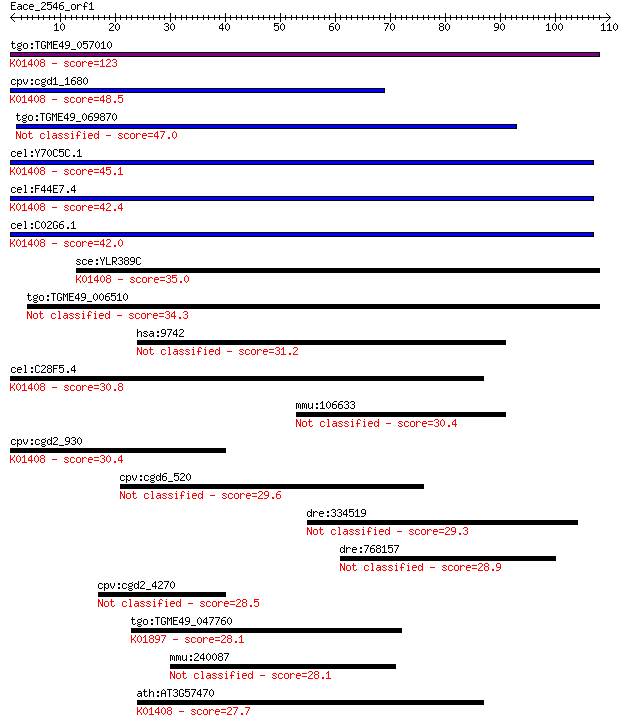
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
Query= Eace_2546_orf1
Length=109
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_057010 insulysin, putative (EC:3.4.24.56); K01408 i... 123 1e-28
cpv:cgd1_1680 insulinase like protease, signal peptide ; K0140... 48.5 5e-06
tgo:TGME49_069870 hypothetical protein 47.0 2e-05
cel:Y70C5C.1 hypothetical protein; K01408 insulysin [EC:3.4.24... 45.1 6e-05
cel:F44E7.4 hypothetical protein; K01408 insulysin [EC:3.4.24.56] 42.4 4e-04
cel:C02G6.1 hypothetical protein; K01408 insulysin [EC:3.4.24.56] 42.0 5e-04
sce:YLR389C STE23; Metalloprotease involved, with homolog Axl1... 35.0 0.062
tgo:TGME49_006510 peptidase M16 domain containing protein (EC:... 34.3 0.091
hsa:9742 IFT140, DKFZp564L232, FLJ10306, FLJ30571, KIAA0590, W... 31.2 0.85
cel:C28F5.4 hypothetical protein; K01408 insulysin [EC:3.4.24.56] 30.8 1.1
mmu:106633 Ift140, AI661311, MGC170632, Tce5, Wdtc2, mKIAA0590... 30.4 1.6
cpv:cgd2_930 peptidase'insulinase-like peptidase' ; K01408 ins... 30.4 1.7
cpv:cgd6_520 Ser/Thr protein kinase 29.6 2.6
dre:334519 rnf17, fj98c04, im:7147535, wu:fj98c04; ring finger... 29.3 3.0
dre:768157 cplx4a, CPLX4, si:ch211-233a1.5, zgc:153429; comple... 28.9 4.3
cpv:cgd2_4270 secreted insulinase-like peptidase 28.5 5.6
tgo:TGME49_047760 long chain acyl-CoA synthetase, putative (EC... 28.1 6.5
mmu:240087 Mdc1, 6820401C03, AA413496, Nfbd1, mKIAA0170; media... 28.1 6.7
ath:AT3G57470 peptidase M16 family protein / insulinase family... 27.7 8.9
> tgo:TGME49_057010 insulysin, putative (EC:3.4.24.56); K01408
insulysin [EC:3.4.24.56]
Length=953
Score = 123 bits (309), Expect = 1e-28, Method: Compositional matrix adjust.
Identities = 57/107 (53%), Positives = 77/107 (71%), Gaps = 0/107 (0%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTH 60
LG+PSIE+RVNLAVL Q LNRR++D LRTE QLGYI GA+ +S L+C +EG++ H
Sbjct 778 LGVPSIEERVNLAVLGQMLNRRLFDRLRTEEQLGYIVGARSYIDSSVESLRCVLEGSRKH 837
Query 61 PDEVVKMIDEELTKAKEYLANMPDAEMARWKEAAHAKLTKREATFSE 107
PDE+ +ID+EL K ++L ++ D E+ WKE+A A+L K TF E
Sbjct 838 PDEIADLIDKELWKMNDHLQSISDGELDHWKESARAELEKPTETFYE 884
> cpv:cgd1_1680 insulinase like protease, signal peptide ; K01408
insulysin [EC:3.4.24.56]
Length=1033
Score = 48.5 bits (114), Expect = 5e-06, Method: Composition-based stats.
