bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
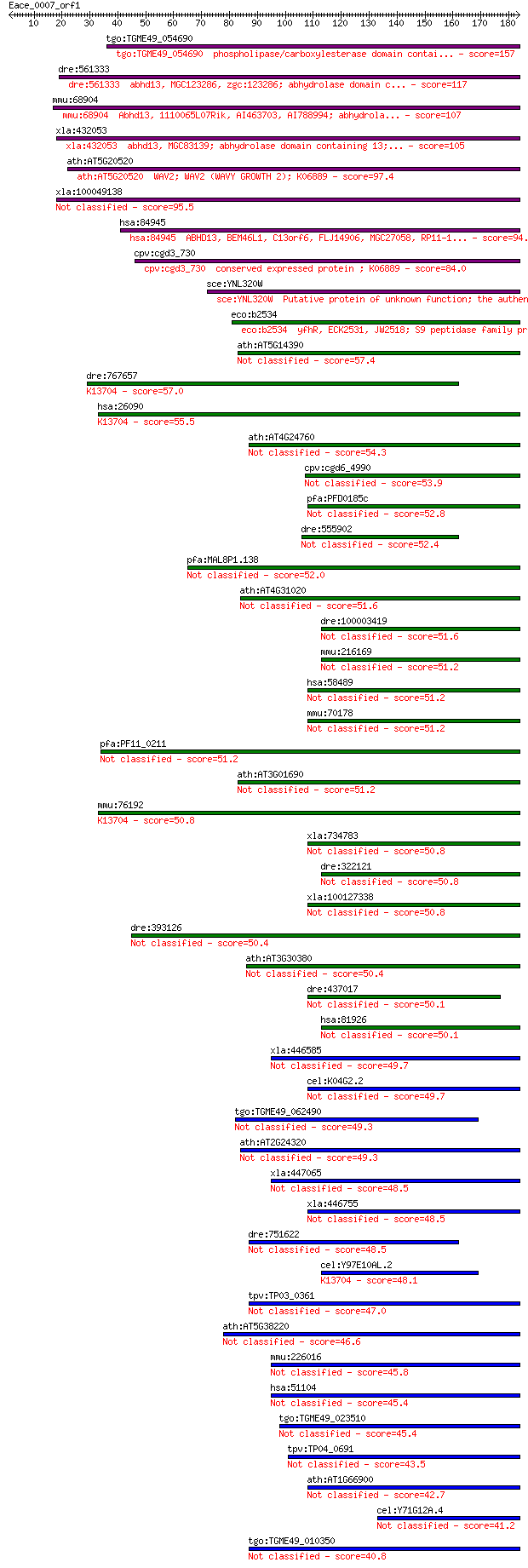
Query= Eace_0007_orf1
Length=183
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_054690 phospholipase/carboxylesterase domain contai... 157 2e-38
dre:561333 abhd13, MGC123286, zgc:123286; abhydrolase domain c... 117 2e-26
mmu:68904 Abhd13, 1110065L07Rik, AI463703, AI788994; abhydrola... 107 3e-23
xla:432053 abhd13, MGC83139; abhydrolase domain containing 13;... 105 8e-23
ath:AT5G20520 WAV2; WAV2 (WAVY GROWTH 2); K06889 97.4 2e-20
xla:100049138 hypothetical protein LOC100049138 95.5 9e-20
hsa:84945 ABHD13, BEM46L1, C13orf6, FLJ14906, MGC27058, RP11-1... 94.7 2e-19
cpv:cgd3_730 conserved expressed protein ; K06889 84.0 2e-16
sce:YNL320W Putative protein of unknown function; the authenti... 84.0 3e-16
eco:b2534 yfhR, ECK2531, JW2518; S9 peptidase family protein, ... 72.4 8e-13
ath:AT5G14390 hypothetical protein 57.4 2e-08
dre:767657 abhd12, MGC153367, zgc:153367; abhydrolase domain c... 57.0 4e-08
hsa:26090 ABHD12, ABHD12A, BEM46L2, C20orf22, DKFZp434P106, PH... 55.5 1e-07
ath:AT4G24760 hypothetical protein 54.3 2e-07
cpv:cgd6_4990 peptidase of the alpha/beta-hydrolase fold 53.9 3e-07
pfa:PFD0185c conserved Plasmodium protein, unknown function 52.8 6e-07
dre:555902 Bem46-like 52.4 8e-07
pfa:MAL8P1.138 alpha/beta hydrolase, putative 52.0 1e-06
ath:AT4G31020 hypothetical protein 51.6 2e-06
dre:100003419 si:rp71-61h23.3 51.6 2e-06
mmu:216169 Fam108a, 1700013O15Rik, BC005632, D10Bwg1364e, MGC1... 51.2 2e-06
hsa:58489 FAM108C1, FLJ34461, MGC131546; family with sequence ... 51.2 2e-06
mmu:70178 Fam108c, 2210412D01Rik, AL023007, Fam108c1; family w... 51.2 2e-06
pfa:PF11_0211 conserved Plasmodium protein 51.2 2e-06
ath:AT3G01690 hypothetical protein 51.2 2e-06
mmu:76192 Abhd12, 1500011G07Rik, 6330583M11Rik, AI431047, AW54... 50.8 2e-06
xla:734783 fam108a1, MGC131027, fam108a2; family with sequence... 50.8 3e-06
dre:322121 fb50g01, wu:fb50g01; zgc:162293 50.8 3e-06
xla:100127338 hypothetical protein LOC100127338 50.8 3e-06
dre:393126 MGC55468, fam108c1; zgc:55468 50.4 3e-06
ath:AT3G30380 hypothetical protein 50.4 3e-06
dre:437017 zgc:100937 50.1 4e-06
hsa:81926 FAM108A1, C19orf27, MGC5244; family with sequence si... 50.1 5e-06
xla:446585 fam108b1, MGC81688; family with sequence similarity... 49.7 6e-06
cel:K04G2.2 hypothetical protein 49.7 6e-06
tgo:TGME49_062490 hypothetical protein 49.3 7e-06
ath:AT2G24320 hypothetical protein 49.3 7e-06
xla:447065 fam108b1, MGC83647; abhydrolase domain-containing p... 48.5 1e-05
xla:446755 fam108c1, MGC79044; family with sequence similarity... 48.5 1e-05
dre:751622 MGC153037, zgc:153037; si:ch211-117n7.7 48.5 1e-05
cel:Y97E10AL.2 hypothetical protein; K13704 abhydrolase domain... 48.1 2e-05
tpv:TP03_0361 hypothetical protein 47.0 3e-05
ath:AT5G38220 hydrolase 46.6 5e-05
mmu:226016 Fam108b, 5730446C15Rik, Cgi67, Fam108b1, MGC40949; ... 45.8 7e-05
hsa:51104 FAM108B1, C9orf77, RP11-409O11.2; family with sequen... 45.4 1e-04
tgo:TGME49_023510 hypothetical protein 45.4 1e-04
tpv:TP04_0691 hypothetical protein 43.5 4e-04
ath:AT1G66900 hypothetical protein 42.7 7e-04
cel:Y71G12A.4 hypothetical protein 41.2 0.002
tgo:TGME49_010350 hypothetical protein 40.8 0.002
> tgo:TGME49_054690 phospholipase/carboxylesterase domain containing
protein (EC:3.1.-.-); K06889
Length=497
Score = 157 bits (397), Expect = 2e-38, Method: Compositional matrix adjust.
