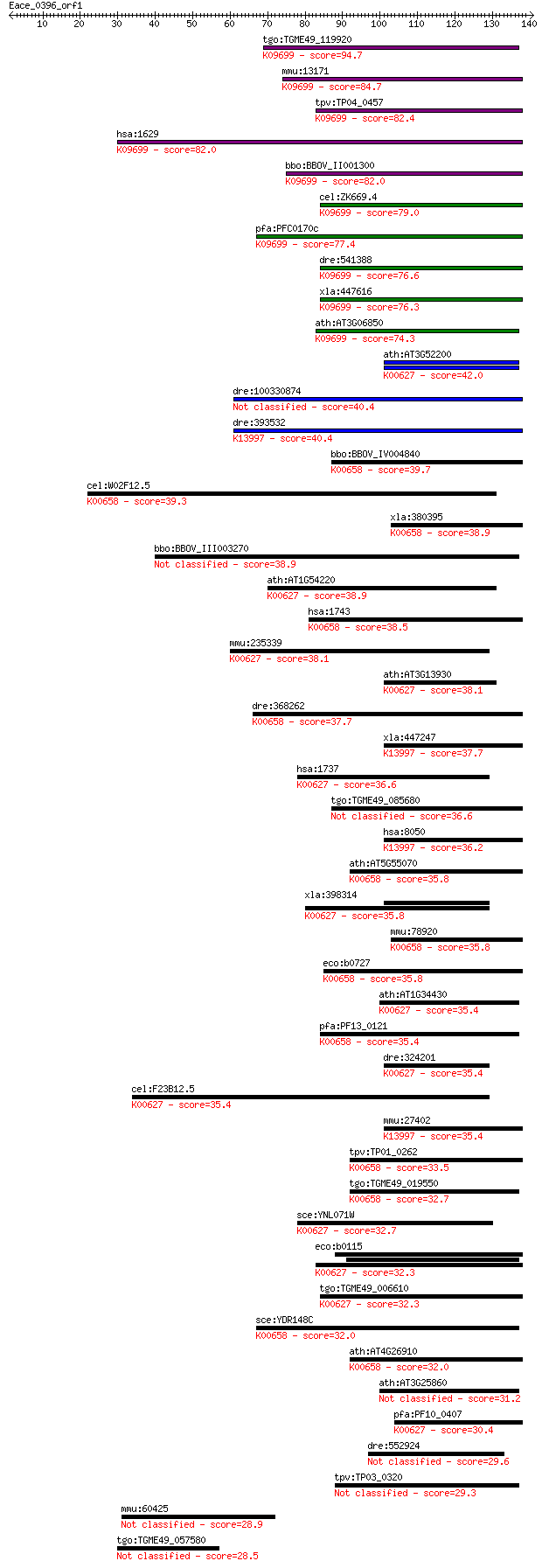
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
Query= Eace_0396_orf1
Length=140
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_119920 dihydrolipoamide branched chain transacylase... 94.7 9e-20
mmu:13171 Dbt, D3Wsu60e; dihydrolipoamide branched chain trans... 84.7 9e-17
tpv:TP04_0457 lipoamide transferase (EC:2.3.1.-); K09699 2-oxo... 82.4 3e-16
hsa:1629 DBT, BCATE2, E2, E2B, MGC9061; dihydrolipoamide branc... 82.0 5e-16
bbo:BBOV_II001300 18.m06098; lipoamide acyltransferase compone... 82.0 5e-16
cel:ZK669.4 hypothetical protein; K09699 2-oxoisovalerate dehy... 79.0 5e-15
pfa:PFC0170c dihydrolipoamide acyltransferase, putative (EC:2.... 77.4 1e-14
dre:541388 dbt, im:7147214, zgc:103768; dihydrolipoamide branc... 76.6 2e-14
xla:447616 dbt, MGC85493; dihydrolipoamide branched chain tran... 76.3 3e-14
ath:AT3G06850 BCE2; BCE2; acetyltransferase/ alpha-ketoacid de... 74.3 1e-13
ath:AT3G52200 LTA3; LTA3; ATP binding / dihydrolipoyllysine-re... 42.0 6e-04
dre:100330874 pyruvate dehydrogenase complex, component X-like 40.4 0.002
dre:393532 pdhx, MGC66110, zgc:66110, zgc:85977; pyruvate dehy... 40.4 0.002
bbo:BBOV_IV004840 23.m06243; dihydrolipoamide succinyltransfer... 39.7 0.003
cel:W02F12.5 hypothetical protein; K00658 2-oxoglutarate dehyd... 39.3 0.004
xla:380395 dlst, MGC53142; dihydrolipoamide S-succinyltransfer... 38.9 0.004
bbo:BBOV_III003270 17.m07312; biotin-requiring enzyme family p... 38.9 0.005
ath:AT1G54220 dihydrolipoamide S-acetyltransferase, putative (... 38.9 0.005
hsa:1743 DLST, DLTS; dihydrolipoamide S-succinyltransferase (E... 38.5 0.007
mmu:235339 Dlat, 6332404G05Rik, DLTA, PDC-E2; dihydrolipoamide... 38.1 0.008
ath:AT3G13930 dihydrolipoamide S-acetyltransferase, putative (... 38.1 0.009
dre:368262 dlst, cb60, wu:fc39d01; dihydrolipoamide S-succinyl... 37.7 0.010
xla:447247 pdhx, MGC86218; pyruvate dehydrogenase complex, com... 37.7 0.011
hsa:1737 DLAT, DLTA, PDC-E2, PDCE2; dihydrolipoamide S-acetylt... 36.6 0.026
tgo:TGME49_085680 dihydrolipoamide acyltransferase, putative (... 36.6 0.028
hsa:8050 PDHX, DLDBP, E3BP, OPDX, PDX1, proX; pyruvate dehydro... 36.2 0.036
ath:AT5G55070 2-oxoacid dehydrogenase family protein (EC:2.3.1... 35.8 0.040
xla:398314 dlat; dihydrolipoamide S-acetyltransferase (EC:2.3.... 35.8 0.041
mmu:78920 Dlst, 1600017E01Rik, 4632413C10Rik, 4930529O08Rik, D... 35.8 0.042
eco:b0727 sucB, ECK0715, JW0716; dihydrolipoyltranssuccinase (... 35.8 0.043
ath:AT1G34430 EMB3003 (embryo defective 3003); acyltransferase... 35.4 0.050
pfa:PF13_0121 dihydrolipamide succinyltransferase component of... 35.4 0.051
dre:324201 dlat, wu:fc14f10, wu:fc21f08, wu:fc86g11, wu:fj57d0... 35.4 0.052
cel:F23B12.5 hypothetical protein; K00627 pyruvate dehydrogena... 35.4 0.052
mmu:27402 Pdhx, AI481367, E3bp, Pdx1; pyruvate dehydrogenase c... 35.4 0.054
