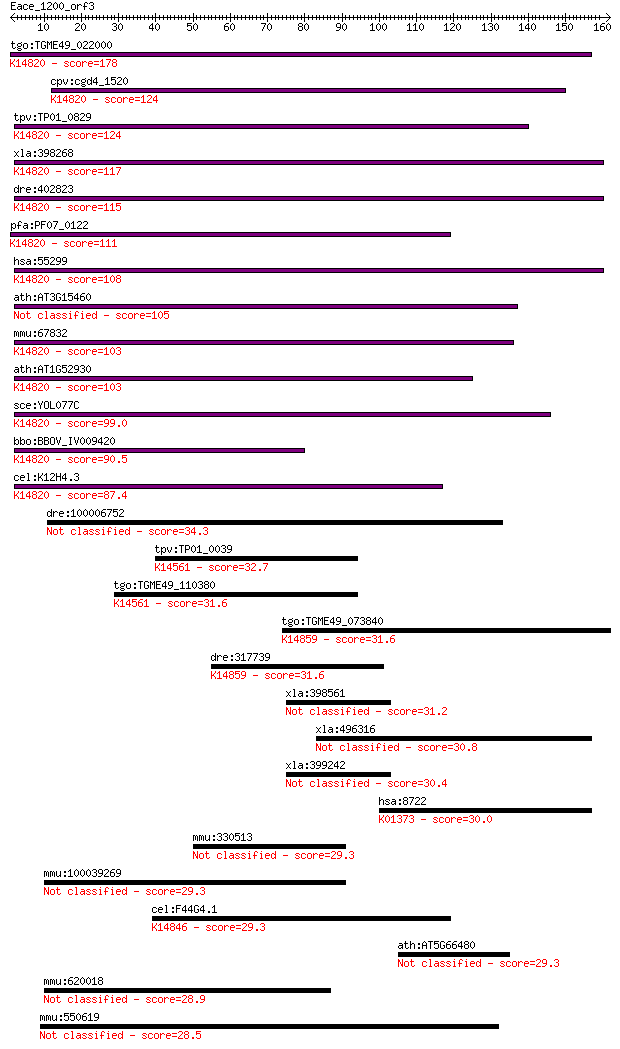
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
Query= Eace_1200_orf3
Length=161
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_022000 ribosome biogenesis protein Brix, putative ;... 178 7e-45
cpv:cgd4_1520 Brx1p nucleolar protein required for biogenesis ... 124 2e-28
tpv:TP01_0829 hypothetical protein; K14820 ribosome biogenesis... 124 2e-28
xla:398268 brix1, brix, bxdc2; BRX1, biogenesis of ribosomes, ... 117 2e-26
dre:402823 bxdc2; brix domain containing 2; K14820 ribosome bi... 115 7e-26
pfa:PF07_0122 nucleolus BRIX protein, putative; K14820 ribosom... 111 1e-24
hsa:55299 BRIX1, BRIX, BXDC2, FLJ11100; BRX1, biogenesis of ri... 108 1e-23
ath:AT3G15460 brix domain-containing protein 105 7e-23
mmu:67832 Brix1, 1110064N10Rik, Bxdc2, C76935; BRX1, biogenesi... 103 2e-22
ath:AT1G52930 brix domain-containing protein; K14820 ribosome ... 103 2e-22
sce:YOL077C BRX1; Brx1p; K14820 ribosome biogenesis protein BRX1 99.0 6e-21
bbo:BBOV_IV009420 23.m05729; ribosome biogenesis Brix protein;... 90.5 2e-18
cel:K12H4.3 hypothetical protein; K14820 ribosome biogenesis p... 87.4 2e-17
dre:100006752 hypothetical protein LOC100006752 34.3 0.18
tpv:TP01_0039 U3 small nuclear ribonucleoprotein; K14561 U3 sm... 32.7 0.55
tgo:TGME49_110380 U3 small nucleolar ribonucleoprotein protein... 31.6 1.1
tgo:TGME49_073840 ribosome biogenesis protein SSF1, putative ;... 31.6 1.2
dre:317739 ppan, cb642; peter pan homolog (Drosophila); K14859... 31.6 1.2
xla:398561 ppan-a; peter pan homolog 31.2 1.4
xla:496316 hypothetical LOC496316 30.8 1.8
xla:399242 ppan-b, MGC81614, ppan; peter pan homolog 30.4 2.6
hsa:8722 CTSF, CATSF; cathepsin F (EC:3.4.22.41); K01373 cathe... 30.0 3.1
mmu:330513 Gm5114, 4921507A12, EG330513; predicted gene 5114 29.3
mmu:100039269 Gm2128; predicted gene 2128 29.3
cel:F44G4.1 hypothetical protein; K14846 ribosome production f... 29.3 5.9
ath:AT5G66480 hypothetical protein 29.3 6.0
mmu:620018 Gm6124; predicted gene 6124 28.9
mmu:550619 Arid3c, OTTMUSG00000006683; AT rich interactive dom... 28.5 8.7
> tgo:TGME49_022000 ribosome biogenesis protein Brix, putative
; K14820 ribosome biogenesis protein BRX1
Length=517
Score = 178 bits (451), Expect = 7e-45, Method: Compositional matrix adjust.
