bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_3076_orf1
Length=111
Score E
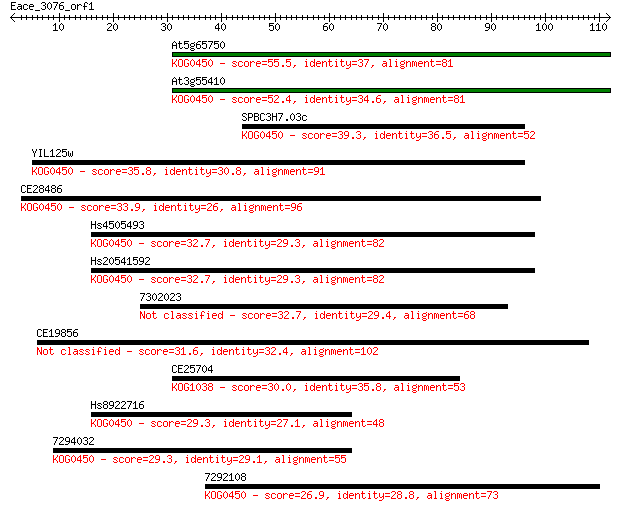
Sequences producing significant alignments: (Bits) Value
At5g65750 55.5 3e-08
At3g55410 52.4 2e-07
SPBC3H7.03c 39.3 0.002
YIL125w 35.8 0.018
CE28486 33.9 0.082
Hs4505493 32.7 0.16
Hs20541592 32.7 0.17
7302023 32.7 0.17
CE19856 31.6 0.35
CE25704 30.0 1.1
Hs8922716 29.3 1.8
7294032 29.3 2.1
7292108 26.9 8.7
> At5g65750
Length=1025
Score = 55.5 bits (132), Expect = 3e-08, Method: Composition-based stats.
Identities = 30/81 (37%), Positives = 45/81 (55%), Gaps = 9/81 (11%)
Query 31 SSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLPRKPPYCSLRAAQQPLQLGPEHYGFTA 90
S Q++ ++ RL+ +VR YQ+ GH A ++PL L ++ + P L P YGFT
Sbjct 114 SGQTIQESMRLLLLVRAYQVNGHMKAKLDPLGLEKR---------EIPEDLTPGLYGFTE 164
Query 91 ADFDKVYVARVPGMQGFLSPD 111
AD D+ + V M GFLS +
Sbjct 165 ADLDREFFLGVWRMSGFLSEN 185
> At3g55410
Length=1009
Score = 52.4 bits (124), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 28/81 (34%), Positives = 44/81 (54%), Gaps = 9/81 (11%)
Query 31 SSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLPRKPPYCSLRAAQQPLQLGPEHYGFTA 90
S Q++ ++ RL+ +VR YQ+ GH A ++PL L ++ + P L YGFT
Sbjct 111 SGQTIQESMRLLLLVRAYQVNGHMKAKLDPLGLEQR---------EIPEDLDLALYGFTE 161
Query 91 ADFDKVYVARVPGMQGFLSPD 111
AD D+ + V M GF+S +
Sbjct 162 ADLDREFFLGVWQMSGFMSEN 182
> SPBC3H7.03c
Length=1009
Score = 39.3 bits (90), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 19/52 (36%), Positives = 28/52 (53%), Gaps = 8/52 (15%)
Query 44 MVRGYQMIGHELAAINPLSLPRKPPYCSLRAAQQPLQLGPEHYGFTAADFDK 95
+VR YQ GH LA ++PL + +P +L EHYGFT +D ++
Sbjct 131 LVRAYQSRGHHLAKLDPLGINVN--------HNRPSELTLEHYGFTESDLNR 174
> YIL125w
Length=1014
Score = 35.8 bits (81), Expect = 0.018, Method: Composition-based stats.
Identities = 28/91 (30%), Positives = 42/91 (46%), Gaps = 14/91 (15%)
Query 5 PAATTAGTTGAAAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLP 64
P T A G A GS + + S+H +L + R YQ+ GH A I+PL +
Sbjct 99 PQGTEAAPLGTAMTGSVD--------ENVSIHLKVQL--LCRAYQVRGHLKAHIDPLGI- 147
Query 65 RKPPYCSLRAAQQPLQLGPEHYGFTAADFDK 95
+ S + P +L ++YGF+ D DK
Sbjct 148 ---SFGSNKNNPVPPELTLDYYGFSKHDLDK 175
> CE28486
Length=1029
Score = 33.9 bits (76), Expect = 0.082, Method: Compositional matrix adjust.
Identities = 25/96 (26%), Positives = 41/96 (42%), Gaps = 11/96 (11%)
Query 3 ASPAATTAGTTGAAAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLS 62
SPAA T+ A A S QS+ D ++ ++R YQ GH +A ++PL
Sbjct 109 VSPAAAQVTTSSAPATRLDTN------ASVQSISDHLKIQLLIRSYQTRGHNIADLDPLG 162
Query 63 LPRKPPYCSLRAAQQPLQLGPEHYGFTAADFDKVYV 98
+ ++ P +L YG D D+ ++
Sbjct 163 INSADLDDTI-----PPELELSFYGLGERDLDREFL 193
> Hs4505493
Length=1002
Score = 32.7 bits (73), Expect = 0.16, Method: Compositional matrix adjust.