tpv:TP01_0262 dihydrolipoamide succinyltransferase; K00658 2-o... 33.5 0.23
tgo:TGME49_019550 dihydrolipoamide succinyltransferase compone... 32.7 0.32
sce:YNL071W LAT1, ODP2, PDA2; Dihydrolipoamide acetyltransfera... 32.7 0.33
eco:b0115 aceF, ECK0114, JW0111; pyruvate dehydrogenase, dihyd... 32.3 0.49
tgo:TGME49_006610 biotin requiring domain-containing protein /... 32.3 0.54
sce:YDR148C KGD2; Dihydrolipoyl transsuccinylase, component of... 32.0 0.54
ath:AT4G26910 2-oxoacid dehydrogenase family protein (EC:2.3.1... 32.0 0.63
ath:AT3G25860 LTA2; LTA2; dihydrolipoyllysine-residue acetyltr... 31.2 0.92
pfa:PF10_0407 dihydrolipoamide acyltransferase, putative (EC:2... 30.4 1.7
dre:552924 slc2a15b, zgc:113085; solute carrier family 2 (faci... 29.6 2.8
tpv:TP03_0320 hypothetical protein 29.3 3.6
mmu:60425 Doc2g, D830013O18Rik; double C2, gamma 28.9 4.6
tgo:TGME49_057580 hypothetical protein 28.5 7.7
> tgo:TGME49_119920 dihydrolipoamide branched chain transacylase,
E2 subunit, putative (EC:2.3.1.168); K09699 2-oxoisovalerate
dehydrogenase E2 component (dihydrolipoyl transacylase)
[EC:2.3.1.168]
Length=510
Score = 94.7 bits (234), Expect = 9e-20, Method: Compositional matrix adjust.
Identities = 43/68 (63%), Positives = 53/68 (77%), Gaps = 0/68 (0%)
Query 69 PSARGIFSISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
P R + ++S+P + FKLADIGEGIA VEL KW+K GD +EEM+E+CEVQSDKAAV
Sbjct 56 PGKRCLLTVSRPALAVKTFKLADIGEGIAQVELLKWHKGVGDHVEEMDELCEVQSDKAAV 115
Query 129 EITSRYPG 136
EITSR+ G
Sbjct 116 EITSRFTG 123
> mmu:13171 Dbt, D3Wsu60e; dihydrolipoamide branched chain transacylase
E2 (EC:2.3.1.168); K09699 2-oxoisovalerate dehydrogenase
E2 component (dihydrolipoyl transacylase) [EC:2.3.1.168]
Length=482
Score = 84.7 bits (208), Expect = 9e-17, Method: Composition-based stats.
Identities = 40/74 (54%), Positives = 50/74 (67%), Gaps = 10/74 (13%)
Query 74 IFSISQPRHG----------IVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQS 123
+F SQPRH +V FKL+DIGEGI V + +WY KEGDT+ + + +CEVQS
Sbjct 44 LFKYSQPRHSLRTAAVLQGQVVQFKLSDIGEGIREVTIKEWYVKEGDTVSQFDSICEVQS 103
Query 124 DKAAVEITSRYPGV 137
DKA+V ITSRY GV
Sbjct 104 DKASVTITSRYDGV 117
> tpv:TP04_0457 lipoamide transferase (EC:2.3.1.-); K09699 2-oxoisovalerate
dehydrogenase E2 component (dihydrolipoyl transacylase)
[EC:2.3.1.168]
Length=420
Score = 82.4 bits (202), Expect = 3e-16, Method: Compositional matrix adjust.
Identities = 38/55 (69%), Positives = 42/55 (76%), Gaps = 0/55 (0%)
Query 83 GIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+ FKL+DIGEGI V+L KW K GD +EEME VC VQSDKAAVEITSRY GV
Sbjct 39 ALTTFKLSDIGEGINEVQLVKWEKNVGDEVEEMESVCTVQSDKAAVEITSRYTGV 93
> hsa:1629 DBT, BCATE2, E2, E2B, MGC9061; dihydrolipoamide branched
chain transacylase E2 (EC:2.3.1.168); K09699 2-oxoisovalerate
dehydrogenase E2 component (dihydrolipoyl transacylase)
[EC:2.3.1.168]
Length=482
Score = 82.0 bits (201), Expect = 5e-16, Method: Composition-based stats.