Identities = 90/163 (55%), Positives = 117/163 (71%), Gaps = 10/163 (6%)
Query 1 IFGVEFDRE-----PHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTD-NRIWFRHY 54
+F EF E PH ++KE+F+QVFGTPRNHP++KPFFDHA++F+ D NRIWFRHY
Sbjct 338 LFSPEFGSEHGPAQPHLALIKEVFVQVFGTPRNHPKAKPFFDHALAFYKFDGNRIWFRHY 397
Query 55 QIAPLVMGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRP 114
QIAPL+ GEGGDA+ P QTFIEIGPR VL+ VKIL+GSF GKT+W N ++ +R+L+
Sbjct 398 QIAPLIGGEGGDADTPERQTFIEIGPRCVLEIVKILDGSFSGKTIWSNRNYICSRDLVAL 457
Query 115 RRLREALDFARRQGQKEKRRLYLESM-LESEPRNKLATEEVFG 156
+R+ A +A+R KEKR L+ + +E P LA E VFG
Sbjct 458 QRMGRAQSYAQRVQAKEKRTERLDKLHIEESP---LAMENVFG 497
> cpv:cgd4_1520 Brx1p nucleolar protein required for biogenesis
of the 60S ribosomal subunit ; K14820 ribosome biogenesis
protein BRX1
Length=353
Score = 124 bits (310), Expect = 2e-28, Method: Compositional matrix adjust.
Identities = 64/141 (45%), Positives = 88/141 (62%), Gaps = 6/141 (4%)
Query 12 FRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVMGEGGDANNPH 71
+++K+LF+QVFGTPR HP+SKPF DH +SF D +IWFRHYQIAP D NN
Sbjct 202 LKLIKDLFIQVFGTPRYHPKSKPFHDHVISFNFLDGKIWFRHYQIAPTT---ERDHNNVD 258
Query 72 GQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREALDFARRQGQKE 131
Q IEIGPRFVL+P+ IL+G+F G+ L+RN ++ L + + + +R K+
Sbjct 259 KQVLIEIGPRFVLEPILILDGTFSGQVLYRNPDYKSPTLLRNIIKQGYSSSYLQRVQSKQ 318
Query 132 KRRLY---LESMLESEPRNKL 149
KR Y L+S L + P K+
Sbjct 319 KRSDYKDDLDSSLMNVPEKKM 339
> tpv:TP01_0829 hypothetical protein; K14820 ribosome biogenesis
protein BRX1
Length=294
Score = 124 bits (310), Expect = 2e-28, Method: Compositional matrix adjust.
Identities = 62/138 (44%), Positives = 84/138 (60%), Gaps = 5/138 (3%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F F EP+ +++++L Q+FGTP HP+S PF DH +SFF DNRI RHYQI P
Sbjct 156 FDKSFSDEPYLKLVQKLISQIFGTPNQHPKSMPFHDHCLSFFYLDNRIHLRHYQITP--- 212
Query 62 GEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREAL 121
N+P Q +EIGP+FVL+P+ IL+GSF G TLWRN++++ LM R R+
Sbjct 213 KNEFSVNSPDNQLLVEIGPQFVLEPILILDGSFSGITLWRNSKYISP--LMVKRYERDMK 270
Query 122 DFARRQGQKEKRRLYLES 139
RR K+K L + S
Sbjct 271 LKKRRISNKKKVILTIIS 288
> xla:398268 brix1, brix, bxdc2; BRX1, biogenesis of ribosomes,
homolog; K14820 ribosome biogenesis protein BRX1
Length=339
Score = 117 bits (293), Expect = 2e-26, Method: Compositional matrix adjust.
