bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
Query= Eace_1529_orf1
Length=119
Score E
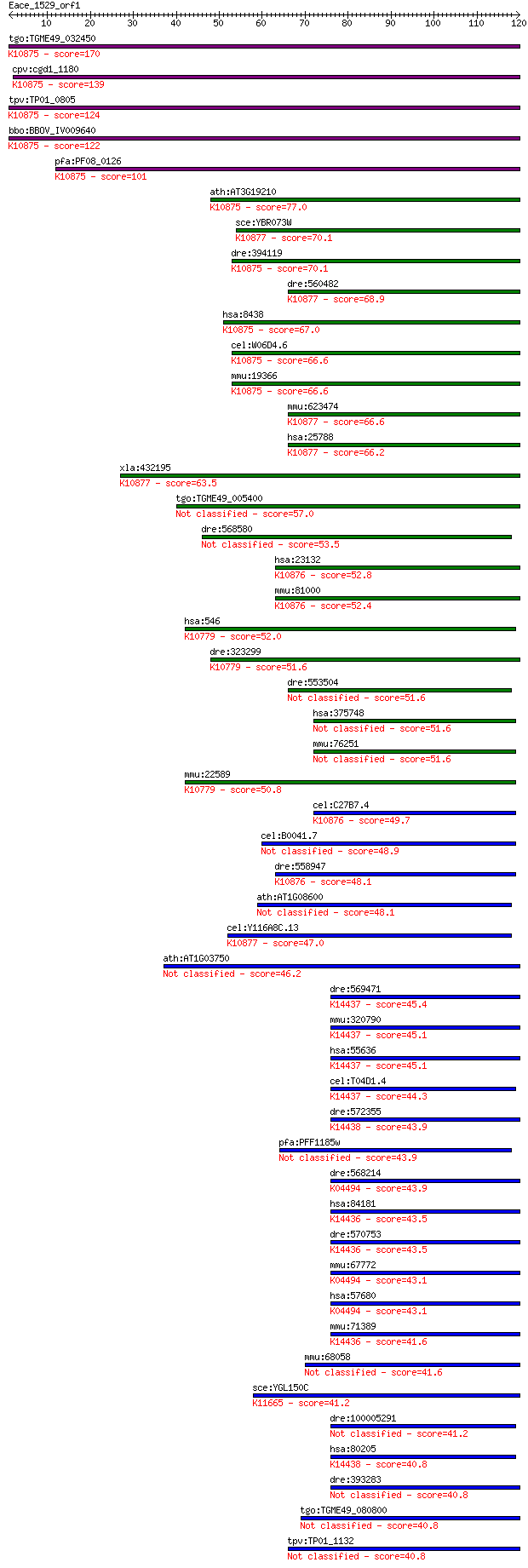
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_032450 DNA repair protein RAD54, putative (EC:2.7.1... 170 1e-42
cpv:cgd1_1180 RAD54 like SWI/SNF2 ATpase ; K10875 DNA repair a... 139 3e-33
tpv:TP01_0805 DNA repair protein Rad54; K10875 DNA repair and ... 124 8e-29
bbo:BBOV_IV009640 23.m05735; DNA repair and recombination prot... 122 2e-28
pfa:PF08_0126 DNA repair protein rad54, putative; K10875 DNA r... 101 7e-22
ath:AT3G19210 ATRAD54 (ARABIDOPSIS HOMOLOG OF RAD54); ATP bind... 77.0 1e-14
sce:YBR073W RDH54, TID1; DNA-dependent ATPase, stimulates stra... 70.1 2e-12
dre:394119 rad54l, MGC56289, zgc:56289; RAD54-like (S. cerevis... 70.1 2e-12
dre:560482 rad54b, im:7137737, im:7153525; RAD54 homolog B (S.... 68.9 3e-12
hsa:8438 RAD54L, HR54, RAD54A, hHR54, hRAD54; RAD54-like (S. c... 67.0 1e-11
cel:W06D4.6 rad-54; RADiation sensitivity abnormal/yeast RAD-r... 66.6 2e-11
mmu:19366 Rad54l, RAD54; RAD54 like (S. cerevisiae); K10875 DN... 66.6 2e-11
mmu:623474 Rad54b, E130016E03Rik, Fsbp, MGC67261; RAD54 homolo... 66.6 2e-11
hsa:25788 RAD54B, FSBP, RDH54; RAD54 homolog B (S. cerevisiae)... 66.2 3e-11
xla:432195 rad54b, MGC81308, fsbp, rdh54; RAD54 homolog B; K10... 63.5 1e-10
tgo:TGME49_005400 DNA repair protein RAD54, putative (EC:2.7.1... 57.0 1e-08
dre:568580 atrxl, wu:fb94e07; alpha thalassemia/mental retarda... 53.5 2e-07
hsa:23132 RAD54L2, ARIP4, FLJ21396, FLJ22400, HSPC325, KIAA080... 52.8 3e-07
mmu:81000 Rad54l2, Arip4, D130058C05, G630026H09Rik, Srisnf2l;... 52.4 3e-07
hsa:546 ATRX, ATR2, MGC2094, MRXHF1, RAD54, RAD54L, SFM1, SHS,... 52.0 5e-07
dre:323299 atrx, MGC66223, atrxl, wu:fb26e12, wu:fb52h08, wu:f... 51.6 5e-07
dre:553504 hypothetical protein LOC553504 51.6 6e-07
hsa:375748 C9orf102, FLJ37706, MGC30192, MGC43364, RAD26L, SR2... 51.6 6e-07
mmu:76251 0610007P08Rik, 1700019D06Rik, 9330134C04Rik, MGC1001... 51.6 6e-07
mmu:22589 Atrx, 4833408C14Rik, AI447451, ATR2, DXHXS6677E, HP1... 50.8 1e-06
cel:C27B7.4 rad-26; RADiation sensitivity abnormal/yeast RAD-r... 49.7 2e-06
cel:B0041.7 xnp-1; human XNP gene related family member (xnp-1) 48.9 4e-06
dre:558947 fc27a09, wu:fc27a09; si:dkey-42i9.10; K10876 RAD54-... 48.1 6e-06
ath:AT1G08600 ATRX; ATRX; ATP binding / DNA binding / helicase... 48.1 7e-06
cel:Y116A8C.13 hypothetical protein; K10877 DNA repair and rec... 47.0 1e-05
ath:AT1G03750 SWI2; SWI2 (SWITCH 2); ATP binding / DNA binding... 46.2 2e-05
dre:569471 chd7, KIAA0308, fd19h06, si:ch211-197o6.2, wu:cegs2... 45.4 5e-05
mmu:320790 Chd7, A730019I05Rik, Cycn, Cyn, Dz, Edy, Flo, Lda, ... 45.1 6e-05
hsa:55636 CHD7, FLJ20357, FLJ20361, IS3, KAL5, KIAA1416; chrom... 45.1 6e-05
cel:T04D1.4 tag-192; Temporarily Assigned Gene name family mem... 44.3 9e-05