Identities = 50/121 (41%), Positives = 62/121 (51%), Gaps = 17/121 (14%)
Query 30 MGRVAAATSYSSNAGGLCTFLAPVAVAGVRVPRQG---LFKTPSARGIFSISQPRH---- 82
M V ++S NAG L V V + F PS F S P H
Sbjct 1 MAAVRMLRTWSRNAGKLICVRYFQTCGNVHVLKPNYVCFFGYPS----FKYSHPHHFLKT 56
Query 83 ------GIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
+V FKL+DIGEGI V + +WY KEGDT+ + + +CEVQSDKA+V ITSRY G
Sbjct 57 TAALRGQVVQFKLSDIGEGIREVTVKEWYVKEGDTVSQFDSICEVQSDKASVTITSRYDG 116
Query 137 V 137
V
Sbjct 117 V 117
> bbo:BBOV_II001300 18.m06098; lipoamide acyltransferase component
of branched-chain alpha-keto acid dehydrogenase complex
(EC:2.3.1.12); K09699 2-oxoisovalerate dehydrogenase E2 component
(dihydrolipoyl transacylase) [EC:2.3.1.168]
Length=417
Score = 82.0 bits (201), Expect = 5e-16, Method: Compositional matrix adjust.
Identities = 38/63 (60%), Positives = 46/63 (73%), Gaps = 0/63 (0%)
Query 75 FSISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRY 134
F S R+ + F L+DIGEGI+ VEL +W K GD +EEME VC VQSDKAAV+ITSRY
Sbjct 22 FHRSVHRNKLTTFHLSDIGEGISEVELVRWNKNVGDEVEEMETVCTVQSDKAAVDITSRY 81
Query 135 PGV 137
G+
Sbjct 82 TGL 84
> cel:ZK669.4 hypothetical protein; K09699 2-oxoisovalerate dehydrogenase
E2 component (dihydrolipoyl transacylase) [EC:2.3.1.168]
Length=448
Score = 79.0 bits (193), Expect = 5e-15, Method: Composition-based stats.
Identities = 35/54 (64%), Positives = 45/54 (83%), Gaps = 0/54 (0%)
Query 84 IVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+V FKL+DIGEGIA V++ +WY KEGDTI + ++VCEVQSDKAAV I+ RY G+
Sbjct 30 VVQFKLSDIGEGIAEVQVKEWYVKEGDTISQFDKVCEVQSDKAAVTISCRYDGI 83
> pfa:PFC0170c dihydrolipoamide acyltransferase, putative (EC:2.3.1.-);
K09699 2-oxoisovalerate dehydrogenase E2 component
(dihydrolipoyl transacylase) [EC:2.3.1.168]
Length=448
Score = 77.4 bits (189), Expect = 1e-14, Method: Composition-based stats.
Identities = 39/71 (54%), Positives = 48/71 (67%), Gaps = 0/71 (0%)
Query 67 KTPSARGIFSISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKA 126
K+ R + S IV KL DIGEGI+ VE+TKW+K EGD + EME + VQSDKA
Sbjct 16 KSFKGRHYLNTSAIHFKIVKCKLFDIGEGISEVEITKWHKNEGDQVSEMESLLTVQSDKA 75
Query 127 AVEITSRYPGV 137
AV+ITS+Y GV
Sbjct 76 AVDITSKYNGV 86
> dre:541388 dbt, im:7147214, zgc:103768; dihydrolipoamide branched
chain transacylase E2 (EC:2.3.1.168); K09699 2-oxoisovalerate
dehydrogenase E2 component (dihydrolipoyl transacylase)
[EC:2.3.1.168]
Length=493
Score = 76.6 bits (187), Expect = 2e-14, Method: Compositional matrix adjust.
Identities = 34/54 (62%), Positives = 42/54 (77%), Gaps = 0/54 (0%)
Query 84 IVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
IV FKL+DIGEGI V + +WY KEGD + + + +CEVQSDKA+V ITSRY GV
Sbjct 63 IVQFKLSDIGEGIMEVTVKEWYVKEGDKVSQFDSICEVQSDKASVTITSRYDGV 116
> xla:447616 dbt, MGC85493; dihydrolipoamide branched chain transacylase
E2 (EC:2.3.1.168); K09699 2-oxoisovalerate dehydrogenase
E2 component (dihydrolipoyl transacylase) [EC:2.3.1.168]
Length=492
Score = 76.3 bits (186), Expect = 3e-14, Method: Composition-based stats.
Identities = 34/54 (62%), Positives = 42/54 (77%), Gaps = 0/54 (0%)
Query 84 IVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
IV FKL+DIGEGI V + WY KEGD++ + + +CEVQSDKA+V ITSRY GV
Sbjct 63 IVQFKLSDIGEGITEVTVKDWYVKEGDSVSQFDSICEVQSDKASVTITSRYDGV 116
> ath:AT3G06850 BCE2; BCE2; acetyltransferase/ alpha-ketoacid
dehydrogenase/ dihydrolipoamide branched chain acyltransferase
(EC:2.3.1.168); K09699 2-oxoisovalerate dehydrogenase E2
component (dihydrolipoyl transacylase) [EC:2.3.1.168]
Length=483
Score = 74.3 bits (181), Expect = 1e-13, Method: Composition-based stats.
Identities = 32/54 (59%), Positives = 41/54 (75%), Gaps = 0/54 (0%)
Query 83 GIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
G++ LA GEGIA EL KW+ KEGD++EE + +CEVQSDKA +EITSR+ G
Sbjct 74 GLIDVPLAQTGEGIAECELLKWFVKEGDSVEEFQPLCEVQSDKATIEITSRFKG 127
> ath:AT3G52200 LTA3; LTA3; ATP binding / dihydrolipoyllysine-residue
acetyltransferase; K00627 pyruvate dehydrogenase E2
component (dihydrolipoamide acetyltransferase) [EC:2.3.1.12]
Length=637
Score = 42.0 bits (97), Expect = 6e-04, Method: Composition-based stats.