Identities = 70/158 (44%), Positives = 87/158 (55%), Gaps = 14/158 (8%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F FDREP + +LKEL Q+FGTPR HPRS+PF DH +F + DNRIWFR+YQI
Sbjct 161 FDPAFDREPQYALLKELLTQIFGTPRYHPRSQPFVDHIFTFSIADNRIWFRNYQII---- 216
Query 62 GEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREAL 121
DA +EIGPRFVL+ +KI +GSF G TL+ N + R RL A
Sbjct 217 --EEDA------ALVEIGPRFVLNLIKIFKGSFGGPTLYENPHYQSPNMHRRMIRLATAA 268
Query 122 DFARRQGQKEKRRLYLESMLESEPRNKLATEEVFGTDA 159
+Q KE ++L E P + TE VF T A
Sbjct 269 KVKEKQQVKEVQKLKKEEDRPVIPVD--PTEAVFYTPA 304
> dre:402823 bxdc2; brix domain containing 2; K14820 ribosome
biogenesis protein BRX1
Length=383
Score = 115 bits (288), Expect = 7e-26, Method: Compositional matrix adjust.
Identities = 69/159 (43%), Positives = 89/159 (55%), Gaps = 16/159 (10%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F FD EP + +LKELF QVF TP +HP+S+PF DH +SF + D+RIWFR+YQI
Sbjct 204 FDPAFDSEPQYALLKELFTQVFSTPLHHPKSQPFVDHVISFSIADHRIWFRNYQIV---- 259
Query 62 GEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREAL 121
DA + +EIGPRFVL+P++I +GSF G TL+ N F R RL A
Sbjct 260 --EEDA------SLVEIGPRFVLNPIRIFQGSFGGPTLYENPHFQSPNAHRRMLRLAAAA 311
Query 122 DFARRQGQKEKR-RLYLESMLESEPRNKLATEEVFGTDA 159
RQ K+ R +E + +P TE VF T A
Sbjct 312 RQKERQMVKQVRMEKRMEECVLLQPD---LTEAVFNTPA 347
> pfa:PF07_0122 nucleolus BRIX protein, putative; K14820 ribosome
biogenesis protein BRX1
Length=416
Score = 111 bits (277), Expect = 1e-24, Method: Compositional matrix adjust.
Identities = 49/118 (41%), Positives = 76/118 (64%), Gaps = 6/118 (5%)
Query 1 IFGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLV 60
IF FD H +++KE+F+ VFG P HP SKPF+DH +FF ++ I+FRHYQI P+
Sbjct 270 IFSKHFDELDHLKLIKEMFIHVFGVPNYHPLSKPFYDHCYNFFYVNDLIYFRHYQILPVT 329
Query 61 MGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLR 118
+ D+NN + Q +EIGP+F L +KI E F G+ L+ N ++ +N + P++++
Sbjct 330 L---ADSNNVNKQQLVEIGPQFTLHIIKIFEECFKGRILYENEKY---QNYVTPQQVK 381
> hsa:55299 BRIX1, BRIX, BXDC2, FLJ11100; BRX1, biogenesis of
ribosomes, homolog (S. cerevisiae); K14820 ribosome biogenesis
protein BRX1
Length=353
Score = 108 bits (269), Expect = 1e-23, Method: Compositional matrix adjust.
Identities = 65/161 (40%), Positives = 93/161 (57%), Gaps = 20/161 (12%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F FD PH+ +LKEL +Q+F TPR HP+S+PF DH +F + DNRIWFR++QI
Sbjct 168 FDPAFDELPHYALLKELLIQIFSTPRYHPKSQPFVDHVFTFTILDNRIWFRNFQII---- 223
Query 62 GEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREAL 121
DA +EIGPRFVL+ +KI +GSF G TL+ N + ++ N+ R R +R
Sbjct 224 --EEDA------ALVEIGPRFVLNLIKIFQGSFGGPTLYENPHY-QSPNMHR-RVIRSIT 273
Query 122 DFARRQGQKEKRRLYLESMLESEPRNKL---ATEEVFGTDA 159
R+ Q+ K ++ + + EP+ L T +VF T A
Sbjct 274 AAKYREKQQVKD---VQKLRKKEPKTLLPHDPTADVFVTPA 311
> ath:AT3G15460 brix domain-containing protein
Length=315
Score = 105 bits (262), Expect = 7e-23, Method: Compositional matrix adjust.