Identities = 27/68 (39%), Positives = 38/68 (55%), Gaps = 0/68 (0%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTH 60
G+PS E++++L L + IYD+LRT QLGYI A +ST LL VEG +
Sbjct 829 FGVPSFEEKLHLMALQPIIQGYIYDNLRTNKQLGYIIFANIVPISSTRLLVVGVEGDNNN 888
Query 61 PDEVVKMI 68
E ++ I
Sbjct 889 SVEKIESI 896
> tgo:TGME49_069870 hypothetical protein
Length=413
Score = 47.0 bits (110), Expect = 2e-05, Method: Composition-based stats.
Identities = 26/96 (27%), Positives = 50/96 (52%), Gaps = 5/96 (5%)
Query 2 GIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGA---K 58
+P+I DR L ++++++++R ++ LRTE QLGY+ S+ + F+
Sbjct 215 ALPNIHDRAVLYMVSRWMSQRFFNKLRTEQQLGYLTAMHSSRLEDRFYYRFFITSTYDPA 274
Query 59 THPDEVVKMIDEELTK--AKEYLANMPDAEMARWKE 92
D +V+ I+ E +K +E A + A + WK+
Sbjct 275 EVADRIVEFINAERSKIPTQEEFATLKQAAIDVWKQ 310
> cel:Y70C5C.1 hypothetical protein; K01408 insulysin [EC:3.4.24.56]
Length=985
Score = 45.1 bits (105), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 27/106 (25%), Positives = 49/106 (46%), Gaps = 1/106 (0%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTH 60
+G+ + D + ++ + +D+LRT+ LGYI + T LQ V+G K+
Sbjct 764 IGVQNTYDNAVIGLIKNLITEPAFDTLRTKESLGYIVWTRTHFNCGTVALQILVQGPKS- 822
Query 61 PDEVVKMIDEELTKAKEYLANMPDAEMARWKEAAHAKLTKREATFS 106
D V++ I+ L ++ + MP E A+L ++ T S
Sbjct 823 VDHVLERIEAFLESVRKEIVEMPQEEFENRVSGLIAQLEEKPKTLS 868
> cel:F44E7.4 hypothetical protein; K01408 insulysin [EC:3.4.24.56]
Length=1051
Score = 42.4 bits (98), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 27/106 (25%), Positives = 48/106 (45%), Gaps = 1/106 (0%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTH 60
+G+ + D + ++ Q + +++LRT LGYI T L V+G K+
Sbjct 823 IGVQNTYDNAVVGLIDQLIREPAFNTLRTNEALGYIVWTGSRLNCGTVALNVIVQGPKS- 881
Query 61 PDEVVKMIDEELTKAKEYLANMPDAEMARWKEAAHAKLTKREATFS 106
D V++ I+ L ++ +A MP E A+L ++ T S
Sbjct 882 VDHVLERIEVFLESVRKEIAEMPQEEFDNQVSGMIARLEEKPKTLS 927
> cel:C02G6.1 hypothetical protein; K01408 insulysin [EC:3.4.24.56]
Length=980
Score = 42.0 bits (97), Expect = 5e-04, Method: Compositional matrix adjust.
Identities = 27/106 (25%), Positives = 49/106 (46%), Gaps = 1/106 (0%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTH 60
+G+ + D + ++ Q + ++D+LRT LGYI L FV+G K+
Sbjct 753 IGVQNKYDNAVVGLIDQLIKEPVFDTLRTNEALGYIVWTGCRFNCGAVALNIFVQGPKS- 811
Query 61 PDEVVKMIDEELTKAKEYLANMPDAEMARWKEAAHAKLTKREATFS 106
D V++ I+ L ++ + MP E + A+L ++ T S
Sbjct 812 VDYVLERIEVFLESVRKEIIEMPQDEFEKKVAGMIARLEEKPKTLS 857
> sce:YLR389C STE23; Metalloprotease involved, with homolog Axl1p,
in N-terminal processing of pro-A-factor to the mature
form; member of the insulin-degrading enzyme family (EC:3.4.24.-);
K01408 insulysin [EC:3.4.24.56]
Length=1027
Score = 35.0 bits (79), Expect = 0.062, Method: Composition-based stats.