Identities = 17/36 (47%), Positives = 25/36 (69%), Gaps = 0/36 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
+ KW KKEGD +E + +CE+++DKA VE S+ G
Sbjct 102 VVKWMKKEGDKVEVGDVLCEIETDKATVEFESQEEG 137
Score = 37.7 bits (86), Expect = 0.010, Method: Composition-based stats.
Identities = 16/36 (44%), Positives = 24/36 (66%), Gaps = 0/36 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
+ KW+KKEGD IE + + E+++DKA +E S G
Sbjct 229 IAKWWKKEGDKIEVGDVIGEIETDKATLEFESLEEG 264
> dre:100330874 pyruvate dehydrogenase complex, component X-like
Length=181
Score = 40.4 bits (93), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 22/79 (27%), Positives = 40/79 (50%), Gaps = 2/79 (2%)
Query 61 PRQGLFKTPSARGIFSISQPRHGIVCFK--LADIGEGIAAVELTKWYKKEGDTIEEMEEV 118
PR + +AR ++ + R G+ K + + + + KW KKEG+ + + +
Sbjct 37 PRTAGWSRSAARPLYQSCRARAGVCPLKVQMPALSPTMEEGNIVKWLKKEGEDVAAGDAL 96
Query 119 CEVQSDKAAVEITSRYPGV 137
CE+++DKA V + S GV
Sbjct 97 CEIETDKAVVVMESNEDGV 115
> dre:393532 pdhx, MGC66110, zgc:66110, zgc:85977; pyruvate dehydrogenase
complex, component X; K13997 dihydrolipoamide dehydrogenase-binding
protein of pyruvate dehydrogenase complex
Length=490
Score = 40.4 bits (93), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 22/79 (27%), Positives = 40/79 (50%), Gaps = 2/79 (2%)
Query 61 PRQGLFKTPSARGIFSISQPRHGIVCFK--LADIGEGIAAVELTKWYKKEGDTIEEMEEV 118
PR + +AR ++ + R G+ K + + + + KW KKEG+ + + +
Sbjct 37 PRTAGWSRSAARPLYQSCRARAGVCPLKVQMPALSPTMEEGNIVKWLKKEGEDVAAGDAL 96
Query 119 CEVQSDKAAVEITSRYPGV 137
CE+++DKA V + S GV
Sbjct 97 CEIETDKAVVVMESNEDGV 115
> bbo:BBOV_IV004840 23.m06243; dihydrolipoamide succinyltransferase
(EC:2.3.1.61); K00658 2-oxoglutarate dehydrogenase E2
component (dihydrolipoamide succinyltransferase) [EC:2.3.1.61]
Length=402
Score = 39.7 bits (91), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 18/51 (35%), Positives = 30/51 (58%), Gaps = 0/51 (0%)
Query 87 FKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
KL +G+ I+ L++W K G+++E E + V++DK V+I S GV
Sbjct 58 MKLPSLGDSISEGTLSEWKKNVGESVEVDEPIAIVETDKVTVDINSTLSGV 108
> cel:W02F12.5 hypothetical protein; K00658 2-oxoglutarate dehydrogenase
E2 component (dihydrolipoamide succinyltransferase)
[EC:2.3.1.61]
Length=463
Score = 39.3 bits (90), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 29/114 (25%), Positives = 52/114 (45%), Gaps = 15/114 (13%)
Query 22 RMACVARYMGR-----VAAATSYSSNAGGLCTFLAPVAVAGVRVPRQGLFKTPSARGIFS 76
R V R++ R V AA+S + + L P+ V VR+ F + R
Sbjct 5 RAVSVHRFLSRAARQSVTAASSAQPSLQAKTSLLEPL-VQNVRITSSANFHMSAVR---- 59
Query 77 ISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEI 130
++ + E I+ ++ +W K++GD + E E V E+++DK +VE+
Sbjct 60 ----MSDVITVEGPAFAESISEGDI-RWLKQKGDHVNEDELVAEIETDKTSVEV 108
> xla:380395 dlst, MGC53142; dihydrolipoamide S-succinyltransferase
(E2 component of 2-oxo-glutarate complex) (EC:2.3.1.61);
K00658 2-oxoglutarate dehydrogenase E2 component (dihydrolipoamide
succinyltransferase) [EC:2.3.1.61]
Length=452
Score = 38.9 bits (89), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 16/35 (45%), Positives = 24/35 (68%), Gaps = 0/35 (0%)
Query 103 KWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+W K GDT+ E E VCE+++DK +V++ S GV
Sbjct 86 RWEKAVGDTVSEDEVVCEIETDKTSVQVPSPSAGV 120
> bbo:BBOV_III003270 17.m07312; biotin-requiring enzyme family
protein
Length=177
Score = 38.9 bits (89), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 24/102 (23%), Positives = 44/102 (43%), Gaps = 14/102 (13%)
Query 40 SSNAGGLCTFLA-----PVAVAGVRVPRQGLFKTPSARGIFSISQPRHGIVCFKLADIGE 94
S+N C LA V G+ P+ TP G I K+ IG
Sbjct 41 STNDIAYCQDLALRTHSTVDAKGLLQPKNDELGTPVDTGTLFI---------VKVPHIGR 91
Query 95 GIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
+ ++ +W+K+ GD ++ + +C +++D+ V + S+ G
Sbjct 92 DVKHSKIQQWHKQRGDEVDVGDLICVLETDQVLVNVQSQLSG 133
> ath:AT1G54220 dihydrolipoamide S-acetyltransferase, putative
(EC:2.3.1.12); K00627 pyruvate dehydrogenase E2 component (dihydrolipoamide
acetyltransferase) [EC:2.3.1.12]
Length=539
Score = 38.9 bits (89), Expect = 0.005, Method: Composition-based stats.