dre:572355 kismet-like; K14438 chromodomain-helicase-DNA-bindi... 43.9 1e-04
pfa:PFF1185w Smarca-related protein 43.9 1e-04
dre:568214 chd8, fi45h08, si:ch211-10e2.6, wu:fi45h08; chromod... 43.9 1e-04
hsa:84181 CHD6, CHD5, KIAA1335, RIGB; chromodomain helicase DN... 43.5 2e-04
dre:570753 chd6; chromodomain helicase DNA binding protein 6; ... 43.5 2e-04
mmu:67772 Chd8, 5830451P18Rik, AU015341, Duplin, HELSNF1, mKIA... 43.1 2e-04
hsa:57680 CHD8, DKFZp686N17164, HELSNF1, KIAA1564; chromodomai... 43.1 2e-04
mmu:71389 Chd6, 5430439G14Rik, 6330406J24Rik; chromodomain hel... 41.6 5e-04
mmu:68058 Chd1l, 4432404A22Rik, Alc1, Snf2p; chromodomain heli... 41.6 7e-04
sce:YGL150C INO80; ATPase, subunit of a complex containing act... 41.2 7e-04
dre:100005291 ercc6l, fc22h03, wu:fc22h03; si:ch211-278b8.3 (E... 41.2 7e-04
hsa:80205 CHD9, AD013, CReMM, KISH2, PRIC320; chromodomain hel... 40.8 0.001
dre:393283 chd1l, MGC56084, zgc:56084; chromodomain helicase D... 40.8 0.001
tgo:TGME49_080800 SNF2 family N-terminal domain-containing pro... 40.8 0.001
tpv:TP01_1132 ATP-dependent helicase 40.8 0.001
> tgo:TGME49_032450 DNA repair protein RAD54, putative (EC:2.7.11.1);
K10875 DNA repair and recombination protein RAD54 and
RAD54-like protein [EC:3.6.4.-]
Length=873
Score = 170 bits (430), Expect = 1e-42, Method: Compositional matrix adjust.
Identities = 84/121 (69%), Positives = 96/121 (79%), Gaps = 4/121 (3%)
Query 1 KGHHPGCSEESLRKTLGIRMKRSACFLTNTKQFKSPPPLSAAAAPEVPAGEPLILY--EP 58
KGH PG SEESLRKTLG+R+KRS+ F +KQFKSPPP + A +P PL+LY P
Sbjct 87 KGHQPGVSEESLRKTLGVRLKRSSAFAVASKQFKSPPPAPTSQA--LPEANPLVLYIPPP 144
Query 59 KEVGERKVEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
E+KVEV PMLTRWLREHQR+GV+F+F+CLMGLKEF G GCILADDMGLGKTLQSIT
Sbjct 145 DRPHEKKVEVDPMLTRWLREHQRQGVKFMFDCLMGLKEFQGEGCILADDMGLGKTLQSIT 204
Query 119 I 119
I
Sbjct 205 I 205
> cpv:cgd1_1180 RAD54 like SWI/SNF2 ATpase ; K10875 DNA repair
and recombination protein RAD54 and RAD54-like protein [EC:3.6.4.-]
Length=877
Score = 139 bits (349), Expect = 3e-33, Method: Composition-based stats.
Identities = 75/129 (58%), Positives = 88/129 (68%), Gaps = 15/129 (11%)
Query 2 GHHPGCSEESLRKTLGIRMKRSACFLTNTKQ--FKSPPPLSAAAAPEVPAGEP------- 52
GH PG S ES + TLG+R+ R + + + KQ FK PP A P GEP
Sbjct 56 GHVPGVSLESRKMTLGVRIPRDSTYCNSIKQPGFKLPP----KADDSSPKGEPEPQVNVN 111
Query 53 -LILYEPKEVGERKV-EVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGL 110
LIL+ E GE++V EV MLT+WLREHQR+GV FIFECLMGL++FDG GCILADDMGL
Sbjct 112 PLILWISSEDGEKRVIEVDSMLTKWLREHQRQGVTFIFECLMGLRDFDGNGCILADDMGL 171
Query 111 GKTLQSITI 119
GKTLQSITI
Sbjct 172 GKTLQSITI 180
> tpv:TP01_0805 DNA repair protein Rad54; K10875 DNA repair and
recombination protein RAD54 and RAD54-like protein [EC:3.6.4.-]
Length=786
Score = 124 bits (311), Expect = 8e-29, Method: Composition-based stats.
