 2978
> Hs22041525
Length=1231
Score = 39.7 bits (91), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 31/89 (34%), Positives = 49/89 (55%), Gaps = 10/89 (11%)
Query 47 AANAVDSVSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKN-IGEGVWS-L 104
AA ++S GLG T D+ + + +RE +R A+ L AG + + G+ L
Sbjct 952 AAKFAGTLSDGLGK-----TMDNRH--QSEREYIRYHAATSGEHLVAGIHGLAHGIIGGL 1004
Query 105 TNIVTKPIEGAQREG-VGGFFKGIGKGIV 132
T+++T +EG + EG V GF G+GKG+V
Sbjct 1005 TSVITSTVEGVKTEGGVSGFISGLGKGLV 1033
> CE13443
Length=3212
Score = 35.8 bits (81), Expect = 0.021, Method: Composition-based stats.
Identities = 23/66 (34%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Query 54 VSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIG-EGVWSLTNIVTKPI 112
++ +G ++ TFD +Y+ +RQ + R S +G K +G V +T +VTKPI
Sbjct 2963 ITGTVGKGVAALTFDDDYMKKRQEDLNRKPQSFGEGMARGLKGLGMGVVGGITGVVTKPI 3022
Query 113 EGAQRE 118
EGA++E
Sbjct 3023 EGAKQE 3028
> At4g17120
Length=1661
Score = 35.0 bits (79), Expect = 0.041, Method: Compositional matrix adjust.
Identities = 23/83 (27%), Positives = 43/83 (51%), Gaps = 8/83 (9%)
Query 51 VDSVSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIGEGV-WSLTNIVT 109
V+S + GL SLL ++ +RR + D + + + +GV + ++ +VT
Sbjct 1513 VNSCNYGLNSLLKRGVNMNQVWSRR-------ITGVGDAIVQGTEALAQGVAFGVSGVVT 1565
Query 110 KPIEGAQREGVGGFFKGIGKGIV 132
KP+E A+ G+ GF G+G+ +
Sbjct 1566 KPVESARENGILGFAHGVGRAFL 1588
> 7294545
Length=1902
Score = 33.5 bits (75), Expect = 0.12, Method: Composition-based stats.
Identities = 31/98 (31%), Positives = 43/98 (43%), Gaps = 7/98 (7%)
Query 39 LGKNTIGLAANAVDSVSAGLGSLLSTFTFDSEYINRRQR--ERVRNTASMRDGFLSAGKN 96
L KN +N+ ++ L L D RQR E NT+ G L+AG
Sbjct 1722 LVKNVTHGISNSTAKLTETLSDSLGKVVLDDHDNETRQRILELQSNTSG---GHLAAGLK 1778
Query 97 IGEGVWS--LTNIVTKPIEGAQREGVGGFFKGIGKGIV 132
+T+IV +GA +GV GF G+GKG+V
Sbjct 1779 GLGFGLLGGVTSIVRHTYDGATSDGVPGFLSGLGKGLV 1816
> ECU04g0690
Length=2371
Score = 32.0 bits (71), Expect = 0.36, Method: Compositional matrix adjust.
Identities = 16/29 (55%), Positives = 20/29 (68%), Gaps = 1/29 (3%)
Query 104 LTNIVTKPIEGAQREGVGGFFKGIGKGIV 132
+ I T PIEGA +GV G KG+GKGI+
Sbjct 2200 IAGIATSPIEGAS-QGVTGVVKGLGKGIL 2227
> SPBC31F10.18c
Length=600
Score = 31.6 bits (70), Expect = 0.36, Method: Compositional matrix adjust.
Identities = 13/23 (56%), Positives = 18/23 (78%), Gaps = 0/23 (0%)
Query 110 KPIEGAQREGVGGFFKGIGKGIV 132
+PI GA+R G+ G KG+GKG+V
Sbjct 408 QPIIGARRNGLPGLVKGLGKGLV 430
> At4g30160
Length=974
Score = 30.8 bits (68), Expect = 0.65, Method: Composition-based stats.
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 6/47 (12%)
Query 83 TASMRD---GFLSAGKNIGEGVWSLTNIVTKPIEGAQREGVGGFFKG 126
+ SMRD F AG+ G +W + N + PI + +G FF G
Sbjct 2 SVSMRDLDPAFQGAGQKAGIEIWRIENFIPTPIP---KSSIGKFFTG 45
> Hs22048785
Length=1687
Score = 30.4 bits (67), Expect = 0.96, Method: Composition-based stats.
Identities = 19/93 (20%), Positives = 42/93 (45%), Gaps = 10/93 (10%)
Query 50 AVDSVSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKNIGEGVW-SLTNIV 108
++ +++ L + + D E+ NR++ R + S+ +G +G + ++ IV
Sbjct 1376 SITNLATSLARNMDRLSLDEEHYNRQEEWRRQLPESLGEGLRQGLSRLGISLLGAIAGIV 1435
Query 109 TKPIEGAQREG---------VGGFFKGIGKGIV 132
+P++ Q+ G G+GKGI+
Sbjct 1436 DQPMQNFQKTSEAQASAGHKAKGVISGVGKGIM 1468
> CE05596
Length=667
Score = 29.6 bits (65), Expect = 1.8, Method: Composition-based stats.