Identities = 23/62 (37%), Positives = 34/62 (54%), Gaps = 3/62 (4%)
Query 70 SARGIFSISQ-PRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
SARG S S P H + + + + + +W KKEGD + E +CEV++DKA V
Sbjct 98 SARGFSSGSDLPPHQEI--GMPSLSPTMTEGNIARWLKKEGDKVAPGEVLCEVETDKATV 155
Query 129 EI 130
E+
Sbjct 156 EM 157
> hsa:1743 DLST, DLTS; dihydrolipoamide S-succinyltransferase
(E2 component of 2-oxo-glutarate complex) (EC:2.3.1.61); K00658
2-oxoglutarate dehydrogenase E2 component (dihydrolipoamide
succinyltransferase) [EC:2.3.1.61]
Length=453
Score = 38.5 bits (88), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 19/57 (33%), Positives = 32/57 (56%), Gaps = 1/57 (1%)
Query 81 RHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+ +V K E + ++ +W K GDT+ E E VCE+++DK +V++ S GV
Sbjct 67 KDDLVTVKTPAFAESVTEGDV-RWEKAVGDTVAEDEVVCEIETDKTSVQVPSPANGV 122
> mmu:235339 Dlat, 6332404G05Rik, DLTA, PDC-E2; dihydrolipoamide
S-acetyltransferase (E2 component of pyruvate dehydrogenase
complex) (EC:2.3.1.12); K00627 pyruvate dehydrogenase E2
component (dihydrolipoamide acetyltransferase) [EC:2.3.1.12]
Length=642
Score = 38.1 bits (87), Expect = 0.008, Method: Compositional matrix adjust.
Identities = 27/73 (36%), Positives = 38/73 (52%), Gaps = 9/73 (12%)
Query 60 VPRQGLFK----TPSARGIFSISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEM 115
VPR L + +PS R S S P H V L + + A + +W KKEG+ I E
Sbjct 67 VPRNRLLRQLLGSPSRR---SYSLPPHQKV--PLPSLSPTMQAGTIARWEKKEGEKISEG 121
Query 116 EEVCEVQSDKAAV 128
+ + EV++DKA V
Sbjct 122 DLIAEVETDKATV 134
> ath:AT3G13930 dihydrolipoamide S-acetyltransferase, putative
(EC:2.3.1.12); K00627 pyruvate dehydrogenase E2 component (dihydrolipoamide
acetyltransferase) [EC:2.3.1.12]
Length=539
Score = 38.1 bits (87), Expect = 0.009, Method: Compositional matrix adjust.
Identities = 15/30 (50%), Positives = 22/30 (73%), Gaps = 0/30 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAVEI 130
+ +W KKEGD + E +CEV++DKA VE+
Sbjct 128 IARWLKKEGDKVAPGEVLCEVETDKATVEM 157
> dre:368262 dlst, cb60, wu:fc39d01; dihydrolipoamide S-succinyltransferase
(EC:2.3.1.61); K00658 2-oxoglutarate dehydrogenase
E2 component (dihydrolipoamide succinyltransferase) [EC:2.3.1.61]
Length=458
Score = 37.7 bits (86), Expect = 0.010, Method: Compositional matrix adjust.
Identities = 22/72 (30%), Positives = 38/72 (52%), Gaps = 9/72 (12%)
Query 66 FKTPSARGIFSISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDK 125
FKT +A R+ ++ K E + ++ +W K GD++ E E VCE+++DK
Sbjct 60 FKTTAAH--------RNEVITVKTPAFAESVTEGDV-RWEKAVGDSVAEDEVVCEIETDK 110
Query 126 AAVEITSRYPGV 137
+V++ S GV
Sbjct 111 TSVQVPSPAAGV 122
> xla:447247 pdhx, MGC86218; pyruvate dehydrogenase complex, component
X; K13997 dihydrolipoamide dehydrogenase-binding protein
of pyruvate dehydrogenase complex
Length=478
Score = 37.7 bits (86), Expect = 0.011, Method: Composition-based stats.
Identities = 15/37 (40%), Positives = 25/37 (67%), Gaps = 0/37 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+ KW KKEG+++ + +CE+++DKA V + S GV
Sbjct 60 IVKWLKKEGESVSAGDALCEIETDKAVVTMESNDDGV 96
> hsa:1737 DLAT, DLTA, PDC-E2, PDCE2; dihydrolipoamide S-acetyltransferase
(EC:2.3.1.12); K00627 pyruvate dehydrogenase E2
component (dihydrolipoamide acetyltransferase) [EC:2.3.1.12]
Length=647
Score = 36.6 bits (83), Expect = 0.026, Method: Compositional matrix adjust.
Identities = 20/51 (39%), Positives = 28/51 (54%), Gaps = 2/51 (3%)
Query 78 SQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
S P H V L + + A + +W KKEGD I E + + EV++DKA V
Sbjct 87 SLPPHQKV--PLPSLSPTMQAGTIARWEKKEGDKINEGDLIAEVETDKATV 135
> tgo:TGME49_085680 dihydrolipoamide acyltransferase, putative
(EC:2.4.1.115 2.3.1.61)
Length=470
Score = 36.6 bits (83), Expect = 0.028, Method: Compositional matrix adjust.
Identities = 18/51 (35%), Positives = 28/51 (54%), Gaps = 0/51 (0%)
Query 87 FKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
K+ +G+ I L +W KK GD + E +C +++DK VEI S G+
Sbjct 235 IKVPSLGDSITEGGLLEWRKKVGDFVLVDEVLCVIETDKVTVEIHSDCSGI 285
> hsa:8050 PDHX, DLDBP, E3BP, OPDX, PDX1, proX; pyruvate dehydrogenase
complex, component X; K13997 dihydrolipoamide dehydrogenase-binding
protein of pyruvate dehydrogenase complex
Length=486
Score = 36.2 bits (82), Expect = 0.036, Method: Composition-based stats.