Identities = 67/123 (54%), Positives = 86/123 (69%), Gaps = 7/123 (5%)
Query 1 KGHHPGCSEESLRKTLGIRMKR-SACFLTNTKQFKSPPPLSAAAAPE-VPAGEPLILYEP 58
+GH PG SEES RKTLG R+K + FL + F++ L + E +P PL+LY
Sbjct 61 EGHQPGVSEESQRKTLGCRIKSDTKSFL---RDFRAGIYLEKPESLEDLPPDNPLVLYTS 117
Query 59 KEVGERKVE--VGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQS 116
E +VE V +L+R+LR+HQR+GV+FIF+CLMGLK F+G GCILADDMGLGKTLQS
Sbjct 118 DPDAEVQVEIKVDSILSRFLRDHQRQGVQFIFDCLMGLKGFNGRGCILADDMGLGKTLQS 177
Query 117 ITI 119
IT+
Sbjct 178 ITV 180
> bbo:BBOV_IV009640 23.m05735; DNA repair and recombination protein
RAD54-like (EC:3.6.1.-); K10875 DNA repair and recombination
protein RAD54 and RAD54-like protein [EC:3.6.4.-]
Length=824
Score = 122 bits (307), Expect = 2e-28, Method: Composition-based stats.
Identities = 66/124 (53%), Positives = 84/124 (67%), Gaps = 8/124 (6%)
Query 1 KGHHPGCSEESLRKTLGIRMK---RSACFLTNTKQFKSPPPLSAAAAPEVPAGEPLILYE 57
+GH PG SE + RKTLG R++ R L + P ++ P V PL+LY
Sbjct 61 EGHLPGVSELAKRKTLGCRIRNLGRPLNLLHSPVGAAKPTEADDSSLPPV---NPLVLYT 117
Query 58 PKEVGER--KVEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQ 115
E E K+EV PML+R+LR+HQR+GV+F+F+CLMGLKEF+G GCILADDMGLGKTLQ
Sbjct 118 SPEDAEVQVKIEVDPMLSRFLRDHQRQGVQFVFDCLMGLKEFNGQGCILADDMGLGKTLQ 177
Query 116 SITI 119
SIT+
Sbjct 178 SITV 181
> pfa:PF08_0126 DNA repair protein rad54, putative; K10875 DNA
repair and recombination protein RAD54 and RAD54-like protein
[EC:3.6.4.-]
Length=1239
Score = 101 bits (251), Expect = 7e-22, Method: Composition-based stats.
Identities = 55/111 (49%), Positives = 75/111 (67%), Gaps = 6/111 (5%)
Query 12 LRKTLGIRMKRSAC--FLTNTKQFKSPPPLSAAAAPEV-PAGEPLILYEPKEVGERKVEV 68
L+KTLG++++R+ + N+ +S EV EPLILY+ + K+EV
Sbjct 69 LKKTLGVKVRRTGWINLMKNSIPRRSDETEEQEKIEEVIKKHEPLILYKDEN---DKIEV 125
Query 69 GPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
P+L ++LREHQREGV+F+FECLM +K+ GCILADDMGLGKTLQSIT+
Sbjct 126 DPILAQYLREHQREGVQFVFECLMNIKDDKISGCILADDMGLGKTLQSITV 176
> ath:AT3G19210 ATRAD54 (ARABIDOPSIS HOMOLOG OF RAD54); ATP binding
/ DNA binding / helicase/ nucleic acid binding; K10875
DNA repair and recombination protein RAD54 and RAD54-like protein
[EC:3.6.4.-]
Length=910
Score = 77.0 bits (188), Expect = 1e-14, Method: Composition-based stats.
Identities = 41/76 (53%), Positives = 54/76 (71%), Gaps = 4/76 (5%)
Query 48 PAGEPLILYEPKEVGERKVE---VGPMLTRWLREHQREGVRFIFECLMGLK-EFDGCGCI 103
P EPL+L++ +E G V V +L ++LR HQREGV+F+F+C+ GL + GCI
Sbjct 149 PDIEPLVLWQSEEDGMSNVTTIMVHSVLVKFLRPHQREGVQFMFDCVSGLHGSANINGCI 208
Query 104 LADDMGLGKTLQSITI 119
LADDMGLGKTLQSIT+
Sbjct 209 LADDMGLGKTLQSITL 224
> sce:YBR073W RDH54, TID1; DNA-dependent ATPase, stimulates strand
exchange by modifying the topology of double-stranded DNA;
involved in recombinational repair of DNA double-strand
breaks during mitosis and meiosis; proposed to be involved in
crossover interference (EC:5.99.-.- 3.6.1.-); K10877 DNA repair
and recombination protein RAD54B [EC:3.6.4.-]
Length=958
Score = 70.1 bits (170), Expect = 2e-12, Method: Composition-based stats.
Identities = 37/86 (43%), Positives = 48/86 (55%), Gaps = 20/86 (23%)
Query 54 ILYEPKEVGERKVEVGPMLTRWLREHQREGVRFIFECLMGL------------------- 94
I+ E V V P+L ++LR HQREGV+F+++CLMGL
Sbjct 274 IVMNKNAAAEVDVIVDPLLGKFLRPHQREGVKFMYDCLMGLARPTIENPDIDCTTKSLVL 333
Query 95 -KEFDGCGCILADDMGLGKTLQSITI 119
+ D GC+LADDMGLGKTL SIT+
Sbjct 334 ENDSDISGCLLADDMGLGKTLMSITL 359
> dre:394119 rad54l, MGC56289, zgc:56289; RAD54-like (S. cerevisiae);
K10875 DNA repair and recombination protein RAD54 and
RAD54-like protein [EC:3.6.4.-]
Length=738
Score = 70.1 bits (170), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 35/79 (44%), Positives = 50/79 (63%), Gaps = 12/79 (15%)
Query 53 LILYEPKEVGERK------------VEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGC 100
L+LYEP + V V P+L++ LR HQREGV+F+++C+ G + +
Sbjct 112 LVLYEPPAISAHDLIKADKEKLPVHVVVDPVLSKVLRPHQREGVKFLWDCVTGRRIENSY 171
Query 101 GCILADDMGLGKTLQSITI 119
GCI+AD+MGLGKTLQ IT+
Sbjct 172 GCIMADEMGLGKTLQCITL 190
> dre:560482 rad54b, im:7137737, im:7153525; RAD54 homolog B (S.
cerevisiae); K10877 DNA repair and recombination protein
RAD54B [EC:3.6.4.-]
Length=1174
Score = 68.9 bits (167), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 31/54 (57%), Positives = 40/54 (74%), Gaps = 0/54 (0%)
Query 66 VEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
V + P LT LR HQ+EGV F++ECLMG++ CG ILAD+MGLGKTLQ + +
Sbjct 550 VVIDPHLTNHLRPHQKEGVVFLYECLMGMRLAGRCGAILADEMGLGKTLQCVCV 603
> hsa:8438 RAD54L, HR54, RAD54A, hHR54, hRAD54; RAD54-like (S.
cerevisiae); K10875 DNA repair and recombination protein RAD54
and RAD54-like protein [EC:3.6.4.-]
Length=747
Score = 67.0 bits (162), Expect = 1e-11, Method: Compositional matrix adjust.