Identities = 22/95 (23%), Positives = 44/95 (46%), Gaps = 1/95 (1%)
Query 13 AVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTHPDEVVKMIDEEL 72
+ Q ++ +D+LRT+ QLGY+ + TA ++ ++ T P + I+
Sbjct 818 GLFAQLIHEPCFDTLRTKEQLGYVVFSSSLNNHGTANIRILIQSEHTTP-YLEWRINNFY 876
Query 73 TKAKEYLANMPDAEMARWKEAAHAKLTKREATFSE 107
+ L +MP+ + + KEA L ++ +E
Sbjct 877 ETFGQVLRDMPEEDFEKHKEALCNSLLQKFKNMAE 911
> tgo:TGME49_006510 peptidase M16 domain containing protein (EC:4.1.1.70
3.4.24.13 3.4.24.56 3.4.21.10 3.4.24.35 3.2.1.91)
Length=2435
Score = 34.3 bits (77), Expect = 0.091, Method: Compositional matrix adjust.
Identities = 22/106 (20%), Positives = 53/106 (50%), Gaps = 2/106 (1%)
Query 4 PSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTHPDE 63
P + + V +++ + ++ +D++RT GY+A A + L V+G++ PDE
Sbjct 1203 PDMMEMVVYSLIGEVISSPFFDTIRTHWMDGYVAAAAVREVPPAMTLATIVQGSQRKPDE 1262
Query 64 VVKMIDEELTKAKEYLAN--MPDAEMARWKEAAHAKLTKREATFSE 107
+ + + L + +E + + +A + R + + +K + +FS+
Sbjct 1263 LERHVCAFLAEMEENIGSSMTTEAFLERLRWLSSSKFHRSATSFSD 1308
> hsa:9742 IFT140, DKFZp564L232, FLJ10306, FLJ30571, KIAA0590,
WDTC2, c305C8.4, c380F5.1, gs114; intraflagellar transport
140 homolog (Chlamydomonas)
Length=1462
Score = 31.2 bits (69), Expect = 0.85, Method: Compositional matrix adjust.
Identities = 19/67 (28%), Positives = 32/67 (47%), Gaps = 1/67 (1%)
Query 24 YDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTHPDEVVKMIDEELTKAKEYLANMP 83
+D + + AG E+ A + L + E + TH EV +M+ E+L + Y+ M
Sbjct 895 HDRVHLRSTYHRYAGHLEASADCSRALS-YYEKSDTHRFEVPRMLSEDLPSLELYVNKMK 953
Query 84 DAEMARW 90
D + RW
Sbjct 954 DKTLWRW 960
> cel:C28F5.4 hypothetical protein; K01408 insulysin [EC:3.4.24.56]
Length=856
Score = 30.8 bits (68), Expect = 1.1, Method: Composition-based stats.
Identities = 21/86 (24%), Positives = 37/86 (43%), Gaps = 1/86 (1%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTH 60
+G+ S + +L + + Y LRT LGY + L V+G ++
Sbjct 695 IGVQSTYNNSVNKLLNELIKNPAYTILRTNEALGYNVSTESRLNDGNVYLHVIVQGPES- 753
Query 61 PDEVVKMIDEELTKAKEYLANMPDAE 86
D V++ I+ L A+E + MP +
Sbjct 754 ADHVLERIEVFLESAREEIVAMPQED 779
> mmu:106633 Ift140, AI661311, MGC170632, Tce5, Wdtc2, mKIAA0590;
intraflagellar transport 140 homolog (Chlamydomonas)
Length=1464
Score = 30.4 bits (67), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 13/38 (34%), Positives = 21/38 (55%), Gaps = 0/38 (0%)
Query 53 FVEGAKTHPDEVVKMIDEELTKAKEYLANMPDAEMARW 90
+ E + TH EV +M+ E+L + Y+ M D + RW
Sbjct 924 YYEKSDTHRFEVPRMLSEDLQSLELYINRMKDKTLWRW 961
> cpv:cgd2_930 peptidase'insulinase-like peptidase' ; K01408 insulysin
[EC:3.4.24.56]
Length=1013
Score = 30.4 bits (67), Expect = 1.7, Method: Composition-based stats.
Identities = 15/39 (38%), Positives = 24/39 (61%), Gaps = 0/39 (0%)
Query 1 LGIPSIEDRVNLAVLTQFLNRRIYDSLRTEAQLGYIAGA 39
+G ++ + V L +L+++L+ Y LRT QLGYI A
Sbjct 812 MGEYNLRNYVLLELLSKYLDSNCYLELRTNQQLGYIVHA 850
> cpv:cgd6_520 Ser/Thr protein kinase
Length=1103
Score = 29.6 bits (65), Expect = 2.6, Method: Composition-based stats.