Identities = 13/37 (35%), Positives = 24/37 (64%), Gaps = 0/37 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+ KW KKEG+ + + +CE+++DKA V + + G+
Sbjct 58 IVKWLKKEGEAVSAGDALCEIETDKAVVTLDASDDGI 94
> ath:AT5G55070 2-oxoacid dehydrogenase family protein (EC:2.3.1.61);
K00658 2-oxoglutarate dehydrogenase E2 component (dihydrolipoamide
succinyltransferase) [EC:2.3.1.61]
Length=464
Score = 35.8 bits (81), Expect = 0.040, Method: Compositional matrix adjust.
Identities = 16/46 (34%), Positives = 26/46 (56%), Gaps = 0/46 (0%)
Query 92 IGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+GE I L + KK GD +E E + ++++DK ++I S GV
Sbjct 101 MGESITDGTLAAFLKKPGDRVEADEAIAQIETDKVTIDIASPASGV 146
> xla:398314 dlat; dihydrolipoamide S-acetyltransferase (EC:2.3.1.12);
K00627 pyruvate dehydrogenase E2 component (dihydrolipoamide
acetyltransferase) [EC:2.3.1.12]
Length=628
Score = 35.8 bits (81), Expect = 0.041, Method: Compositional matrix adjust.
Identities = 14/28 (50%), Positives = 20/28 (71%), Gaps = 0/28 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
+ +W KKEGD I E + + EV++DKA V
Sbjct 89 IARWEKKEGDKINEGDLIAEVETDKATV 116
Score = 30.4 bits (67), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 14/49 (28%), Positives = 26/49 (53%), Gaps = 2/49 (4%)
Query 80 PRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
P H +C L + + + KW KK G+ + E + + E+++DKA +
Sbjct 193 PNHMKIC--LPALSPTMTMGTVQKWEKKVGEKLSEGDLLAEIETDKATI 239
> mmu:78920 Dlst, 1600017E01Rik, 4632413C10Rik, 4930529O08Rik,
DLTS; dihydrolipoamide S-succinyltransferase (E2 component
of 2-oxo-glutarate complex) (EC:2.3.1.61); K00658 2-oxoglutarate
dehydrogenase E2 component (dihydrolipoamide succinyltransferase)
[EC:2.3.1.61]
Length=454
Score = 35.8 bits (81), Expect = 0.042, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 23/35 (65%), Gaps = 0/35 (0%)
Query 103 KWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+W K GD + E E VCE+++DK +V++ S G+
Sbjct 89 RWEKAVGDAVAEDEVVCEIETDKTSVQVPSPANGI 123
> eco:b0727 sucB, ECK0715, JW0716; dihydrolipoyltranssuccinase
(EC:2.3.1.61); K00658 2-oxoglutarate dehydrogenase E2 component
(dihydrolipoamide succinyltransferase) [EC:2.3.1.61]
Length=405
Score = 35.8 bits (81), Expect = 0.043, Method: Compositional matrix adjust.
Identities = 15/53 (28%), Positives = 29/53 (54%), Gaps = 0/53 (0%)
Query 85 VCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
V + D+ E +A + W+KK GD + E + E+++DK +E+ + G+
Sbjct 4 VDILVPDLPESVADATVATWHKKPGDAVVRDEVLVEIETDKVVLEVPASADGI 56
> ath:AT1G34430 EMB3003 (embryo defective 3003); acyltransferase/
dihydrolipoyllysine-residue acetyltransferase/ protein binding;
K00627 pyruvate dehydrogenase E2 component (dihydrolipoamide
acetyltransferase) [EC:2.3.1.12]
Length=465
Score = 35.4 bits (80), Expect = 0.050, Method: Compositional matrix adjust.
Identities = 14/37 (37%), Positives = 23/37 (62%), Gaps = 0/37 (0%)
Query 100 ELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
++ W K EGD + + E V V+SDKA +++ + Y G
Sbjct 55 KIVSWVKSEGDKLNKGESVVVVESDKADMDVETFYDG 91
> pfa:PF13_0121 dihydrolipamide succinyltransferase component
of 2-oxoglutarate dehydrogenase complex (EC:2.3.1.61); K00658
2-oxoglutarate dehydrogenase E2 component (dihydrolipoamide
succinyltransferase) [EC:2.3.1.61]
Length=421
Score = 35.4 bits (80), Expect = 0.051, Method: Composition-based stats.
Identities = 16/53 (30%), Positives = 29/53 (54%), Gaps = 0/53 (0%)
Query 84 IVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
I K+ +G+ I + +W KK GD ++ E + + +DK +V+I S+ G
Sbjct 45 IETIKVPRLGDSITEGTINEWKKKVGDYVKADETITIIDTDKVSVDINSKVSG 97
> dre:324201 dlat, wu:fc14f10, wu:fc21f08, wu:fc86g11, wu:fj57d06;
dihydrolipoamide S-acetyltransferase (E2 component of pyruvate
dehydrogenase complex) (EC:2.3.1.12); K00627 pyruvate
dehydrogenase E2 component (dihydrolipoamide acetyltransferase)
[EC:2.3.1.12]
Length=652
Score = 35.4 bits (80), Expect = 0.052, Method: Compositional matrix adjust.