Identities = 50/136 (36%), Positives = 79/136 (58%), Gaps = 1/136 (0%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIA-PLV 60
F F+ + H+++LKE+ Q+FG P H +SKP+ DH F + D+ IWFR+YQI+ P
Sbjct 162 FSSNFENDAHWKLLKEMLTQIFGIPEGHRKSKPYHDHVFVFSIVDDHIWFRNYQISVPHN 221
Query 61 MGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREA 120
+ + T IE+GPRF L+P+KI GSF G TL+ N +V + + +A
Sbjct 222 ESDKIARGDLDKMTLIEVGPRFCLNPIKIFGGSFGGTTLYENPFYVSPNQIRALEKRNKA 281
Query 121 LDFARRQGQKEKRRLY 136
FA++ K +R+++
Sbjct 282 GKFAKKIKAKTRRKMH 297
> mmu:67832 Brix1, 1110064N10Rik, Bxdc2, C76935; BRX1, biogenesis
of ribosomes, homolog (S. cerevisiae); K14820 ribosome biogenesis
protein BRX1
Length=353
Score = 103 bits (258), Expect = 2e-22, Method: Compositional matrix adjust.
Identities = 56/134 (41%), Positives = 76/134 (56%), Gaps = 12/134 (8%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F FD PH+ +LKE +Q+F TPR HP+S+PF DH +F + DNRIWFR++QI
Sbjct 168 FDPAFDDLPHYALLKEFLIQIFSTPRYHPKSQPFVDHVFTFTILDNRIWFRNFQII---- 223
Query 62 GEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREAL 121
DA +EIGPRFVL+ +KI +GSF G TL+ N + R R A
Sbjct 224 --EEDA------ALVEIGPRFVLNLIKIFQGSFGGPTLYENPHYQSPNMHRRVIRSITAA 275
Query 122 DFARRQGQKEKRRL 135
+ RQ K+ ++L
Sbjct 276 KYRERQQVKDVQKL 289
> ath:AT1G52930 brix domain-containing protein; K14820 ribosome
biogenesis protein BRX1
Length=320
Score = 103 bits (258), Expect = 2e-22, Method: Compositional matrix adjust.
Identities = 50/124 (40%), Positives = 71/124 (57%), Gaps = 1/124 (0%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIA-PLV 60
F FD++ H+++LKE+ QVFG P+ H +SKP+ DH F + D IWFR+YQI+ P
Sbjct 166 FSSNFDKDAHWKLLKEMLTQVFGIPKEHRKSKPYHDHVFVFSIVDEHIWFRNYQISVPHN 225
Query 61 MGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREA 120
+ T IE+GPRF L+P+KI GSF G TL+ N +V + + +A
Sbjct 226 ESDKIAKGGLDKMTLIEVGPRFCLNPIKIFAGSFGGPTLYENPLYVSPNQIRALEKRNKA 285
Query 121 LDFA 124
FA
Sbjct 286 GKFA 289
> sce:YOL077C BRX1; Brx1p; K14820 ribosome biogenesis protein
BRX1
Length=291
Score = 99.0 bits (245), Expect = 6e-21, Method: Compositional matrix adjust.
Identities = 52/149 (34%), Positives = 88/149 (59%), Gaps = 9/149 (6%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F F+ PH++++KEL + F P N +SKPF DH MSF + D++IW R Y+I+
Sbjct 139 FDQRFESSPHYQLIKELLVHNFCVPPNARKSKPFIDHVMSFSIVDDKIWVRTYEISHSTK 198
Query 62 GEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREAL 121
+ + + +EIGPRFV+ + ILEGSF G ++ N ++V + N++R + ++A
Sbjct 199 NKEEYEDGEEDISLVEIGPRFVMTVILILEGSFGGPKIYENKQYV-SPNVVRAQIKQQAA 257
Query 122 DFARRQGQ-----KEKRRLYLESMLESEP 145
+ A+ + + K KRR E++L ++P
Sbjct 258 EEAKSRAEAAVERKIKRR---ENVLAADP 283
> bbo:BBOV_IV009420 23.m05729; ribosome biogenesis Brix protein;
K14820 ribosome biogenesis protein BRX1
Length=281
Score = 90.5 bits (223), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 42/78 (53%), Positives = 52/78 (66%), Gaps = 3/78 (3%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVM 61
F FD PH +++K L QVFGTP NHP+SKPF DH +SFF D RI+FRH+QI+P+
Sbjct 201 FDAGFDTAPHLQLIKSLISQVFGTPNNHPKSKPFHDHCLSFFYFDGRIFFRHFQISPI-- 258
Query 62 GEGGDANNPHGQTFIEIG 79
E G N P Q+ EIG
Sbjct 259 NEYG-INKPEQQSLTEIG 275
> cel:K12H4.3 hypothetical protein; K14820 ribosome biogenesis
protein BRX1
Length=352
Score = 87.4 bits (215), Expect = 2e-17, Method: Compositional matrix adjust.