Identities = 18/55 (32%), Positives = 25/55 (45%), Gaps = 1/55 (1%)
Query 21 RRIYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTHPDEVVKMIDEELTKA 75
RR Y T YIA K + +T LL CF G HP E++ + + + K
Sbjct 235 RRKYYVKVTPINTKYIAIVK-TITNNTPLLLCFEHGKDRHPKEIINLKNSNIYKG 288
> dre:334519 rnf17, fj98c04, im:7147535, wu:fj98c04; ring finger
protein 17
Length=1485
Score = 29.3 bits (64), Expect = 3.0, Method: Composition-based stats.
Identities = 20/49 (40%), Positives = 25/49 (51%), Gaps = 2/49 (4%)
Query 55 EGAKTHPDEVVKMIDEELTKAKEYLANMPDAEMARWKEAAHAKLTKREA 103
E A+ PD VVK++DE LTKA E L N D H +L K +
Sbjct 114 ENAEGKPD-VVKVLDEALTKATENL-NQLDKLHQTLANGVHGQLRKERS 160
> dre:768157 cplx4a, CPLX4, si:ch211-233a1.5, zgc:153429; complexin
4a
Length=159
Score = 28.9 bits (63), Expect = 4.3, Method: Compositional matrix adjust.
Identities = 16/45 (35%), Positives = 28/45 (62%), Gaps = 6/45 (13%)
Query 61 PDEVVKMIDEELTKAKE------YLANMPDAEMARWKEAAHAKLT 99
P+E++KM+DE+ T+ +E + N+ + +M + KE A A LT
Sbjct 102 PEELLKMVDEDATEEEEKDSIMGQIQNLQNMDMDQIKEKASATLT 146
> cpv:cgd2_4270 secreted insulinase-like peptidase
Length=1257
Score = 28.5 bits (62), Expect = 5.6, Method: Composition-based stats.
Identities = 13/23 (56%), Positives = 16/23 (69%), Gaps = 0/23 (0%)
Query 17 QFLNRRIYDSLRTEAQLGYIAGA 39
+ LN YD+LRTE Q GY+A A
Sbjct 950 EILNSPFYDTLRTEWQDGYVAFA 972
> tgo:TGME49_047760 long chain acyl-CoA synthetase, putative (EC:6.2.1.3);
K01897 long-chain acyl-CoA synthetase [EC:6.2.1.3]
Length=793
Score = 28.1 bits (61), Expect = 6.5, Method: Compositional matrix adjust.
Identities = 21/49 (42%), Positives = 26/49 (53%), Gaps = 10/49 (20%)
Query 23 IYDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTHPDEVVKMIDEE 71
+YD+L EA L YI G T L FVEGAK HP + ++ EE
Sbjct 266 LYDTLGHEALL-YIIGL-------TKLQVLFVEGAKLHP--ALNLVTEE 304
> mmu:240087 Mdc1, 6820401C03, AA413496, Nfbd1, mKIAA0170; mediator
of DNA damage checkpoint 1
Length=1708
Score = 28.1 bits (61), Expect = 6.7, Method: Composition-based stats.
Identities = 18/46 (39%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Query 30 EAQLGYIAGAKESQAASTALL-----QCFVEGAKTHPDEVVKMIDE 70
EA +G AKE A L QCFVEG HP+ V + +E
Sbjct 631 EAHVGRTKSAKECCDAEPEDLCLPATQCFVEGESQHPEAVQSLENE 676
> ath:AT3G57470 peptidase M16 family protein / insulinase family
protein; K01408 insulysin [EC:3.4.24.56]
Length=891
Score = 27.7 bits (60), Expect = 8.9, Method: Composition-based stats.
Identities = 16/63 (25%), Positives = 26/63 (41%), Gaps = 0/63 (0%)
Query 24 YDSLRTEAQLGYIAGAKESQAASTALLQCFVEGAKTHPDEVVKMIDEELTKAKEYLANMP 83
+ LRT QLGYI S + +Q ++ + P + ++ L + NM
Sbjct 711 FHQLRTIEQLGYITSLSLSNDSGVYGVQFIIQSSVKGPGHIDSRVESLLKDLESKFYNMS 770
Query 84 DAE 86
D E
Sbjct 771 DEE 773
Lambda K H
0.315 0.128 0.351
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2072286120
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40