Identities = 14/28 (50%), Positives = 20/28 (71%), Gaps = 0/28 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
+ +W KKEGD I E + + EV++DKA V
Sbjct 109 IARWEKKEGDKINEGDLIAEVETDKATV 136
> cel:F23B12.5 hypothetical protein; K00627 pyruvate dehydrogenase
E2 component (dihydrolipoamide acetyltransferase) [EC:2.3.1.12]
Length=507
Score = 35.4 bits (80), Expect = 0.052, Method: Composition-based stats.
Identities = 25/95 (26%), Positives = 42/95 (44%), Gaps = 8/95 (8%)
Query 34 AAATSYSSNAGGLCTFLAPVAVAGVRVPRQGLFKTPSARGIFSISQPRHGIVCFKLADIG 93
A +T ++ + GL V + P F R S + P+H V L +
Sbjct 35 ALSTGAAAKSSGL------VGQVARQYPNAAAFSIKQVRLYSSGNLPKHNRVA--LPALS 86
Query 94 EGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAV 128
+ + W KKEGD + E + +CE+++DKA +
Sbjct 87 PTMELGTVVSWQKKEGDQLSEGDLLCEIETDKATM 121
> mmu:27402 Pdhx, AI481367, E3bp, Pdx1; pyruvate dehydrogenase
complex, component X; K13997 dihydrolipoamide dehydrogenase-binding
protein of pyruvate dehydrogenase complex
Length=501
Score = 35.4 bits (80), Expect = 0.054, Method: Composition-based stats.
Identities = 12/37 (32%), Positives = 24/37 (64%), Gaps = 0/37 (0%)
Query 101 LTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+ KW +KEG+ + + +CE+++DKA V + + G+
Sbjct 73 IVKWLRKEGEAVSAGDSLCEIETDKAVVTLDANDDGI 109
> tpv:TP01_0262 dihydrolipoamide succinyltransferase; K00658 2-oxoglutarate
dehydrogenase E2 component (dihydrolipoamide succinyltransferase)
[EC:2.3.1.61]
Length=456
Score = 33.5 bits (75), Expect = 0.23, Method: Composition-based stats.
Identities = 15/46 (32%), Positives = 27/46 (58%), Gaps = 0/46 (0%)
Query 92 IGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+G+ I+ LTKW GD + + + V++DK +V++ S + GV
Sbjct 80 LGDSISEGTLTKWAVSVGDYLNVDDLIAVVETDKVSVDVNSPFSGV 125
> tgo:TGME49_019550 dihydrolipoamide succinyltransferase component
of 2-oxoglutaratedehydrogenase complex, putative (EC:2.3.1.61);
K00658 2-oxoglutarate dehydrogenase E2 component (dihydrolipoamide
succinyltransferase) [EC:2.3.1.61]
Length=470
Score = 32.7 bits (73), Expect = 0.32, Method: Compositional matrix adjust.
Identities = 14/45 (31%), Positives = 26/45 (57%), Gaps = 0/45 (0%)
Query 92 IGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
+G+ I L +W K+ G+ ++E E V + +DK +V+I + G
Sbjct 101 MGDSITEGSLNEWKKQPGEYVKEGELVAVIDTDKVSVDINAPQAG 145
> sce:YNL071W LAT1, ODP2, PDA2; Dihydrolipoamide acetyltransferase
component (E2) of pyruvate dehydrogenase complex, which
catalyzes the oxidative decarboxylation of pyruvate to acetyl-CoA
(EC:2.3.1.12); K00627 pyruvate dehydrogenase E2 component
(dihydrolipoamide acetyltransferase) [EC:2.3.1.12]
Length=482
Score = 32.7 bits (73), Expect = 0.33, Method: Composition-based stats.
Identities = 16/52 (30%), Positives = 27/52 (51%), Gaps = 2/52 (3%)
Query 78 SQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVE 129
S P H I+ + + + L W KKEGD + E + E+++DKA ++
Sbjct 30 SYPEHTII--GMPALSPTMTQGNLAAWTKKEGDQLSPGEVIAEIETDKAQMD 79
> eco:b0115 aceF, ECK0114, JW0111; pyruvate dehydrogenase, dihydrolipoyltransacetylase
component E2 (EC:2.3.1.12); K00627
pyruvate dehydrogenase E2 component (dihydrolipoamide acetyltransferase)
[EC:2.3.1.12]
Length=630
Score = 32.3 bits (72), Expect = 0.49, Method: Composition-based stats.
Identities = 18/50 (36%), Positives = 29/50 (58%), Gaps = 2/50 (4%)
Query 88 KLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
K+ DIG VE+T+ K GD +E + + V+ DKA++E+ S G+
Sbjct 6 KVPDIGAD--EVEITEILVKVGDKVEAEQSLITVEGDKASMEVPSPQAGI 53
Score = 30.0 bits (66), Expect = 2.3, Method: Composition-based stats.
Identities = 16/46 (34%), Positives = 27/46 (58%), Gaps = 2/46 (4%)
Query 91 DIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
DIG VE+T+ K GD +E + + V+ DKA++E+ + + G
Sbjct 112 DIGSD--EVEVTEILVKVGDKVEAEQSLITVEGDKASMEVPAPFAG 155
Score = 28.9 bits (63), Expect = 5.4, Method: Composition-based stats.
Identities = 17/55 (30%), Positives = 30/55 (54%), Gaps = 2/55 (3%)
Query 83 GIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
G+ + DIG VE+T+ K GD + + + V+ DKA++E+ + + GV
Sbjct 205 GVKEVNVPDIGGD--EVEVTEVMVKVGDKVAAEQSLITVEGDKASMEVPAPFAGV 257
> tgo:TGME49_006610 biotin requiring domain-containing protein
/ 2-oxo acid dehydrogenases acyltransferase catalytic domain-containing
protein (EC:6.4.1.4 2.3.1.12 2.3.1.168); K00627
pyruvate dehydrogenase E2 component (dihydrolipoamide acetyltransferase)
[EC:2.3.1.12]
Length=932
Score = 32.3 bits (72), Expect = 0.54, Method: Compositional matrix adjust.