Identities = 79/154 (51%), Positives = 105/154 (68%), Gaps = 6/154 (3%)
Query 36 VVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGIPFKDVSFTTRDGVKLSA 95
++ +V LLW FQEKLLF P G++TP+ NP+G RSPAERG+PF+++ T DGVKL
Sbjct 37 LLCMVVLLWYFQEKLLFYPGVPQGFETPDKNPKGLRSPAERGLPFEELWLRTVDGVKLHC 96
Query 96 WLLLGEKP---LEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTP 152
WL+ + P APT I FHGNAGN+GF + E+L +G NVL++ YRGYG SEG+P
Sbjct 97 WLIKQKLPQVAAHAPTLIFFHGNAGNVGFRLPNVELLYKHVGVNVLIVSYRGYGFSEGSP 156
Query 153 SESGVYADADAALDFLLSTS---LVNNKDIFLFG 183
+E+GVY D +AALD L+ ++ IFLFG
Sbjct 157 TEAGVYRDGEAALDMLVERQNELHIDANKIFLFG 190
> dre:561333 abhd13, MGC123286, zgc:123286; abhydrolase domain
containing 13; K06889
Length=337
Score = 117 bits (293), Expect = 2e-26, Method: Compositional matrix adjust.
Identities = 69/175 (39%), Positives = 99/175 (56%), Gaps = 18/175 (10%)
Query 19 FGGVSILSATLLVAFIGVVVLVGL--------LWRFQEKLLFLNAFPVGYKTPNLNPRGY 70
F VS+L+ L G VL+GL L++FQ+ LL+ P +
Sbjct 25 FCRVSLLALILTFHLYGGFVLLGLILASLAGILYKFQDVLLYFPDQPSSSRL-------- 76
Query 71 RSPAERGIPFKDVSFTTRDGVKLSAWLL--LGEKPLEAPTFIQFHGNAGNIGFHQGHAEV 128
P GIP ++V T+DG++L+ LL GE P APT + FHGNAGNIG +A +
Sbjct 77 YVPMPTGIPHENVYIRTKDGIRLNLILLRYTGENPAGAPTILYFHGNAGNIGHRVPNALL 136
Query 129 LCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
+ + ANV+L+DYRGYG SEG PSE G+Y DA+A LD++++ ++ + LFG
Sbjct 137 MLVNLKANVVLVDYRGYGKSEGDPSEDGLYQDAEATLDYVMTRPDIDKTKVVLFG 191
> mmu:68904 Abhd13, 1110065L07Rik, AI463703, AI788994; abhydrolase
domain containing 13; K06889
Length=337
Score = 107 bits (266), Expect = 3e-23, Method: Compositional matrix adjust.
Identities = 64/169 (37%), Positives = 103/169 (60%), Gaps = 17/169 (10%)
Query 17 RLFGGVSILSATLLVAFIGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAER 76
L+GG+ +L L+ F+ + G+L++FQ+ LL+ FP + P+ + R Y P
Sbjct 38 HLYGGIVLL----LLIFVSIA---GILYKFQDVLLY---FP---EQPS-SSRLY-VPMPT 82
Query 77 GIPFKDVSFTTRDGVKLSAWLL--LGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMG 134
GIP +++ T+DGV+L+ L+ G+ PT I FHGNAGNIG +A ++ +
Sbjct 83 GIPHENIFIRTKDGVRLNLILVRYTGDNSPYCPTIIYFHGNAGNIGHRLPNALLMLVNLR 142
Query 135 ANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
N++L+DYRGYG SEG SE G+Y D++A LD++++ ++ +FLFG
Sbjct 143 VNLVLVDYRGYGKSEGEASEEGLYLDSEAVLDYVMTRPDLDKTKVFLFG 191
> xla:432053 abhd13, MGC83139; abhydrolase domain containing 13;
K06889
Length=336
Score = 105 bits (262), Expect = 8e-23, Method: Compositional matrix adjust.
Identities = 61/168 (36%), Positives = 99/168 (58%), Gaps = 17/168 (10%)
Query 18 LFGGVSILSATLLVAFIGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERG 77
++GGV +L L+ F+ + G+L++FQ+ LL+ FP + L P G
Sbjct 39 MYGGVILL----LLIFVSIA---GILFKFQDVLLY---FPDQPSSSRL-----YIPMPTG 83
Query 78 IPFKDVSFTTRDGVKLSAWLL--LGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGA 135
IP +++ T+D ++L+ LL G+ +PT I FHGNAGNIG +A ++ +
Sbjct 84 IPHENIFIKTKDNIRLNLILLRYTGDNSSFSPTIIYFHGNAGNIGHRLPNALLMLVNLKV 143
Query 136 NVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
N++L+DYRGYG S+G PSE G+Y D++A LD++++ ++ I LFG
Sbjct 144 NLILVDYRGYGKSDGEPSEEGLYMDSEAVLDYVMTRPDIDKTKIILFG 191
> ath:AT5G20520 WAV2; WAV2 (WAVY GROWTH 2); K06889
Length=308
Score = 97.4 bits (241), Expect = 2e-20, Method: Compositional matrix adjust.