Identities = 43/116 (37%), Positives = 65/116 (56%), Gaps = 13/116 (11%)
Query 2 FGVEFDREPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTD-NRIWFRHYQIAPLV 60
F FD++P +++K + MQ GTP +HPRS+PF DH +F V + ++IWFR++QI
Sbjct 171 FDDAFDKKPQLKLIKAVLMQTLGTPHHHPRSQPFVDHVFNFSVGEGDKIWFRNFQIVDES 230
Query 61 MGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRR 116
+ E+GPRFVL+ V++ GSF G L+ N +V + R R
Sbjct 231 L------------QLQEVGPRFVLEMVRLFAGSFEGAVLYDNPNYVSPNVIRREHR 274
> dre:100006752 hypothetical protein LOC100006752
Length=400
Score = 34.3 bits (77), Expect = 0.18, Method: Compositional matrix adjust.
Identities = 34/149 (22%), Positives = 56/149 (37%), Gaps = 30/149 (20%)
Query 11 HFRVLKELFMQVFGTPRNHPRSKPF-----FDHAMSFFVTDNR------IWFRHYQIAPL 59
H E+++ V PRN S + + ++D+ WF Q + +
Sbjct 218 HTLTSPEVYLNVTYPPRNVSVSTNYSGQLWLNSVSLMCISDSNPPALSFSWFEENQSSAV 277
Query 60 VMGEG----------GDANNPHGQ------TFIEIGPRFVLDPVKILEGSFCGKTLWRNA 103
G+ +A+NPHG T GPR + I G CG ++
Sbjct 278 GSGQSFSAVQSGRFYCEAHNPHGAQRSDAVTVTVKGPRELF---YICFGVVCGAAVFITM 334
Query 104 EFVRTRNLMRPRRLREALDFARRQGQKEK 132
+ M+ R+ ++ LD +RRQ K K
Sbjct 335 GIIWGIRRMKERKAKQQLDCSRRQDDKRK 363
> tpv:TP01_0039 U3 small nuclear ribonucleoprotein; K14561 U3
small nucleolar ribonucleoprotein protein IMP4
Length=302
Score = 32.7 bits (73), Expect = 0.55, Method: Compositional matrix adjust.
Identities = 21/54 (38%), Positives = 25/54 (46%), Gaps = 5/54 (9%)
Query 40 MSFFVTDNRIWFRHYQIAPLVMGEGGDANNPHGQTFIEIGPRFVLDPVKILEGS 93
MSF ++I FRH+ V E EIGPRFVL P KI G+
Sbjct 229 MSFINEKDKIIFRHH-----VWTEKKIGAEDKKVALKEIGPRFVLKPFKIELGT 277
> tgo:TGME49_110380 U3 small nucleolar ribonucleoprotein protein,
putative (EC:3.1.2.15); K14561 U3 small nucleolar ribonucleoprotein
protein IMP4
Length=314
Score = 31.6 bits (70), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 20/81 (24%), Positives = 30/81 (37%), Gaps = 16/81 (19%)
Query 29 HPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVMGEGGDANNPHGQT-------------- 74
+P + P +F + I FRHY G +N P +
Sbjct 209 YPPANPIAQRVQAFVNVGDEIHFRHYTWTDNRKKTSGASNTPGSSSADAEEEESSEKKGH 268
Query 75 --FIEIGPRFVLDPVKILEGS 93
+E+GPRF L P +I G+
Sbjct 269 LELVEVGPRFTLKPYRIDLGT 289
> tgo:TGME49_073840 ribosome biogenesis protein SSF1, putative
; K14859 ribosome biogenesis protein SSF1/2
Length=673
Score = 31.6 bits (70), Expect = 1.2, Method: Compositional matrix adjust.