Identities = 37/81 (45%), Positives = 50/81 (61%), Gaps = 12/81 (14%)
Query 51 EPLILYEP------------KEVGERKVEVGPMLTRWLREHQREGVRFIFECLMGLKEFD 98
+ L+LYEP KE V V P+L++ LR HQREGV+F++EC+ +
Sbjct 116 DALVLYEPPPLSAHDQLKLDKEKLPVHVVVDPILSKVLRPHQREGVKFLWECVTSRRIPG 175
Query 99 GCGCILADDMGLGKTLQSITI 119
GCI+AD+MGLGKTLQ IT+
Sbjct 176 SHGCIMADEMGLGKTLQCITL 196
> cel:W06D4.6 rad-54; RADiation sensitivity abnormal/yeast RAD-related
family member (rad-54); K10875 DNA repair and recombination
protein RAD54 and RAD54-like protein [EC:3.6.4.-]
Length=818
Score = 66.6 bits (161), Expect = 2e-11, Method: Composition-based stats.
Identities = 36/78 (46%), Positives = 52/78 (66%), Gaps = 11/78 (14%)
Query 53 LILYEP---------KEVGERKVEV--GPMLTRWLREHQREGVRFIFECLMGLKEFDGCG 101
LILY P KE +RKV V P++ + LR HQR+GV+F+++C+ G+ + G
Sbjct 168 LILYAPEHLSEHAQLKEDKDRKVHVVADPVVGKILRPHQRDGVKFMWDCVTGINIPEFHG 227
Query 102 CILADDMGLGKTLQSITI 119
CI+AD+MGLGKTLQ I++
Sbjct 228 CIMADEMGLGKTLQCISL 245
> mmu:19366 Rad54l, RAD54; RAD54 like (S. cerevisiae); K10875
DNA repair and recombination protein RAD54 and RAD54-like protein
[EC:3.6.4.-]
Length=747
Score = 66.6 bits (161), Expect = 2e-11, Method: Compositional matrix adjust.
Identities = 37/79 (46%), Positives = 49/79 (62%), Gaps = 12/79 (15%)
Query 53 LILYEP------------KEVGERKVEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGC 100
L+LYEP KE V V P+L++ LR HQREGV+F++EC+ +
Sbjct 118 LVLYEPPPLSAHDQLKLDKEKLPVHVVVDPILSKVLRPHQREGVKFLWECVTSRRIPGSH 177
Query 101 GCILADDMGLGKTLQSITI 119
GCI+AD+MGLGKTLQ IT+
Sbjct 178 GCIMADEMGLGKTLQCITL 196
> mmu:623474 Rad54b, E130016E03Rik, Fsbp, MGC67261; RAD54 homolog
B (S. cerevisiae); K10877 DNA repair and recombination protein
RAD54B [EC:3.6.4.-]
Length=886
Score = 66.6 bits (161), Expect = 2e-11, Method: Compositional matrix adjust.
Identities = 28/54 (51%), Positives = 40/54 (74%), Gaps = 0/54 (0%)
Query 66 VEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
V + P L LR HQ++G+ F++EC+MG++ CG ILAD+MGLGKTLQ I++
Sbjct 264 VVIDPHLVHHLRPHQKDGIIFLYECVMGMRAVGKCGAILADEMGLGKTLQCISL 317
> hsa:25788 RAD54B, FSBP, RDH54; RAD54 homolog B (S. cerevisiae);
K10877 DNA repair and recombination protein RAD54B [EC:3.6.4.-]
Length=910
Score = 66.2 bits (160), Expect = 3e-11, Method: Composition-based stats.
Identities = 29/54 (53%), Positives = 40/54 (74%), Gaps = 0/54 (0%)
Query 66 VEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
V + P L LR HQ+EG+ F++EC+MG++ CG ILAD+MGLGKTLQ I++
Sbjct 286 VVIDPYLVYHLRPHQKEGIIFLYECVMGMRMNGRCGAILADEMGLGKTLQCISL 339
> xla:432195 rad54b, MGC81308, fsbp, rdh54; RAD54 homolog B; K10877
DNA repair and recombination protein RAD54B [EC:3.6.4.-]
Length=895
Score = 63.5 bits (153), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 36/93 (38%), Positives = 53/93 (56%), Gaps = 1/93 (1%)
Query 27 LTNTKQFKSPPPLSAAAAPEVPAGEPLILYEPKEVGERKVEVGPMLTRWLREHQREGVRF 86
L N K P +A P+ P+ + + + V V P L LR HQ+EG+ F
Sbjct 231 LQNIKPRHDPSAPNAFVMPK-PSQQHQWAFNKSGLPIVDVVVDPYLAVHLRPHQKEGILF 289
Query 87 IFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
++EC+MG++ + G ILAD+MGLGKTLQ I++
Sbjct 290 LYECVMGMRVNERFGAILADEMGLGKTLQCISL 322
> tgo:TGME49_005400 DNA repair protein RAD54, putative (EC:2.7.11.1)
Length=693
Score = 57.0 bits (136), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 32/81 (39%), Positives = 46/81 (56%), Gaps = 6/81 (7%)
Query 40 SAAAAPEVPAGEPLILYEPKEVGERK-VEVGPMLTRWLREHQREGVRFIFECLMGLKEFD 98
+ +A P G I GER + + L + LR HQREG+ +++ L+ +
Sbjct 208 TTSAEKACPHGAFEIAVSETATGERHPIWIDAFLNKHLRPHQREGISWMYNQLL-----E 262
Query 99 GCGCILADDMGLGKTLQSITI 119
G GCILAD MGLGKTLQ+I++
Sbjct 263 GGGCILADTMGLGKTLQAISL 283
> dre:568580 atrxl, wu:fb94e07; alpha thalassemia/mental retardation
syndrome X-linked, like
Length=1757
Score = 53.5 bits (127), Expect = 2e-07, Method: Composition-based stats.