Identities = 61/163 (37%), Positives = 88/163 (53%), Gaps = 8/163 (4%)
Query 22 VSILSATLLVAFIGVVVL-VGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGIPF 80
V+ +SA L F G+VV V LL FQEKL+++ PV P L+ +PA + +
Sbjct 2 VTYVSA-LFYGFGGIVVAGVALLVAFQEKLVYV---PV---LPGLSKSYPITPARLNLIY 54
Query 81 KDVSFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLL 140
+D+ + DGV+L AW + PT + F NAGNI ++ ++ NV +L
Sbjct 55 EDIWLQSSDGVRLHAWFIKMFPECRGPTILFFQENAGNIAHRLEMVRIMIQKLKCNVFML 114
Query 141 DYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
YRGYG SEG PS+ G+ DA AALD L + ++ I +FG
Sbjct 115 SYRGYGASEGYPSQQGIIKDAQAALDHLSGRTDIDTSRIVVFG 157
> xla:100049138 hypothetical protein LOC100049138
Length=336
Score = 95.5 bits (236), Expect = 9e-20, Method: Compositional matrix adjust.
Identities = 61/168 (36%), Positives = 99/168 (58%), Gaps = 17/168 (10%)
Query 18 LFGGVSILSATLLVAFIGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERG 77
++GGV +L L+ F+ + G+L++FQ+ LL+ FP + L P G
Sbjct 39 MYGGVILL----LLIFVSIA---GILFKFQDVLLY---FPDQPSSSRL-----YIPMPTG 83
Query 78 IPFKDVSFTTRDGVKLSAWLL--LGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGA 135
IP +++ T+D ++L+ LL G+ +PT + FHGNAGNIG +A ++ +
Sbjct 84 IPHENIFIKTKDNIRLNLILLRYTGDNSNFSPTIVYFHGNAGNIGHRLPNALLMLVNLKV 143
Query 136 NVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
N+LL+DYRGYG S+G PSE G+Y D++A LD++++ ++ I LFG
Sbjct 144 NLLLVDYRGYGKSDGEPSEEGLYLDSEAVLDYIMTRPDIDKTKIILFG 191
> hsa:84945 ABHD13, BEM46L1, C13orf6, FLJ14906, MGC27058, RP11-153I24.2,
bA153I24.2; abhydrolase domain containing 13; K06889
Length=337
Score = 94.7 bits (234), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 57/145 (39%), Positives = 88/145 (60%), Gaps = 10/145 (6%)
Query 41 GLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGIPFKDVSFTTRDGVKLSAWLL-- 98
G+L++FQ+ LL+ P + R Y P GIP +++ T+DG++L+ L+
Sbjct 55 GILYKFQDVLLYFPEQPS-------SSRLY-VPMPTGIPHENIFIRTKDGIRLNLILIRY 106
Query 99 LGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVY 158
G+ +PT I FHGNAGNIG +A ++ + N+LL+DYRGYG SEG SE G+Y
Sbjct 107 TGDNSPYSPTIIYFHGNAGNIGHRLPNALLMLVNLKVNLLLVDYRGYGKSEGEASEEGLY 166
Query 159 ADADAALDFLLSTSLVNNKDIFLFG 183
D++A LD++++ ++ IFLFG
Sbjct 167 LDSEAVLDYVMTRPDLDKTKIFLFG 191
> cpv:cgd3_730 conserved expressed protein ; K06889
Length=419
Score = 84.0 bits (206), Expect = 2e-16, Method: Compositional matrix adjust.
Identities = 64/207 (30%), Positives = 86/207 (41%), Gaps = 78/207 (37%)
Query 46 FQEKLLFLNAFPVGYKTPNLNPR--------GYRSPAERGIPFKDVSFTTRDGVKLSAWL 97
FQEKLLF PN+N + GYR P E + +DV TT D VKL W
Sbjct 45 FQEKLLFF---------PNINGKRDLASNFPGYRHPNENKMNSEDVVLTTDDNVKLYCWF 95
Query 98 LLGEK---------------------------------------PLEAPTFIQF------ 112
++ +K +AP +++F
Sbjct 96 IIHKKFDRYSHGQSSAMHFSSNLTEPHHSEQVKKDGPEKKILDRIRKAPKYMKFHFPEFS 155
Query 113 ---------------HGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGV 157
HGNAGNIG +G N+ + YRGYG+SEGTPSE G
Sbjct 156 DRYEQQEKAPTIVFFHGNAGNIGHRLPRFLEFYNLIGVNIFAVSYRGYGDSEGTPSEEGF 215
Query 158 YADADAALDFLLS-TSLVNNKDIFLFG 183
Y DA A+L+++LS T +V+ IFL+G
Sbjct 216 YLDAKASLEYVLSRTDVVDKNMIFLYG 242
> sce:YNL320W Putative protein of unknown function; the authentic,
non-tagged protein is detected in highly purified mitochondria
in high-throughput studies; K06889
Length=284
Score = 84.0 bits (206), Expect = 3e-16, Method: Compositional matrix adjust.
Identities = 41/112 (36%), Positives = 64/112 (57%), Gaps = 2/112 (1%)
Query 72 SPAERGIPFKDVSFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCA 131
+P RGIP++ ++ T+D +KL AW + E T + NAGNIG+ ++
Sbjct 46 TPDSRGIPYEKLTLITQDHIKLEAWDIKNEN--STSTVLILCPNAGNIGYFILIIDIFYR 103
Query 132 EMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
+ G +V + YRGYGNSEG+PSE G+ DAD + L + S + + + L+G
Sbjct 104 QFGMSVFIYSYRGYGNSEGSPSEKGLKLDADCVISHLSTDSFHSKRKLVLYG 155
> eco:b2534 yfhR, ECK2531, JW2518; S9 peptidase family protein,
function unknown; K06889
Length=284
Score = 72.4 bits (176), Expect = 8e-13, Method: Compositional matrix adjust.