Identities = 28/101 (27%), Positives = 48/101 (47%), Gaps = 15/101 (14%)
Query 74 TFIEIGPRFVLDPVKILEGSFCGKTLWRNAEFV-RTRNLMRPRRLREALDFARRQGQKEK 132
+E+GPR L VKI E G+ L+ + F+ +T +R + +EA A RQ Q+E+
Sbjct 410 ALVELGPRMELQLVKIEEDVCAGRVLYHH--FISKTPEELRMLKAKEAAVKAERQRQREE 467
Query 133 RRLYLESMLESEPRNK------------LATEEVFGTDAMK 161
+E ++ + R++ E+V GT +K
Sbjct 468 MERAVELQMKKKRRDREDDSDGSSAASGSGDEQVSGTKGVK 508
> dre:317739 ppan, cb642; peter pan homolog (Drosophila); K14859
ribosome biogenesis protein SSF1/2
Length=522
Score = 31.6 bits (70), Expect = 1.2, Method: Compositional matrix adjust.
Identities = 16/46 (34%), Positives = 22/46 (47%), Gaps = 0/46 (0%)
Query 55 QIAPLVMGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLW 100
++ + G G A+ EIGPR L VKI+EG G L+
Sbjct 251 ELPQVYSGRGNMASQQSAVRLTEIGPRMTLQLVKIVEGMGEGSVLY 296
> xla:398561 ppan-a; peter pan homolog
Length=564
Score = 31.2 bits (69), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 15/28 (53%), Positives = 16/28 (57%), Gaps = 0/28 (0%)
Query 75 FIEIGPRFVLDPVKILEGSFCGKTLWRN 102
EIGPR L VKI EG GK L+ N
Sbjct 271 LTEIGPRMTLQLVKIEEGLSDGKVLYHN 298
> xla:496316 hypothetical LOC496316
Length=483
Score = 30.8 bits (68), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 20/78 (25%), Positives = 41/78 (52%), Gaps = 4/78 (5%)
Query 83 VLDPVKILEGSFCGKTLWRNAEFVRTRNLMRPRRLREALDFAR--RQGQKEKRRLYLE-- 138
V++ +K F G+ + ++ ++ R N +R+RE +D + G ++ Y+E
Sbjct 257 VVEKIKETLKEFYGEDVKKSPDYERIINKRHFKRIRELMDGQKIVHGGSYDEGTCYIEPT 316
Query 139 SMLESEPRNKLATEEVFG 156
M++ P +K+ EE+FG
Sbjct 317 VMVDVNPESKVMQEEIFG 334
> xla:399242 ppan-b, MGC81614, ppan; peter pan homolog
Length=600
Score = 30.4 bits (67), Expect = 2.6, Method: Compositional matrix adjust.
Identities = 15/28 (53%), Positives = 16/28 (57%), Gaps = 0/28 (0%)
Query 75 FIEIGPRFVLDPVKILEGSFCGKTLWRN 102
EIGPR L VKI EG GK L+ N
Sbjct 271 LTEIGPRMTLQLVKIEEGLSDGKVLFHN 298
> hsa:8722 CTSF, CATSF; cathepsin F (EC:3.4.22.41); K01373 cathepsin
F [EC:3.4.22.41]
Length=484
Score = 30.0 bits (66), Expect = 3.1, Method: Composition-based stats.
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 12/66 (18%)
Query 100 WRNAEFVRTRNLMRPRRLREALDFARRQ---------GQKEKRRLYLESMLESEPRNKLA 150
WR + FV N++R +++ +ALD Q ++E R +YL ++L EP NK+
Sbjct 206 WRLSVFVN--NMVRAQKI-QALDRGTAQYGVTKFSDLTEEEFRTIYLNTLLRKEPGNKMK 262
Query 151 TEEVFG 156
+ G
Sbjct 263 QAKSVG 268
> mmu:330513 Gm5114, 4921507A12, EG330513; predicted gene 5114
Length=858
Score = 29.3 bits (64), Expect = 5.1, Method: Composition-based stats.