Identities = 32/78 (41%), Positives = 44/78 (56%), Gaps = 6/78 (7%)
Query 46 EVPAGE--PLILYEPKEVGERKVEVGPMLTRWLREHQREGVRFIFEC----LMGLKEFDG 99
E PA PL+L + + V+V L++HQR+GVRF+++C + +K G
Sbjct 953 ERPANSIAPLVLEQDDATNKPVVQVHMHFLSKLKQHQRDGVRFMWDCCCESVQSVKSTPG 1012
Query 100 CGCILADDMGLGKTLQSI 117
GCILA MGLGKT Q I
Sbjct 1013 SGCILAHCMGLGKTFQVI 1030
> hsa:23132 RAD54L2, ARIP4, FLJ21396, FLJ22400, HSPC325, KIAA0809,
SRISNF2L; RAD54-like 2 (S. cerevisiae) (EC:3.6.4.12); K10876
RAD54-like protein 2 [EC:3.6.4.12]
Length=1467
Score = 52.8 bits (125), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 28/61 (45%), Positives = 35/61 (57%), Gaps = 4/61 (6%)
Query 63 ERKVEVGPMLTRWLREHQREGVRFIF----ECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
E V + P L R ++ HQ G+RF++ E L K G GCILA MGLGKTLQ I+
Sbjct 258 EENVFLAPQLARAVKPHQIGGIRFLYDNLVESLERFKTSSGFGCILAHSMGLGKTLQVIS 317
Query 119 I 119
Sbjct 318 F 318
> mmu:81000 Rad54l2, Arip4, D130058C05, G630026H09Rik, Srisnf2l;
RAD54 like 2 (S. cerevisiae) (EC:3.6.4.12); K10876 RAD54-like
protein 2 [EC:3.6.4.12]
Length=1467
Score = 52.4 bits (124), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 28/61 (45%), Positives = 35/61 (57%), Gaps = 4/61 (6%)
Query 63 ERKVEVGPMLTRWLREHQREGVRFIF----ECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
E V + P L R ++ HQ G+RF++ E L K G GCILA MGLGKTLQ I+
Sbjct 258 EENVFLAPQLARAVKPHQIGGIRFLYDNLVESLERFKTSSGFGCILAHSMGLGKTLQVIS 317
Query 119 I 119
Sbjct 318 F 318
> hsa:546 ATRX, ATR2, MGC2094, MRXHF1, RAD54, RAD54L, SFM1, SHS,
XH2, XNP, ZNF-HX; alpha thalassemia/mental retardation syndrome
X-linked (EC:3.6.4.12); K10779 transcriptional regulator
ATRX [EC:3.6.4.12]
Length=2492
Score = 52.0 bits (123), Expect = 5e-07, Method: Composition-based stats.
Identities = 31/81 (38%), Positives = 47/81 (58%), Gaps = 4/81 (4%)
Query 42 AAAPEVPAGEPLILYEPKEVGERKVEVGPMLTRWLREHQREGVRFIFEC----LMGLKEF 97
A+ + P L+L E +E E V+V + L+ HQ +GV+F+++C + K+
Sbjct 1526 ASPTKCPITTKLVLDEDEETKEPLVQVHRNMVIKLKPHQVDGVQFMWDCCCESVKKTKKS 1585
Query 98 DGCGCILADDMGLGKTLQSIT 118
G GCILA MGLGKTLQ ++
Sbjct 1586 PGSGCILAHCMGLGKTLQVVS 1606
> dre:323299 atrx, MGC66223, atrxl, wu:fb26e12, wu:fb52h08, wu:fb94e07,
zgc:66223; alpha thalassemia/mental retardation syndrome
X-linked homolog (human) (EC:3.6.4.12); K10779 transcriptional
regulator ATRX [EC:3.6.4.12]
Length=2013
Score = 51.6 bits (122), Expect = 5e-07, Method: Composition-based stats.
Identities = 30/76 (39%), Positives = 46/76 (60%), Gaps = 4/76 (5%)
Query 48 PAGEPLILYEPKEVGERKVEVGPMLTRWLREHQREGVRFIFEC----LMGLKEFDGCGCI 103
P L+L E +E E V+V + L+ HQ +GV+F+++C + +++ G GCI
Sbjct 1086 PITTKLVLDEDEETKEPLVQVHRNMVTKLKPHQVDGVQFMWDCCCESVRKVEKSAGSGCI 1145
Query 104 LADDMGLGKTLQSITI 119
LA MGLGKTLQ +T+
Sbjct 1146 LAHCMGLGKTLQVVTL 1161
> dre:553504 hypothetical protein LOC553504
Length=1269
Score = 51.6 bits (122), Expect = 6e-07, Method: Composition-based stats.
Identities = 25/52 (48%), Positives = 36/52 (69%), Gaps = 5/52 (9%)
Query 66 VEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSI 117
V+V + R+LR++QREG++FI++ + GCIL DDMGLGKT+Q I
Sbjct 50 VKVPYTINRYLRDYQREGIKFIYQNYAKSR-----GCILGDDMGLGKTVQVI 96
> hsa:375748 C9orf102, FLJ37706, MGC30192, MGC43364, RAD26L, SR278;
chromosome 9 open reading frame 102
Length=712
Score = 51.6 bits (122), Expect = 6e-07, Method: Composition-based stats.