Identities = 40/106 (37%), Positives = 57/106 (53%), Gaps = 4/106 (3%)
Query 81 KDVSFTTRDGVKLSAWLL---LGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANV 137
+ V FT +DG +L W + G T I HGNAGN+ H L E NV
Sbjct 50 ESVEFTAKDGTRLQGWFIPSSTGPADNAIATIIHAHGNAGNMSAHWPLVSWL-PERNFNV 108
Query 138 LLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
+ DYRG+G S+GTPS++G+ D +A++ + S VN + + LFG
Sbjct 109 FMFDYRGFGKSKGTPSQAGLLDDTQSAINVVRHRSDVNPQRLVLFG 154
> ath:AT5G14390 hypothetical protein
Length=369
Score = 57.4 bits (137), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 34/101 (33%), Positives = 48/101 (47%), Gaps = 2/101 (1%)
Query 83 VSFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDY 142
+ TR G ++ A + P+ T + HGNA ++G L + N++ DY
Sbjct 47 LKLPTRRGTEIVAMYV--RHPMATSTLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDY 104
Query 143 RGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
GYG S G PSE YAD +AA L T +DI L+G
Sbjct 105 SGYGQSTGKPSEHHTYADIEAAYKCLEETYGAKQEDIILYG 145
> dre:767657 abhd12, MGC153367, zgc:153367; abhydrolase domain
containing 12 (EC:3.1.1.23); K13704 abhydrolase domain-containing
protein 12 [EC:3.1.1.23]
Length=382
Score = 57.0 bits (136), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 42/152 (27%), Positives = 71/152 (46%), Gaps = 25/152 (16%)
Query 29 LLVAFIGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGIPF-KDVSFTT 87
LL +I + V++ + Q KL+FLN V Y + P ++G+ +
Sbjct 65 LLGIYIAIPVIIKVCPSIQAKLVFLNFVRVPYFI------DLKRPQDQGMNHTHNFYLQP 118
Query 88 RDGVKLSAWLLLG---------------EKPLEA--PTFIQFHGNAGNIG-FHQGHAEVL 129
+G+ + W + EK ++ P + HGNAG G H+ +
Sbjct 119 EEGINIGVWHTVPAGMWREAQAKDAEWYEKSFQSSHPVILYLHGNAGTRGGDHRVQLYKV 178
Query 130 CAEMGANVLLLDYRGYGNSEGTPSESGVYADA 161
+ +G +V+ DYRG+G+SEG+PSE G+ +DA
Sbjct 179 LSSLGYHVVTFDYRGWGDSEGSPSERGMTSDA 210
> hsa:26090 ABHD12, ABHD12A, BEM46L2, C20orf22, DKFZp434P106,
PHARC, dJ965G21.2; abhydrolase domain containing 12 (EC:3.1.1.23);
K13704 abhydrolase domain-containing protein 12 [EC:3.1.1.23]
Length=398
Score = 55.5 bits (132), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 47/170 (27%), Positives = 77/170 (45%), Gaps = 27/170 (15%)
Query 33 FIGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGI---------PFKDV 83
+I + L+ L Q KL+FLN V Y + P ++G+ P +DV
Sbjct 83 YIAIPFLIKLCPGIQAKLIFLNFVRVPYFI------DLKKPQDQGLNHTCNYYLQPEEDV 136
Query 84 SFTTRDGVKLSAW-------LLLGEKPLEA--PTFIQFHGNAGNIG-FHQGHAEVLCAEM 133
+ V W + E L + P + HGNAG G H+ + + +
Sbjct 137 TIGVWHTVPAVWWKNAQGKDQMWYEDALASSHPIILYLHGNAGTRGGDHRVELYKVLSSL 196
Query 134 GANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
G +V+ DYRG+G+S GTPSE G+ DA D++ + S + ++++G
Sbjct 197 GYHVVTFDYRGWGDSVGTPSERGMTYDALHVFDWIKARS--GDNPVYIWG 244
> ath:AT4G24760 hypothetical protein
Length=365
Score = 54.3 bits (129), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 32/97 (32%), Positives = 46/97 (47%), Gaps = 2/97 (2%)
Query 87 TRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYG 146
TR G ++ A + P+ T + HGNA +IG L + N++ DY GYG
Sbjct 51 TRRGTEIVAMYI--RYPMAVTTLLYSHGNAADIGQMYELFIELSIHLRVNLMGYDYSGYG 108
Query 147 NSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
S G P+E YAD +AA L ++I L+G
Sbjct 109 QSSGKPTEQNTYADIEAAYKCLEENYGAKQENIILYG 145
> cpv:cgd6_4990 peptidase of the alpha/beta-hydrolase fold
Length=383
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 31/77 (40%), Positives = 40/77 (51%), Gaps = 0/77 (0%)
Query 107 PTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALD 166
P FI HGNA +IG L ++ A+VL DYR YG S+G P+E G+YAD A +
Sbjct 156 PVFIFSHGNATDIGSMLPWFVNLSLKLNAHVLAYDYRSYGLSKGKPTERGIYADIKAVYE 215
Query 167 FLLSTSLVNNKDIFLFG 183
+ IFL G
Sbjct 216 YARDELNFPTDRIFLLG 232
> pfa:PFD0185c conserved Plasmodium protein, unknown function
Length=734
Score = 52.8 bits (125), Expect = 6e-07, Method: Composition-based stats.
Identities = 27/76 (35%), Positives = 42/76 (55%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA +IG E +G N+ DY GYG S G P+E+ +Y D +AA ++
Sbjct 47 TILFSHGNAEDIGDIVPQFESKLKRLGLNMFAYDYSGYGQSTGYPTETHLYNDVEAAYNY 106
Query 168 LLSTSLVNNKDIFLFG 183
L+S ++ + I +G
Sbjct 107 LISELNISKECIIAYG 122
> dre:555902 Bem46-like
Length=344
Score = 52.4 bits (124), Expect = 8e-07, Method: Compositional matrix adjust.