Identities = 24/84 (28%), Positives = 38/84 (45%), Gaps = 4/84 (4%)
Query 16 AAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLPRKPPYCSLRA- 74
AA + L A + V D + ++R YQ+ GH +A ++PL + S+ A
Sbjct 105 AAVAHAQSLVEAQPNVDKLVEDHLAVQSLIRAYQIRGHHVAQLDPLGILDADLDSSVPAD 164
Query 75 -AQQPLQLGPEHYGFTAADFDKVY 97
+LG YG +D DKV+
Sbjct 165 IISSTDKLG--FYGLDESDLDKVF 186
> Hs20541592
Length=1023
Score = 32.7 bits (73), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 24/84 (28%), Positives = 38/84 (45%), Gaps = 4/84 (4%)
Query 16 AAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLPRKPPYCSLRA- 74
AA + L A + V D + ++R YQ+ GH +A ++PL + S+ A
Sbjct 105 AAVAHAQSLVEAQPNVDKLVEDHLAVQSLIRAYQIRGHHVAQLDPLGILDADLDSSVPAD 164
Query 75 -AQQPLQLGPEHYGFTAADFDKVY 97
+LG YG +D DKV+
Sbjct 165 IISSTDKLG--FYGLDESDLDKVF 186
> 7302023
Length=901
Score = 32.7 bits (73), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 20/69 (28%), Positives = 33/69 (47%), Gaps = 10/69 (14%)
Query 25 YSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLPRKPPYCSLRAAQQPLQ-LGP 83
+ E S + + + + V ++ GH+LAA+NP+ S+R +QQ LQ L P
Sbjct 40 FQVAEDVRASRNSQANVYRFVEAFRQHGHKLAAVNPI---------SIRTSQQELQELSP 90
Query 84 EHYGFTAAD 92
YG +
Sbjct 91 AFYGLQTQE 99
> CE19856
Length=933
Score = 31.6 bits (70), Expect = 0.35, Method: Composition-based stats.
Identities = 33/116 (28%), Positives = 45/116 (38%), Gaps = 20/116 (17%)
Query 6 AATTAGTTGAAAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLP- 64
A G G+ G +S ++GS S SR V G+Q G + I +P
Sbjct 523 GARVPGIQGSRFPGIQGARFSGFQGSRDSEFQGSR----VPGFQ--GARVPGIQGFRVPG 576
Query 65 ----RKPPYCSLRA--AQQPLQLGPEHYG-------FTAADFDKVYVARVPGMQGF 107
R P + R ++ P GP G F +V V+RVPG QGF
Sbjct 577 FQGSRVPGFQGFRVQVSRVPGFQGPGFQGSRVSGSRFPGFQGFRVQVSRVPGFQGF 632
> CE25704
Length=1154
Score = 30.0 bits (66), Expect = 1.1, Method: Composition-based stats.
Identities = 19/53 (35%), Positives = 25/53 (47%), Gaps = 2/53 (3%)
Query 31 SSQSVHDTSRLIQMVRGYQMIGHELAAINPLSLPRKPPYCSLRAAQQPLQLGP 83
SS ++ D RLI +G + + I PL LP PYC L + L L P
Sbjct 997 SSMALKDWFRLI--AKGSSDLMKTVEWITPLGLPVVQPYCKLVERKGKLILAP 1047
> Hs8922716
Length=1010
Score = 29.3 bits (64), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 13/48 (27%), Positives = 25/48 (52%), Gaps = 0/48 (0%)
Query 16 AAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSL 63
+ G S+ +S+ V D + ++R YQ+ GH +A ++PL +
Sbjct 92 SVVHEGRSAVSSRTKTSKLVEDHLAVQSLIRAYQIRGHHVAQLDPLGI 139
> 7294032
Length=1001
Score = 29.3 bits (64), Expect = 2.1, Method: Compositional matrix adjust.
Identities = 16/55 (29%), Positives = 28/55 (50%), Gaps = 8/55 (14%)
Query 9 TAGTTGAAAAGSGEGLYSAYEGSSQSVHDTSRLIQMVRGYQMIGHELAAINPLSL 63
TA G A AG+ S+++ D + ++R YQ+ GH +A ++PL +
Sbjct 106 TAFNFGGAVAGAAP--------DSKTIDDHLAVQAIIRSYQIRGHNIAHLDPLEI 152
> 7292108
Length=1075
Score = 26.9 bits (58), Expect = 8.7, Method: Composition-based stats.
Identities = 21/80 (26%), Positives = 36/80 (45%), Gaps = 7/80 (8%)
Query 37 DTSRLIQ-MVRGYQMIGHELAAINPLSL--PRKPPYCSLRAAQQPLQLGPEHYGFTAADF 93
D +IQ ++R YQ GH A ++PL + P+K ++ +H+ + D
Sbjct 141 DDHHVIQAIIRAYQSRGHLAADLDPLGIVGPKKRTSVDGTQRHAAREVLRQHFSYIFNDL 200
Query 94 DKVY----VARVPGMQGFLS 109
+ V+ + G Q FLS
Sbjct 201 NTVFKLPSSTMIGGDQEFLS 220
Lambda K H
0.314 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1192473466
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40