Identities = 13/54 (24%), Positives = 29/54 (53%), Gaps = 0/54 (0%)
Query 84 IVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+ + + + +T W KKEG+ + + + + V+SDKA +++ + + GV
Sbjct 240 VTDLLMPSLSPSLKTARMTVWRKKEGEKVNKGDVLFVVESDKADMDVEAPHDGV 293
> sce:YDR148C KGD2; Dihydrolipoyl transsuccinylase, component
of the mitochondrial alpha-ketoglutarate dehydrogenase complex,
which catalyzes the oxidative decarboxylation of alpha-ketoglutarate
to succinyl-CoA in the TCA cycle; phosphorylated
(EC:2.3.1.61); K00658 2-oxoglutarate dehydrogenase E2 component
(dihydrolipoamide succinyltransferase) [EC:2.3.1.61]
Length=463
Score = 32.0 bits (71), Expect = 0.54, Method: Composition-based stats.
Identities = 19/70 (27%), Positives = 33/70 (47%), Gaps = 0/70 (0%)
Query 67 KTPSARGIFSISQPRHGIVCFKLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKA 126
K A FSI+ R ++ + E + L ++ K GD I+E E + +++DK
Sbjct 56 KISVAANPFSITSNRFKSTSIEVPPMAESLTEGSLKEYTKNVGDFIKEDELLATIETDKI 115
Query 127 AVEITSRYPG 136
+E+ S G
Sbjct 116 DIEVNSPVSG 125
> ath:AT4G26910 2-oxoacid dehydrogenase family protein (EC:2.3.1.61);
K00658 2-oxoglutarate dehydrogenase E2 component (dihydrolipoamide
succinyltransferase) [EC:2.3.1.61]
Length=464
Score = 32.0 bits (71), Expect = 0.63, Method: Composition-based stats.
Identities = 14/46 (30%), Positives = 26/46 (56%), Gaps = 0/46 (0%)
Query 92 IGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
+GE I L + KK G+ ++ E + ++++DK ++I S GV
Sbjct 100 MGESITDGTLATFLKKPGERVQADEAIAQIETDKVTIDIASPASGV 145
> ath:AT3G25860 LTA2; LTA2; dihydrolipoyllysine-residue acetyltransferase
Length=480
Score = 31.2 bits (69), Expect = 0.92, Method: Composition-based stats.
Identities = 13/37 (35%), Positives = 23/37 (62%), Gaps = 0/37 (0%)
Query 100 ELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
++ W K EG+ + + E V V+SDKA +++ + Y G
Sbjct 71 KIVSWIKTEGEKLAKGESVVVVESDKADMDVETFYDG 107
> pfa:PF10_0407 dihydrolipoamide acyltransferase, putative (EC:2.3.1.12);
K00627 pyruvate dehydrogenase E2 component (dihydrolipoamide
acetyltransferase) [EC:2.3.1.12]
Length=640
Score = 30.4 bits (67), Expect = 1.7, Method: Composition-based stats.
Identities = 11/34 (32%), Positives = 21/34 (61%), Gaps = 0/34 (0%)
Query 104 WYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPGV 137
W K E D +++ + + V+ DK+ +E+ S Y G+
Sbjct 202 WLKNENDFVKKNDLLLYVEDDKSTIEVESPYSGI 235
> dre:552924 slc2a15b, zgc:113085; solute carrier family 2 (facilitated
glucose transporter), member 15b
Length=522
Score = 29.6 bits (65), Expect = 2.8, Method: Composition-based stats.
Identities = 13/36 (36%), Positives = 21/36 (58%), Gaps = 0/36 (0%)
Query 97 AAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITS 132
A + KWY+ +G+ E+EE+ E Q ++VE S
Sbjct 232 ATITALKWYRAKGNIQAEVEEMQEEQRSLSSVETVS 267
> tpv:TP03_0320 hypothetical protein
Length=267
Score = 29.3 bits (64), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 13/49 (26%), Positives = 26/49 (53%), Gaps = 0/49 (0%)
Query 88 KLADIGEGIAAVELTKWYKKEGDTIEEMEEVCEVQSDKAAVEITSRYPG 136
K+ +G I ++ KW KK GD + + +C +++D ++ S+ G
Sbjct 102 KVPALGRNITRCKIYKWLKKPGDYVNLNDLLCIIETDLIFGKVYSKVTG 150
> mmu:60425 Doc2g, D830013O18Rik; double C2, gamma
Length=387
Score = 28.9 bits (63), Expect = 4.6, Method: Compositional matrix adjust.
Identities = 18/47 (38%), Positives = 22/47 (46%), Gaps = 6/47 (12%)
Query 31 GRVAAATSYSSNAGGL------CTFLAPVAVAGVRVPRQGLFKTPSA 71
GR+ + YSS GGL C LAP+ G P LF PS+
Sbjct 245 GRILLSLCYSSERGGLLVGVLRCVHLAPMDANGYSDPFVRLFLHPSS 291
> tgo:TGME49_057580 hypothetical protein
Length=613
Score = 28.5 bits (62), Expect = 7.7, Method: Compositional matrix adjust.
Identities = 12/27 (44%), Positives = 18/27 (66%), Gaps = 0/27 (0%)
Query 30 MGRVAAATSYSSNAGGLCTFLAPVAVA 56
+GR AAT++ G +CTF APV ++
Sbjct 338 VGRNVAATAWREREGRMCTFNAPVPLS 364
Lambda K H
0.322 0.135 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2552834388
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40