Identities = 29/58 (50%), Positives = 39/58 (67%), Gaps = 3/58 (5%)
Query 106 APTFIQFHGNAGN-IGFHQ-GHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADA 161
+P FI HGN GN H+ G A VL A +G +VL++DYRG+G+S G P+E G+ DA
Sbjct 117 SPIFIYLHGNGGNRSALHRIGVANVLSA-LGYHVLVMDYRGFGDSTGEPTEPGLTTDA 173
> pfa:MAL8P1.138 alpha/beta hydrolase, putative
Length=245
Score = 52.0 bits (123), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 37/122 (30%), Positives = 54/122 (44%), Gaps = 6/122 (4%)
Query 65 LNPRGYRSPAERGIPFKDVSFT---TRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGF 121
LN + P E D+ F T + K++A + PL T + HGN N+
Sbjct 5 LNRIIFNGPTEGYYEKFDLDFIYIETENNEKVAAHFINRNAPL---TILFCHGNGENVYM 61
Query 122 HQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFL 181
+ NV L DY GYG S GT SE +Y +A D++++T +N I L
Sbjct 62 LYDYFYETSKIWNVNVFLYDYLGYGESTGTASEKNMYLSGNAVYDYMVNTLKINPNSIVL 121
Query 182 FG 183
+G
Sbjct 122 YG 123
> ath:AT4G31020 hypothetical protein
Length=294
Score = 51.6 bits (122), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 33/100 (33%), Positives = 46/100 (46%), Gaps = 2/100 (2%)
Query 84 SFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYR 143
TT+ G K+ A P T + HGNA ++G L A + N++ DY
Sbjct 48 QLTTKSGNKVVA--TFWRHPFARFTLLYSHGNAADLGQMVELFIELRAHLRVNIMSYDYS 105
Query 144 GYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
GYG S G PSE Y D +A L S + ++I L+G
Sbjct 106 GYGASTGKPSEFNTYYDIEAVYSCLRSDYGIKQEEIILYG 145
> dre:100003419 si:rp71-61h23.3
Length=324
Score = 51.6 bits (122), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 28/71 (39%), Positives = 38/71 (53%), Gaps = 0/71 (0%)
Query 113 HGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTS 172
HGNA ++G L + N+ DY GYG S G PSE +YAD DAA L S
Sbjct 132 HGNAVDLGQMSSFYIGLGTRINCNIFSYDYSGYGVSTGKPSEKNLYADIDAAWQALRSRY 191
Query 173 LVNNKDIFLFG 183
++ ++I L+G
Sbjct 192 GISPENIILYG 202
> mmu:216169 Fam108a, 1700013O15Rik, BC005632, D10Bwg1364e, MGC11699,
MGC90979; family with sequence similarity 108, member
A
Length=310
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 28/71 (39%), Positives = 37/71 (52%), Gaps = 0/71 (0%)
Query 113 HGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTS 172
HGNA ++G L +G N+ DY GYG S G PSE +YAD DAA L +
Sbjct 118 HGNAVDLGQMCSFYVGLGTRIGCNIFSYDYSGYGISSGRPSEKNLYADIDAAWQALRTRY 177
Query 173 LVNNKDIFLFG 183
++ I L+G
Sbjct 178 GISPDSIILYG 188
> hsa:58489 FAM108C1, FLJ34461, MGC131546; family with sequence
similarity 108, member C1
Length=329
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 29/76 (38%), Positives = 41/76 (53%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + + N+ DY GYG S G PSE +YAD DAA
Sbjct 134 TLLFSHGNAVDLGQMCSFYIGLGSRINCNIFSYDYSGYGVSSGKPSEKNLYADIDAAWQA 193
Query 168 LLSTSLVNNKDIFLFG 183
L + V+ ++I L+G
Sbjct 194 LRTRYGVSPENIILYG 209
> mmu:70178 Fam108c, 2210412D01Rik, AL023007, Fam108c1; family
with sequence similarity 108, member C
Length=320
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 29/76 (38%), Positives = 41/76 (53%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + + N+ DY GYG S G PSE +YAD DAA
Sbjct 125 TLLFSHGNAVDLGQMCSFYIGLGSRINCNIFSYDYSGYGVSSGKPSEKNLYADIDAAWQA 184
Query 168 LLSTSLVNNKDIFLFG 183
L + V+ ++I L+G
Sbjct 185 LRTRYGVSPENIILYG 200
> pfa:PF11_0211 conserved Plasmodium protein
Length=382
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 39/159 (24%), Positives = 66/159 (41%), Gaps = 9/159 (5%)
Query 34 IGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGIP------FKDVSFTT 87
+G + + G R +K+ F+ GY N N + + I D+++
Sbjct 67 LGCMTICGFRGRMVKKMAFVPPIIKGYNIENDNKFIFHNSHHEEIKELMQINNIDINYKK 126
Query 88 -RDGVKLSAWLLLGEKPLE--APTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRG 144
+ G + ++L +KPL+ T + HGN +IG+ L NV DY G
Sbjct 127 LKRGSTEVSVIMLYKKPLDLNKQTILYSHGNTTDIGYMTPFLLNLVTSNNVNVFSYDYSG 186
Query 145 YGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
YG S PSE Y + D+L + ++I ++G
Sbjct 187 YGLSNKDPSEKNCYKSIKMSYDYLTKDLNIKPENIIVYG 225
> ath:AT3G01690 hypothetical protein
Length=361
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 31/101 (30%), Positives = 45/101 (44%), Gaps = 2/101 (1%)
Query 83 VSFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDY 142
V TR G ++ + P+ T + HGNA ++G L + N++ DY
Sbjct 47 VKLRTRRGTEIVGMYV--RHPMATSTLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDY 104
Query 143 RGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
GYG S G PSE YAD +A L T + + L+G