Identities = 13/41 (31%), Positives = 19/41 (46%), Gaps = 0/41 (0%)
Query 50 WFRHYQIAPLVMGEGGDANNPHGQTFIEIGPRFVLDPVKIL 90
++ H + PL+ GE G PHG VL P K++
Sbjct 182 YYEHNNLGPLITGEFGQCLQPHGSVSYPGSQTSVLQPEKVM 222
> mmu:100039269 Gm2128; predicted gene 2128
Length=858
Score = 29.3 bits (64), Expect = 5.6, Method: Composition-based stats.
Identities = 22/81 (27%), Positives = 31/81 (38%), Gaps = 4/81 (4%)
Query 10 PHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVMGEGGDANN 69
P L F+QV TP P + A S+ ++ H + PL+ GE G
Sbjct 146 PFHPTLSASFVQV--TPSQMPNQG--YSLAPSYQEGSQVYYYEHNNLGPLIAGEFGQCLQ 201
Query 70 PHGQTFIEIGPRFVLDPVKIL 90
PHG VL P ++
Sbjct 202 PHGSVSYPGSQTAVLQPEMVM 222
> cel:F44G4.1 hypothetical protein; K14846 ribosome production
factor 1
Length=384
Score = 29.3 bits (64), Expect = 5.9, Method: Compositional matrix adjust.
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 10/80 (12%)
Query 39 AMSFFVTDNRIWFRHYQIAPLVMGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKT 98
++F + I+FRH++ EG A +E+GPRF L + +G+F K
Sbjct 312 VVTFHNQRDYIFFRHHRYE--FKKEGSKA------ALLELGPRFTLRLKWLQKGTFDAK- 362
Query 99 LWRNAEFVRTRNLMRPRRLR 118
W E+V R+ M R R
Sbjct 363 -WGEFEWVLKRHEMETSRRR 381
> ath:AT5G66480 hypothetical protein
Length=444
Score = 29.3 bits (64), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 14/30 (46%), Positives = 19/30 (63%), Gaps = 0/30 (0%)
Query 105 FVRTRNLMRPRRLREALDFARRQGQKEKRR 134
+ T N +R +EA++ ARR GQK KRR
Sbjct 372 YYETVNQEMSQRNQEAIEVARRDGQKRKRR 401
> mmu:620018 Gm6124; predicted gene 6124
Length=860
Score = 28.9 bits (63), Expect = 8.1, Method: Composition-based stats.
Identities = 21/77 (27%), Positives = 31/77 (40%), Gaps = 4/77 (5%)
Query 10 PHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFFVTDNRIWFRHYQIAPLVMGEGGDANN 69
P + L F+Q+ TP P + + A S+ ++ H + PL+ GE G
Sbjct 146 PFYPTLPASFVQI--TPPQMPSQE--YSLAPSYQEGSQVYYYEHNILGPLIAGEFGQCLQ 201
Query 70 PHGQTFIEIGPRFVLDP 86
PHG VL P
Sbjct 202 PHGSVSYPGSQTSVLQP 218
> mmu:550619 Arid3c, OTTMUSG00000006683; AT rich interactive domain
3C (BRIGHT-like)
Length=409
Score = 28.5 bits (62), Expect = 8.7, Method: Compositional matrix adjust.
Identities = 34/136 (25%), Positives = 57/136 (41%), Gaps = 15/136 (11%)
Query 9 EPHFRVLKELFMQVFGTPRNHPRSKPFFDHAMSFF----VTDNRIWFRHYQIAPL----- 59
+PH +E F Q++ P+ K F D SF NR+ Q+ L
Sbjct 91 QPHEWTYEEQFKQLYEL-DADPKRKEFLDDLFSFMQKRGTPVNRVPIMAKQVLDLYALFR 149
Query 60 -VMGEGGDANNPHGQTFIEIGPRFVLDPVKILEGSFCGKTLWRNAEF---VRTRNLMRPR 115
V +GG + + + E+ R + P I +F +T + + TR L P
Sbjct 150 LVTAKGGLVEVINRKVWREVT-RGLSLPTTITSAAFTLRTQYMKYLYPYECETRALSSPG 208
Query 116 RLREALDFARRQGQKE 131
L+ A+D RR+G+++
Sbjct 209 ELQAAIDSNRREGRRQ 224
Lambda K H
0.326 0.142 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3702936828
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40