Identities = 24/47 (51%), Positives = 33/47 (70%), Gaps = 5/47 (10%)
Query 72 LTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
+ R+LR++QREG RF++ + G GCIL DDMGLGKT+Q I+
Sbjct 130 INRYLRDYQREGTRFLYGHYI-----HGGGCILGDDMGLGKTVQVIS 171
> mmu:76251 0610007P08Rik, 1700019D06Rik, 9330134C04Rik, MGC100170,
Rad26l, Sr278; RIKEN cDNA 0610007P08 gene
Length=699
Score = 51.6 bits (122), Expect = 6e-07, Method: Composition-based stats.
Identities = 23/47 (48%), Positives = 34/47 (72%), Gaps = 5/47 (10%)
Query 72 LTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
+ R+LR++QREG +F++ + +G GCIL DDMGLGKT+Q I+
Sbjct 118 INRYLRDYQREGAQFLYRHYI-----EGRGCILGDDMGLGKTIQVIS 159
> mmu:22589 Atrx, 4833408C14Rik, AI447451, ATR2, DXHXS6677E, HP1-BP38,
Hp1bp2, Hp1bp38, MRXS3, RAD54L, Rad54, XH2, Xnp, ZNF-HX;
alpha thalassemia/mental retardation syndrome X-linked
homolog (human) (EC:3.6.4.12); K10779 transcriptional regulator
ATRX [EC:3.6.4.12]
Length=2476
Score = 50.8 bits (120), Expect = 1e-06, Method: Composition-based stats.
Identities = 31/81 (38%), Positives = 47/81 (58%), Gaps = 4/81 (4%)
Query 42 AAAPEVPAGEPLILYEPKEVGERKVEVGPMLTRWLREHQREGVRFIFEC----LMGLKEF 97
A+ + P L+L E +E E V+V + L+ HQ +GV+F+++C + K+
Sbjct 1511 ASPTKCPITTKLVLDENEETKEPLVQVHRNMVIKLKPHQVDGVQFMWDCCCESVEKTKKS 1570
Query 98 DGCGCILADDMGLGKTLQSIT 118
G GCILA MGLGKTLQ ++
Sbjct 1571 PGSGCILAHCMGLGKTLQVVS 1591
> cel:C27B7.4 rad-26; RADiation sensitivity abnormal/yeast RAD-related
family member (rad-26); K10876 RAD54-like protein 2
[EC:3.6.4.12]
Length=1274
Score = 49.7 bits (117), Expect = 2e-06, Method: Composition-based stats.
Identities = 27/51 (52%), Positives = 33/51 (64%), Gaps = 4/51 (7%)
Query 72 LTRWLREHQREGVRFIF----ECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
LT L+ HQ G+RF++ E L K+ DG GCILA MGLGKT+Q IT
Sbjct 252 LTHVLQPHQLGGIRFMYDNTIESLGEYKKSDGFGCILAHSMGLGKTIQVIT 302
> cel:B0041.7 xnp-1; human XNP gene related family member (xnp-1)
Length=1359
Score = 48.9 bits (115), Expect = 4e-06, Method: Composition-based stats.
Identities = 29/62 (46%), Positives = 37/62 (59%), Gaps = 3/62 (4%)
Query 60 EVGERKVEVGPMLTRWLREHQREGVRFIFECLM-GLKEFD--GCGCILADDMGLGKTLQS 116
E ++ VEV L R L+ HQ G++F+++C L D G G ILA MGLGKTLQ
Sbjct 447 EESKKPVEVHNSLVRILKPHQAHGIQFMYDCAFESLDRLDTEGSGGILAHCMGLGKTLQV 506
Query 117 IT 118
IT
Sbjct 507 IT 508
> dre:558947 fc27a09, wu:fc27a09; si:dkey-42i9.10; K10876 RAD54-like
protein 2 [EC:3.6.4.12]
Length=1437
Score = 48.1 bits (113), Expect = 6e-06, Method: Composition-based stats.
Identities = 27/60 (45%), Positives = 35/60 (58%), Gaps = 4/60 (6%)
Query 63 ERKVEVGPMLTRWLREHQREGVRFIF----ECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
E + + P L R ++ HQ G+RF++ E L K G GCILA MGLGKTLQ I+
Sbjct 268 EEDLFLAPQLARAVKPHQIGGIRFLYDNLVESLERYKTSSGFGCILAHSMGLGKTLQVIS 327
> ath:AT1G08600 ATRX; ATRX; ATP binding / DNA binding / helicase/
nucleic acid binding
Length=1479
Score = 48.1 bits (113), Expect = 7e-06, Method: Composition-based stats.
Identities = 28/64 (43%), Positives = 39/64 (60%), Gaps = 5/64 (7%)
Query 59 KEVGERKVEVGPMLTRWLREHQREGVRFIFECLMG----LKEFD-GCGCILADDMGLGKT 113
+E+GE V V ++ L+ HQ G+RF++E ++ +K D G GCILA MGLGKT
Sbjct 702 REIGEEAVRVPRSISAKLKVHQVTGIRFMWENIIQSISRVKSGDKGLGCILAHTMGLGKT 761
Query 114 LQSI 117
Q I
Sbjct 762 FQVI 765
> cel:Y116A8C.13 hypothetical protein; K10877 DNA repair and recombination
protein RAD54B [EC:3.6.4.-]
Length=833
Score = 47.0 bits (110), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 33/72 (45%), Positives = 44/72 (61%), Gaps = 7/72 (9%)
Query 52 PLILYEPKEVGERK----VEVGPMLTRWLREHQREGVRFIFECL--MGLKEFDGCGCILA 105
P IL E +++ +RK V V R LR HQ+ G++FIF+ L K G G ILA
Sbjct 179 PFILNE-QDILDRKTTSAVTVDTRFARHLRPHQKSGIQFIFDRLRRGSGKNGGGGGAILA 237
Query 106 DDMGLGKTLQSI 117
DDMGLGK+LQ++
Sbjct 238 DDMGLGKSLQTM 249
> ath:AT1G03750 SWI2; SWI2 (SWITCH 2); ATP binding / DNA binding
/ helicase/ nucleic acid binding
Length=862
Score = 46.2 bits (108), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 32/83 (38%), Positives = 42/83 (50%), Gaps = 7/83 (8%)
Query 37 PPLSAAAAPEVPAGEPLILYEPKEVGERKVEVGPMLTRWLREHQREGVRFIFECLMGLKE 96
P LS A EPL+L E+ + V + L EHQREGV+F++
Sbjct 102 PGLSRAEFDYSGPYEPLMLSSIGEIP--IIHVPASINCRLLEHQREGVKFMYNLYK---- 155
Query 97 FDGCGCILADDMGLGKTLQSITI 119
+ G IL DDMGLGKT+Q+I
Sbjct 156 -NNHGGILGDDMGLGKTIQTIAF 177
> dre:569471 chd7, KIAA0308, fd19h06, si:ch211-197o6.2, wu:cegs2051,
wu:fb37f10, wu:fb39h04, wu:fd19h06; chromodomain helicase
DNA binding protein 7; K14437 chromodomain-helicase-DNA-binding
protein 7 [EC:3.6.4.12]
Length=3094
Score = 45.4 bits (106), Expect = 5e-05, Method: Compositional matrix adjust.