Sbjct 105 SGYGQSTGKPSEHNTYADIEAVYKCLEETFGSKQEGVILYG 145
> mmu:76192 Abhd12, 1500011G07Rik, 6330583M11Rik, AI431047, AW547313;
abhydrolase domain containing 12 (EC:3.1.1.23); K13704
abhydrolase domain-containing protein 12 [EC:3.1.1.23]
Length=398
Score = 50.8 bits (120), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 46/170 (27%), Positives = 75/170 (44%), Gaps = 27/170 (15%)
Query 33 FIGVVVLVGLLWRFQEKLLFLNAFPVGYKTPNLNPRGYRSPAERGI---------PFKDV 83
+I + LV L Q KL+FLN V Y + P ++G+ P DV
Sbjct 83 YIAIPFLVKLCPGIQAKLIFLNFVRVPYFI------DLKKPQDQGLNHTCNYYLQPEDDV 136
Query 84 SFTTRDGVKLSAW-------LLLGEKPLEA--PTFIQFHGNAGNIG-FHQGHAEVLCAEM 133
+ + W + E L + + HGNAG G H+ + + +
Sbjct 137 TIGVWHTIPSVWWKNAQGKDQMWYEDALASNHAIILYLHGNAGTRGGDHRVELYKVLSSL 196
Query 134 GANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
G +V+ DYRG+G+S GTPSE G+ DA D++ + S + ++++G
Sbjct 197 GYHVVTFDYRGWGDSVGTPSERGMTYDALHVFDWIKARS--GDNPVYIWG 244
> xla:734783 fam108a1, MGC131027, fam108a2; family with sequence
similarity 108, member A1
Length=305
Score = 50.8 bits (120), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 28/76 (36%), Positives = 40/76 (52%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + N+ DY GYG S G PSE +YAD DAA
Sbjct 107 TLLFSHGNAVDLGQMTSFYLDLGTRINCNIFSYDYSGYGCSSGRPSEKNLYADIDAAWHA 166
Query 168 LLSTSLVNNKDIFLFG 183
L + ++ ++I L+G
Sbjct 167 LRTRYGISPENILLYG 182
> dre:322121 fb50g01, wu:fb50g01; zgc:162293
Length=336
Score = 50.8 bits (120), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 28/71 (39%), Positives = 38/71 (53%), Gaps = 0/71 (0%)
Query 113 HGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTS 172
HGNA ++G L + N+ DY GYG S G PSE +YAD DAA L S
Sbjct 146 HGNAVDLGQMSSFYIGLGTRINCNIFSYDYSGYGVSTGKPSEKNLYADIDAAWHALRSRY 205
Query 173 LVNNKDIFLFG 183
++ ++I L+G
Sbjct 206 GISPENIILYG 216
> xla:100127338 hypothetical protein LOC100127338
Length=305
Score = 50.8 bits (120), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 28/76 (36%), Positives = 40/76 (52%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + N+ DY GYG S G PSE +YAD DAA
Sbjct 107 TLLFSHGNAVDLGQMTSFYLDLGTRINCNIFSYDYSGYGCSSGRPSEKNLYADIDAAWHA 166
Query 168 LLSTSLVNNKDIFLFG 183
L + ++ ++I L+G
Sbjct 167 LRTRYGISPENILLYG 182
> dre:393126 MGC55468, fam108c1; zgc:55468
Length=294
Score = 50.4 bits (119), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 42/147 (28%), Positives = 62/147 (42%), Gaps = 8/147 (5%)
Query 45 RFQEKLLFL--------NAFPVGYKTPNLNPRGYRSPAERGIPFKDVSFTTRDGVKLSAW 96
R KL FL + P G + +L R ++R + +V T
Sbjct 28 RIAAKLAFLPPEPTYSVHTDPSGATSLHLTERADWQYSQRELDAVEVLVTRTSRGNRVGC 87
Query 97 LLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESG 156
+ + P T + HGNA ++G L + + NV DY GYG S G PSE
Sbjct 88 MFVRCAPASRYTLLFSHGNAVDLGQMCSFYIGLGSRINCNVFSYDYSGYGVSTGKPSEKN 147
Query 157 VYADADAALDFLLSTSLVNNKDIFLFG 183
+YAD +AA L + V ++I L+G
Sbjct 148 LYADIEAAWQVLRNKYGVTPENIILYG 174
> ath:AT3G30380 hypothetical protein
Length=377
Score = 50.4 bits (119), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 30/98 (30%), Positives = 47/98 (47%), Gaps = 3/98 (3%)
Query 86 TTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGY 145
T R ++A++ + P + T + HGNA ++G L + N++ DY GY
Sbjct 50 TKRGNQVVAAYI---KNPTASLTLLYSHGNAADLGQMFELFSELSLHLRVNLIGYDYSGY 106
Query 146 GNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
G S G PSE Y+D +A L V +D+ L+G
Sbjct 107 GRSSGKPSEQNTYSDIEAVYRCLEEKYGVKEQDVILYG 144
> dre:437017 zgc:100937
Length=166
Score = 50.1 bits (118), Expect = 4e-06, Method: Compositional matrix adjust.
Identities = 27/69 (39%), Positives = 38/69 (55%), Gaps = 0/69 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + + NV DY GYG S G PSE +YAD DAA
Sbjct 93 TLLFSHGNAVDLGQMSSFYIGLGSRINCNVFSYDYSGYGASSGKPSEKNLYADVDAAWHA 152
Query 168 LLSTSLVNN 176
L + S++++
Sbjct 153 LRTRSMISS 161
> hsa:81926 FAM108A1, C19orf27, MGC5244; family with sequence
similarity 108, member A1
Length=310
Score = 50.1 bits (118), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 27/71 (38%), Positives = 37/71 (52%), Gaps = 0/71 (0%)
Query 113 HGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTS 172
HGNA ++G L + + N+ DY GYG S G PSE +YAD DAA L +
Sbjct 118 HGNAVDLGQMSSFYIGLGSRLHCNIFSYDYSGYGASSGRPSERNLYADIDAAWQALRTRY 177
Query 173 LVNNKDIFLFG 183
++ I L+G
Sbjct 178 GISPDSIILYG 188
> xla:446585 fam108b1, MGC81688; family with sequence similarity
108, member B1
Length=288
Score = 49.7 bits (117), Expect = 6e-06, Method: Compositional matrix adjust.
