bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
Query= Eace_0384_orf1
Length=149
Score E
Sequences producing significant alignments: (Bits) Value
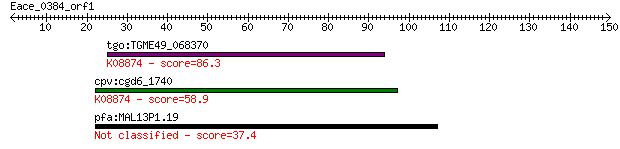
tgo:TGME49_068370 hypothetical protein ; K08874 transformation... 86.3 4e-17
cpv:cgd6_1740 Tra1p-like; C-terminal FAT domain plus phoshoino... 58.9 5e-09
pfa:MAL13P1.19 peptidase, putative 37.4 0.018
> tgo:TGME49_068370 hypothetical protein ; K08874 transformation/transcription
domain-associated protein
Length=8430
Score = 86.3 bits (212), Expect = 4e-17, Method: Compositional matrix adjust.
Identities = 39/69 (56%), Positives = 51/69 (73%), Gaps = 0/69 (0%)
Query 25 LNRLLLRHNPSRRRGLAFPLSAQVSIHPRCRLLEDMPSCITLQQILNEASRAAAEPMKQD 84
+N+LL + +R RGL+F L AQ+ +HPRCRL+ED PS +TLQ+IL E S + +E D
Sbjct 7957 INQLLFKFKDTRGRGLSFTLHAQIPVHPRCRLMEDPPSLLTLQEILQEVSASGSETCLVD 8016
Query 85 PDLPVLLHR 93
PDLPVLLHR
Sbjct 8017 PDLPVLLHR 8025
> cpv:cgd6_1740 Tra1p-like; C-terminal FAT domain plus phoshoinositide
3-kinase domain; very large protein ; K08874 transformation/transcription
domain-associated protein
Length=5542
Score = 58.9 bits (141), Expect = 5e-09, Method: Compositional matrix adjust.
Identities = 26/77 (33%), Positives = 48/77 (62%), Gaps = 3/77 (3%)
Query 22 QLQLNRLLLRHNPSRRRGLAFPLSAQVSIHPRCRLLEDMPSCITLQQILNEASRAA--AE 79
Q+ LN LL ++ +R R ++F + + +HPRCR++ED + + ++ E + ++
Sbjct 5179 QVTLNTLLYKYKETRSRNISFAIQPCIPLHPRCRIIEDTGNKKSFTELFEEEANSSNCTC 5238
Query 80 PMKQDPDLPVLLHRKMM 96
P+K D D P+LLHRK++
Sbjct 5239 PIK-DLDFPILLHRKLI 5254
> pfa:MAL13P1.19 peptidase, putative
Length=9271
Score = 37.4 bits (85), Expect = 0.018, Method: Composition-based stats.
Identities = 23/85 (27%), Positives = 42/85 (49%), Gaps = 1/85 (1%)
Query 22 QLQLNRLLLRHNPSRRRGLAFPLSAQVSIHPRCRLLEDMPSCITLQQILNEASRAAAEPM 81
Q+ LN ++ + +++RGL+F ++ IHP L E ITL ++ NE +
Sbjct 8551 QVLLNNIMFNFSQTKKRGLSFKINFNTQIHPNIILKEQPKDDITLHEVYNEYI-SVLRSD 8609
Query 82 KQDPDLPVLLHRKMMAHEISSKMLQ 106
K D +L HR ++ + +L+
Sbjct 8610 KYSMDNIMLYHRNLVQSSLHRLLLK 8634
Lambda K H
0.315 0.123 0.345
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3068761412
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40