Identities = 22/44 (50%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EGV ++ ++ CILAD+MGLGKT+QSIT
Sbjct 1012 LREYQLEGVNWLL-----FNWYNTRNCILADEMGLGKTIQSITF 1050
> mmu:320790 Chd7, A730019I05Rik, Cycn, Cyn, Dz, Edy, Flo, Lda,
Mt, Obt, Todo, WBE1, Whi; chromodomain helicase DNA binding
protein 7 (EC:3.6.4.12); K14437 chromodomain-helicase-DNA-binding
protein 7 [EC:3.6.4.12]
Length=2986
Score = 45.1 bits (105), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 22/44 (50%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EGV ++ ++ CILAD+MGLGKT+QSIT
Sbjct 958 LREYQLEGVNWLLFNWYNMR-----NCILADEMGLGKTIQSITF 996
> hsa:55636 CHD7, FLJ20357, FLJ20361, IS3, KAL5, KIAA1416; chromodomain
helicase DNA binding protein 7 (EC:3.6.4.12); K14437
chromodomain-helicase-DNA-binding protein 7 [EC:3.6.4.12]
Length=2997
Score = 45.1 bits (105), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 22/44 (50%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EGV ++ ++ CILAD+MGLGKT+QSIT
Sbjct 968 LREYQLEGVNWLLFNWYNMR-----NCILADEMGLGKTIQSITF 1006
> cel:T04D1.4 tag-192; Temporarily Assigned Gene name family member
(tag-192); K14437 chromodomain-helicase-DNA-binding protein
7 [EC:3.6.4.12]
Length=2967
Score = 44.3 bits (103), Expect = 9e-05, Method: Composition-based stats.
Identities = 22/43 (51%), Positives = 30/43 (69%), Gaps = 5/43 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
LRE+Q EGV ++ C ++ CILAD+MGLGKT+Q+IT
Sbjct 1197 LREYQFEGVDWLLYCY-----YNAQNCILADEMGLGKTVQTIT 1234
> dre:572355 kismet-like; K14438 chromodomain-helicase-DNA-binding
protein 9 [EC:3.6.4.12]
Length=2429
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 21/44 (47%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LR++Q EGV ++ ++ CILAD+MGLGKT+QSIT
Sbjct 459 LRDYQLEGVNWLL-----FNWYNRRNCILADEMGLGKTIQSITF 497
> pfa:PFF1185w Smarca-related protein
Length=2719
Score = 43.9 bits (102), Expect = 1e-04, Method: Composition-based stats.
Identities = 24/55 (43%), Positives = 35/55 (63%), Gaps = 7/55 (12%)
Query 64 RKVEVGPMLTR-WLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSI 117
RK+E P++ L+ HQ +GV ++ LK F G ILAD+MGLGKT+Q++
Sbjct 325 RKIERRPLVDNVQLKPHQEDGVEWL------LKSFLTGGAILADEMGLGKTIQTL 373
> dre:568214 chd8, fi45h08, si:ch211-10e2.6, wu:fi45h08; chromodomain
helicase DNA binding protein 8 (EC:3.6.4.12); K04494
chromodomain helicase DNA binding protein 8 [EC:3.6.4.12]
Length=2549
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 21/44 (47%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EGV ++ ++ CILAD+MGLGKT+QSI +
Sbjct 885 LREYQLEGVNWLL-----FNWYNRQNCILADEMGLGKTIQSIAL 923
> hsa:84181 CHD6, CHD5, KIAA1335, RIGB; chromodomain helicase
DNA binding protein 6 (EC:3.6.4.12); K14436 chromodomain-helicase-DNA-binding
protein 6 [EC:3.6.4.12]
Length=2715
Score = 43.5 bits (101), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 21/44 (47%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EG+ ++ ++ CILAD+MGLGKT+QSIT
Sbjct 461 LREYQLEGMNWLL-----FNWYNRKNCILADEMGLGKTIQSITF 499
> dre:570753 chd6; chromodomain helicase DNA binding protein 6;
K14436 chromodomain-helicase-DNA-binding protein 6 [EC:3.6.4.12]
Length=2699
Score = 43.5 bits (101), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 21/44 (47%), Positives = 29/44 (65%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EG+ ++ ++ CILAD+MGLGKT+QSIT
Sbjct 247 LREYQLEGMNWLL-----FNWYNRKNCILADEMGLGKTIQSITF 285
> mmu:67772 Chd8, 5830451P18Rik, AU015341, Duplin, HELSNF1, mKIAA1564;
chromodomain helicase DNA binding protein 8 (EC:3.6.4.12);
K04494 chromodomain helicase DNA binding protein 8 [EC:3.6.4.12]
Length=2582
Score = 43.1 bits (100), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 21/44 (47%), Positives = 28/44 (63%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EGV ++ ++ CILAD+MGLGKT+QSI
Sbjct 813 LREYQLEGVNWLL-----FNWYNRQNCILADEMGLGKTIQSIAF 851
> hsa:57680 CHD8, DKFZp686N17164, HELSNF1, KIAA1564; chromodomain
helicase DNA binding protein 8 (EC:3.6.4.12); K04494 chromodomain
helicase DNA binding protein 8 [EC:3.6.4.12]
Length=2581
Score = 43.1 bits (100), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 21/44 (47%), Positives = 28/44 (63%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EGV ++ ++ CILAD+MGLGKT+QSI
Sbjct 811 LREYQLEGVNWLL-----FNWYNRQNCILADEMGLGKTIQSIAF 849
> mmu:71389 Chd6, 5430439G14Rik, 6330406J24Rik; chromodomain helicase
DNA binding protein 6; K14436 chromodomain-helicase-DNA-binding
protein 6 [EC:3.6.4.12]
Length=2711
Score = 41.6 bits (96), Expect = 5e-04, Method: Compositional matrix adjust.