Identities = 29/89 (32%), Positives = 44/89 (49%), Gaps = 0/89 (0%)
Query 95 AWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSE 154
A + + P T + HGNA ++G L + + N+ DY GYG+S G PSE
Sbjct 80 ACMFVRCSPSAKYTLLFSHGNAVDLGQMSSFYIGLGSRINCNIFSYDYSGYGSSSGKPSE 139
Query 155 SGVYADADAALDFLLSTSLVNNKDIFLFG 183
+YAD DAA L + V + + ++G
Sbjct 140 KNLYADIDAAWIALRTRYGVRPEHVIIYG 168
> cel:K04G2.2 hypothetical protein
Length=332
Score = 49.7 bits (117), Expect = 6e-06, Method: Compositional matrix adjust.
Identities = 30/76 (39%), Positives = 38/76 (50%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + NV DY GYG S G PSE +YAD AA +
Sbjct 114 TLLFSHGNAVDLGQMTSFLYGLGFHLNCNVFSYDYSGYGCSTGKPSEKNLYADITAAFEL 173
Query 168 LLSTSLVNNKDIFLFG 183
L S V + I L+G
Sbjct 174 LKSEFGVPKEKIILYG 189
> tgo:TGME49_062490 hypothetical protein
Length=260
Score = 49.3 bits (116), Expect = 7e-06, Method: Compositional matrix adjust.
Identities = 33/90 (36%), Positives = 46/90 (51%), Gaps = 6/90 (6%)
Query 82 DVSF---TTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVL 138
D SF TTR ++ A+ + L T I HGNA +IG + + + N
Sbjct 21 DASFIWLTTRRRQRIPAFFIDIGASL---TIIFSHGNAEDIGMVIEYFKEVSRLWNCNFF 77
Query 139 LLDYRGYGNSEGTPSESGVYADADAALDFL 168
+ DY GYG+S G PSE GVY +AA ++L
Sbjct 78 VYDYVGYGHSTGKPSEQGVYDSVEAAFEYL 107
> ath:AT2G24320 hypothetical protein
Length=286
Score = 49.3 bits (116), Expect = 7e-06, Method: Compositional matrix adjust.
Identities = 30/100 (30%), Positives = 48/100 (48%), Gaps = 2/100 (2%)
Query 84 SFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYR 143
TT+ G K+ A + P T + HGNA ++G L A + N++ DY
Sbjct 40 QLTTKSGNKVIA--TFWKHPFSRFTLLYSHGNAADLGQMVDLFIELRAHLRVNIMSYDYS 97
Query 144 GYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
GYG S G P+E Y D +A + L + + +++ L+G
Sbjct 98 GYGASTGKPTELNTYYDIEAVYNCLRTEYGIMQEEMILYG 137
> xla:447065 fam108b1, MGC83647; abhydrolase domain-containing
protein FAM108B1
Length=288
Score = 48.5 bits (114), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 28/89 (31%), Positives = 44/89 (49%), Gaps = 0/89 (0%)
Query 95 AWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSE 154
A + + P T + HGNA ++G L + + N+ DY GYG+S G PSE
Sbjct 80 ACMFVRCCPSAKYTLLFSHGNAVDLGQMSSFYIGLGSRINCNIFSYDYSGYGSSSGKPSE 139
Query 155 SGVYADADAALDFLLSTSLVNNKDIFLFG 183
+YAD DAA L + + + + ++G
Sbjct 140 KNLYADIDAAWIALRTRYGIRPEHVIIYG 168
> xla:446755 fam108c1, MGC79044; family with sequence similarity
108, member C1
Length=311
Score = 48.5 bits (114), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 28/76 (36%), Positives = 39/76 (51%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + N+ DY GYG S G PSE +YAD +AA
Sbjct 116 TLLFSHGNAVDLGQMCSFYIGLGTRINCNIFSYDYSGYGVSSGKPSEKNLYADIEAAWHA 175
Query 168 LLSTSLVNNKDIFLFG 183
L + V ++I L+G
Sbjct 176 LRTRYGVTPENIILYG 191
> dre:751622 MGC153037, zgc:153037; si:ch211-117n7.7
Length=347
Score = 48.5 bits (114), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 33/94 (35%), Positives = 50/94 (53%), Gaps = 20/94 (21%)
Query 87 TRDGVKLSAWLLLGE--------KPLE---------APTFIQFHGNAGNIGF-HQ-GHAE 127
T +GV++ W + E K +E +P F+ HGN GN H+ G A
Sbjct 83 TEEGVRVGVWHTVPEHRWKEAQGKNVEWYEKALGDGSPIFMYLHGNTGNRSAPHRIGVAN 142
Query 128 VLCAEMGANVLLLDYRGYGNSEGTPSESGVYADA 161
+L A +G + L++DYRG+G+S G P+E G+ DA
Sbjct 143 ILSA-LGYHALVMDYRGFGDSTGEPTEPGLTTDA 175
> cel:Y97E10AL.2 hypothetical protein; K13704 abhydrolase domain-containing
protein 12 [EC:3.1.1.23]
Length=345
Score = 48.1 bits (113), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 22/57 (38%), Positives = 34/57 (59%), Gaps = 1/57 (1%)
Query 113 HGNAGNIGF-HQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDFL 168
HGN+ + F H+ L ++ +V+ DYRGYG+SEGTP+E G+ D ++L
Sbjct 120 HGNSFDRTFYHRVEMYNLLSDCNYHVVCFDYRGYGDSEGTPTEKGIVEDTKTVYEWL 176
> tpv:TP03_0361 hypothetical protein
Length=315
Score = 47.0 bits (110), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 30/97 (30%), Positives = 50/97 (51%), Gaps = 2/97 (2%)
Query 87 TRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYG 146
T DG ++++ + + T I H NA +IG G+ + N+ + DY GYG
Sbjct 30 TPDGNTIASYFI--KHKFAKFTIIFSHANAEDIGNVFGNLIKRLTKWNCNLFIYDYPGYG 87
Query 147 NSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
S G SE +Y AD + ++L++T VN+ +I +G
Sbjct 88 LSSGVCSEENMYNCADLSYNYLINTLKVNSGNIIAYG 124
> ath:AT5G38220 hydrolase
Length=336
Score = 46.6 bits (109), Expect = 5e-05, Method: Compositional matrix adjust.