Identities = 25/83 (30%), Positives = 39/83 (46%), Gaps = 11/83 (13%)
Query 12 YTYHLLNSVFSSLGQSSLLN----LPRMPFELGKNTIGLAANAVDSVSAGLGSLLSTFTF 67
Y H LNS S SL+N LP+M E G+ +A N +DS+++ ++F
Sbjct 329 YYIHQLNSCEVSTLVISLVNSVMFLPKMDTEHGREINIMAWNTIDSLAS------YVYSF 382
Query 68 DSEYINRRQRERVRNTASMRDGF 90
D + I + + T + DGF
Sbjct 383 DCDNICKTAM-KFLTTTGVYDGF 404
> Hs4503297
Length=1279
Score = 28.9 bits (63), Expect = 2.5, Method: Composition-based stats.
Identities = 16/56 (28%), Positives = 31/56 (55%), Gaps = 3/56 (5%)
Query 70 EYINRRQRERVRNTASMRDGFLSAGKNIGEGVWSLTNIVTKPIEGAQREGVGGFFK 125
+Y +R++ + V+ T + L+AG + G W+L N + I+ Q+E + G +K
Sbjct 147 DYYSRKEEQEVQATLESEEVDLNAGLH---GNWTLENAKARLIQYFQKEKIQGEYK 199
> At4g10690
Length=1515
Score = 28.1 bits (61), Expect = 4.7, Method: Composition-based stats.
Identities = 28/110 (25%), Positives = 46/110 (41%), Gaps = 19/110 (17%)
Query 20 VFSSLGQSSLLNLPRMPFELGKNTIGLAANAVDSVSAGLGSLLSTFTFDSEYINRRQRER 79
+F SL + +L K IGL N+ V GL + F+ +Y +++
Sbjct 83 IFGSLSEEAL-----------KVVIGL--NSAQEVWLGLARRFNRFSTTRKYDLQKRLGT 129
Query 80 VRNTASMRDGFLSAGKNIGEGVWSLTNIVTKPIEGAQREGVGGFFKGIGK 129
D +LS KNI + + S+ VT ++E + G G+GK
Sbjct 130 CSKAGKTMDAYLSEVKNICDQLDSIGFPVT------EQEKIFGVLNGLGK 173
> SPBC16C6.02c
Length=3131
Score = 27.7 bits (60), Expect = 5.4, Method: Composition-based stats.
Identities = 28/93 (30%), Positives = 45/93 (48%), Gaps = 1/93 (1%)
Query 41 KNTIGLAANAVDSVSAGLGSLLSTFTFDSEYINRRQRERVRNTASMRDGFLSAGKN-IGE 99
+ TI +++V ++ + LST T D +Y N R+R R RN ++AG N +
Sbjct 2744 RKTIYGVSDSVSKITGTISKGLSTMTMDPKYQNSRRRFRSRNRPKEAVYGVTAGANSFYD 2803
Query 100 GVWSLTNIVTKPIEGAQREGVGGFFKGIGKGIV 132
+ S + KP + G F KG GKG++
Sbjct 2804 SMSSGFKGLKKPFTDPKNNSAGKFLKGFGKGML 2836
> At4g38030
Length=649
Score = 27.3 bits (59), Expect = 7.6, Method: Composition-based stats.
Identities = 12/32 (37%), Positives = 19/32 (59%), Gaps = 0/32 (0%)
Query 65 FTFDSEYINRRQRERVRNTASMRDGFLSAGKN 96
F S+Y++RR+R V + D FL+ GK+
Sbjct 340 FVASSDYLSRRERGSVTGRLLVNDRFLTPGKS 371
> ECU10g0520
Length=543
Score = 26.9 bits (58), Expect = 9.4, Method: Compositional matrix adjust.
Identities = 27/98 (27%), Positives = 47/98 (47%), Gaps = 18/98 (18%)
Query 18 NSVFSSLGQSSLLNLPRMPFELGKNTIGLAANAVDSVSAGLGSLLSTFTFDSEYINRRQR 77
N +F SL ++ RMP + +NT+GL NA + LG+ +S + I+ R
Sbjct 412 NVLFGSLFKN------RMPTFILENTLGLVTNASMKIGKYLGADVSKEAISMQ-IDPLYR 464
Query 78 ERVRNTASMRDGFL-SAGKNIGE-------GVWSLTNI 107
+ + ++ DG GK++G G+W +T+I
Sbjct 465 AKYK---AVYDGLCGKLGKSLGSIICTVMTGLWDITDI 499
Lambda K H
0.317 0.135 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1319765976
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40