Identities = 20/44 (45%), Positives = 28/44 (63%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LRE+Q EG+ ++ ++ CILAD+MGLGKT+QSI
Sbjct 460 LREYQLEGMNWLL-----FNWYNRKNCILADEMGLGKTIQSIAF 498
> mmu:68058 Chd1l, 4432404A22Rik, Alc1, Snf2p; chromodomain helicase
DNA binding protein 1-like (EC:3.6.4.12)
Length=900
Score = 41.6 bits (96), Expect = 7e-04, Method: Compositional matrix adjust.
Identities = 23/56 (41%), Positives = 31/56 (55%), Gaps = 11/56 (19%)
Query 70 PMLTRW------LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
P L +W LR +Q EGV ++ +C GCIL D+MGLGKT Q+I +
Sbjct 28 PDLQQWGLTGIRLRSYQLEGVNWLVQCFHCQN-----GCILGDEMGLGKTCQTIAL 78
> sce:YGL150C INO80; ATPase, subunit of a complex containing actin
and several actin-related proteins that has chromatin remodeling
activity and 3' to 5' DNA helicase activity in vitro;
has a role in modulating stress gene transcription (EC:3.6.1.-);
K11665 DNA helicase INO80 [EC:3.6.4.12]
Length=1489
Score = 41.2 bits (95), Expect = 7e-04, Method: Composition-based stats.
Identities = 25/62 (40%), Positives = 38/62 (61%), Gaps = 5/62 (8%)
Query 58 PKEVGERKVEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSI 117
P +GE +E +L L+E+Q +G+ + L L + G ILAD+MGLGKT+QSI
Sbjct 688 PTSLGEITIEQPKILACTLKEYQLKGLNW----LANLYD-QGINGILADEMGLGKTVQSI 742
Query 118 TI 119
++
Sbjct 743 SV 744
> dre:100005291 ercc6l, fc22h03, wu:fc22h03; si:ch211-278b8.3
(EC:3.6.4.12)
Length=1451
Score = 41.2 bits (95), Expect = 7e-04, Method: Composition-based stats.
Identities = 21/43 (48%), Positives = 29/43 (67%), Gaps = 4/43 (9%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
L +HQ+EGV F++ ++ G ILADDMGLGKT+Q I+
Sbjct 105 LYDHQKEGVAFLYSLYRDGRK----GGILADDMGLGKTIQVIS 143
> hsa:80205 CHD9, AD013, CReMM, KISH2, PRIC320; chromodomain helicase
DNA binding protein 9 (EC:3.6.4.12); K14438 chromodomain-helicase-DNA-binding
protein 9 [EC:3.6.4.12]
Length=2881
Score = 40.8 bits (94), Expect = 0.001, Method: Composition-based stats.
Identities = 21/43 (48%), Positives = 29/43 (67%), Gaps = 5/43 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSIT 118
LRE+Q EG+ ++ ++ CILAD+MGLGKT+QSIT
Sbjct 860 LREYQLEGLNWLL-----FNWYNRRNCILADEMGLGKTIQSIT 897
> dre:393283 chd1l, MGC56084, zgc:56084; chromodomain helicase
DNA binding protein 1-like (EC:3.6.4.12)
Length=1026
Score = 40.8 bits (94), Expect = 0.001, Method: Composition-based stats.
Identities = 19/44 (43%), Positives = 30/44 (68%), Gaps = 5/44 (11%)
Query 76 LREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
LR +Q +GV+++ C+ + GCIL D+MGLGKT Q+I++
Sbjct 35 LRPYQLDGVKWLSLCMKNQQ-----GCILGDEMGLGKTCQTISL 73
> tgo:TGME49_080800 SNF2 family N-terminal domain-containing protein
(EC:2.7.11.1 2.7.1.127)
Length=2894
Score = 40.8 bits (94), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 24/52 (46%), Positives = 31/52 (59%), Gaps = 6/52 (11%)
Query 69 GPMLTR-WLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
P L R LR +Q EGV+++F G ILAD+MGLGKTLQ+I +
Sbjct 1213 APALVRATLRTYQSEGVQWLFAL-----HDKGLNGILADEMGLGKTLQTIVL 1259
> tpv:TP01_1132 ATP-dependent helicase
Length=1632
Score = 40.8 bits (94), Expect = 0.001, Method: Composition-based stats.
Identities = 25/54 (46%), Positives = 37/54 (68%), Gaps = 5/54 (9%)
Query 66 VEVGPMLTRWLREHQREGVRFIFECLMGLKEFDGCGCILADDMGLGKTLQSITI 119
+EV ++ LR +Q+EG+R+ L+ L E + G ILAD+MGLGKTLQ+I +
Sbjct 688 IEVPFLIKGVLRPYQKEGLRW----LVSLYERNING-ILADEMGLGKTLQTICL 736
Lambda K H
0.320 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2022937320
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40