Identities = 34/110 (30%), Positives = 50/110 (45%), Gaps = 6/110 (5%)
Query 78 IPFKD----VSFTTRDGVKLSAWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEM 133
+P +D + TR G ++ A + + P T + HGNA ++G L +
Sbjct 36 VPRRDDVDVLKLKTRRGNEIVAIYI--KHPKANGTLLYSHGNAADLGQMFELFIELSNRL 93
Query 134 GANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
N++ DY GYG S G SE YAD DAA L V + + L+G
Sbjct 94 RLNLMGYDYSGYGQSTGKASECNTYADIDAAYTCLKEHYGVKDDQLILYG 143
> mmu:226016 Fam108b, 5730446C15Rik, Cgi67, Fam108b1, MGC40949;
family with sequence similarity 108, member B
Length=288
Score = 45.8 bits (107), Expect = 7e-05, Method: Compositional matrix adjust.
Identities = 26/89 (29%), Positives = 44/89 (49%), Gaps = 0/89 (0%)
Query 95 AWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSE 154
A + + P T + HGNA ++G L + + N+ DY GYG S G P+E
Sbjct 80 ACMFVRCSPNAKYTLLFSHGNAVDLGQMSSFYIGLGSRINCNIFSYDYSGYGASSGKPTE 139
Query 155 SGVYADADAALDFLLSTSLVNNKDIFLFG 183
+YAD +AA L + + +++ ++G
Sbjct 140 KNLYADVEAAWLALRTRYGIRPENVIIYG 168
> hsa:51104 FAM108B1, C9orf77, RP11-409O11.2; family with sequence
similarity 108, member B1
Length=288
Score = 45.4 bits (106), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 26/89 (29%), Positives = 44/89 (49%), Gaps = 0/89 (0%)
Query 95 AWLLLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSE 154
A + + P T + HGNA ++G L + + N+ DY GYG S G P+E
Sbjct 80 ACMFVRCSPNAKYTLLFSHGNAVDLGQMSSFYIGLGSRINCNIFSYDYSGYGASSGKPTE 139
Query 155 SGVYADADAALDFLLSTSLVNNKDIFLFG 183
+YAD +AA L + + +++ ++G
Sbjct 140 KNLYADIEAAWLALRTRYGIRPENVIIYG 168
> tgo:TGME49_023510 hypothetical protein
Length=452
Score = 45.4 bits (106), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 29/86 (33%), Positives = 39/86 (45%), Gaps = 0/86 (0%)
Query 98 LLGEKPLEAPTFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGV 157
++ E P I HGN+ +IGF G L + NVL DY GYG S G SE +
Sbjct 190 VMHESAKRLPCIIFSHGNSTDIGFMFGLYYRLAYKCRVNVLAYDYSGYGCSGGKTSEKAL 249
Query 158 YADADAALDFLLSTSLVNNKDIFLFG 183
Y + A + V + I L+G
Sbjct 250 YRNIRAVWTYATQMLHVPPRQIILYG 275
> tpv:TP04_0691 hypothetical protein
Length=378
Score = 43.5 bits (101), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 26/84 (30%), Positives = 41/84 (48%), Gaps = 1/84 (1%)
Query 101 EKPLEAPTFIQF-HGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYA 159
EK +I F HGN +IG LCA + N++ DY GYG+S G SE +Y+
Sbjct 101 EKRKNEEIYILFSHGNNTDIGHMFFKYTRLCAFLNVNLVSYDYSGYGHSSGKASEGNMYS 160
Query 160 DADAALDFLLSTSLVNNKDIFLFG 183
+ ++ + + + I L+G
Sbjct 161 NIANVYKYMTNKMKLGPRQIVLYG 184
> ath:AT1G66900 hypothetical protein
Length=272
Score = 42.7 bits (99), Expect = 7e-04, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 36/76 (47%), Gaps = 0/76 (0%)
Query 108 TFIQFHGNAGNIGFHQGHAEVLCAEMGANVLLLDYRGYGNSEGTPSESGVYADADAALDF 167
T + HGNA ++G L + N++ DY GYG S G SE YAD +A+
Sbjct 71 TLLYSHGNAADLGQMFELFVELSNRLRVNLMGYDYSGYGQSTGQASECNTYADIEASYKC 130
Query 168 LLSTSLVNNKDIFLFG 183
L V + + ++G
Sbjct 131 LKEKYGVKDDQLIVYG 146
> cel:Y71G12A.4 hypothetical protein
Length=463
Score = 41.2 bits (95), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 0/51 (0%)
Query 133 MGANVLLLDYRGYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
+ ++L+ DY GYG SEGT +E VYA +A + + + T + I L G
Sbjct 242 LQCDLLIYDYPGYGVSEGTTNEKNVYAAVEAVMKYAMGTLGYSQDKIILIG 292
> tgo:TGME49_010350 hypothetical protein
Length=2760
Score = 40.8 bits (94), Expect = 0.002, Method: Composition-based stats.
Identities = 30/100 (30%), Positives = 46/100 (46%), Gaps = 3/100 (3%)
Query 87 TRDGVKLSAWLLLG--EKPLEAPTFIQF-HGNAGNIGFHQGHAEVLCAEMGANVLLLDYR 143
+R GV+ + L G + AP + + HGN +IG + +L + A+ LL DY
Sbjct 435 SRFGVESVSALFSGPTAAAVAAPFLVIYAHGNGSDIGDVHARSALLAERLQASFLLFDYP 494
Query 144 GYGNSEGTPSESGVYADADAALDFLLSTSLVNNKDIFLFG 183
GYG G E+ V A + L F + + I L+G
Sbjct 495 GYGKYAGAADEASVDATLRSVLAFAIHCLGWPQQKIVLWG 534
Lambda K H
0.323 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4976880524
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40