bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
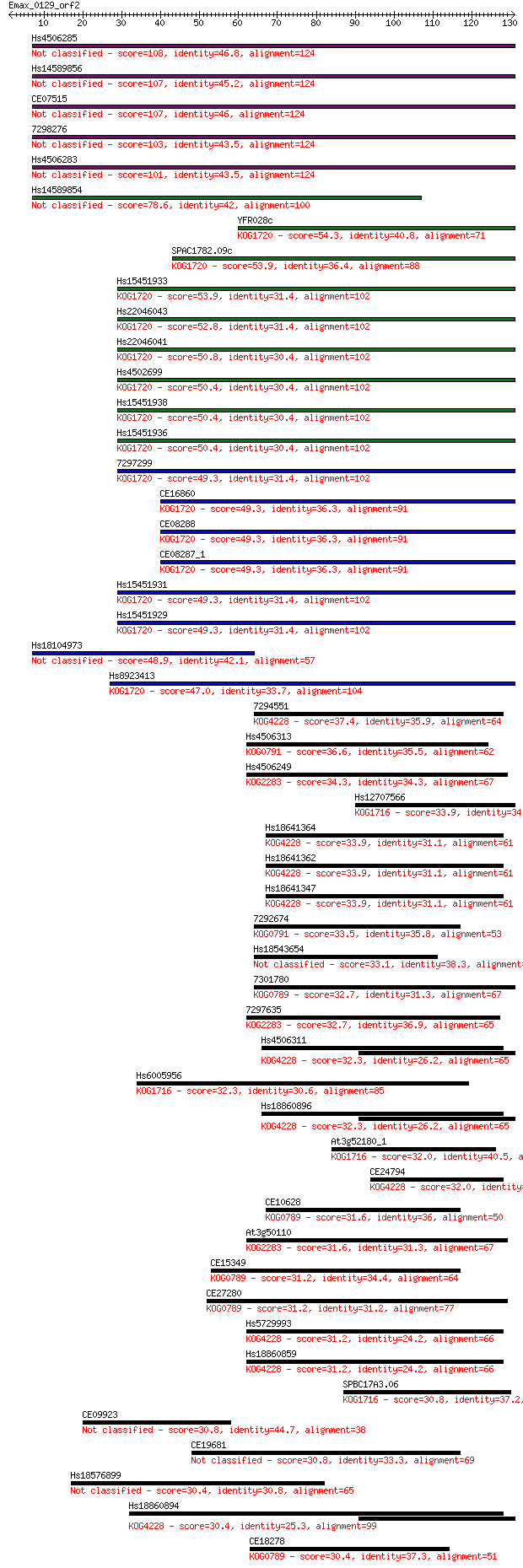
Query= Emax_0129_orf2
Length=130
Score E
Sequences producing significant alignments: (Bits) Value
Hs4506285 108 3e-24
Hs14589856 107 4e-24
CE07515 107 6e-24
7298276 103 7e-23
Hs4506283 101 3e-22
Hs14589854 78.6 3e-15
YFR028c 54.3 6e-08
SPAC1782.09c 53.9 8e-08
Hs15451933 53.9 8e-08
Hs22046043 52.8 1e-07
Hs22046041 50.8 6e-07
Hs4502699 50.4 7e-07
Hs15451938 50.4 7e-07
Hs15451936 50.4 7e-07
7297299 49.3 2e-06
CE16860 49.3 2e-06
CE08288 49.3 2e-06
CE08287_1 49.3 2e-06
Hs15451931 49.3 2e-06
Hs15451929 49.3 2e-06
Hs18104973 48.9 2e-06
Hs8923413 47.0 9e-06
7294551 37.4 0.007
Hs4506313 36.6 0.011
Hs4506249 34.3 0.058
Hs12707566 33.9 0.076
Hs18641364 33.9 0.084
Hs18641362 33.9 0.084
Hs18641347 33.9 0.084
7292674 33.5 0.12
Hs18543654 33.1 0.13
7301780 32.7 0.17
7297635 32.7 0.18
Hs4506311 32.3 0.25
Hs6005956 32.3 0.26
Hs18860896 32.3 0.26
At3g52180_1 32.0 0.28
CE24794 32.0 0.34
CE10628 31.6 0.36
At3g50110 31.6 0.42
CE15349 31.2 0.53
CE27280 31.2 0.57
Hs5729993 31.2 0.58
Hs18860859 31.2 0.58
SPBC17A3.06 30.8 0.63
CE09923 30.8 0.69
CE19681 30.8 0.70
Hs18576899 30.4 0.77
Hs18860894 30.4 0.86
CE18278 30.4 0.88
> Hs4506285
Length=167
Score = 108 bits (269), Expect = 3e-24, Method: Compositional matrix adjust.
Identities = 58/126 (46%), Positives = 73/126 (57%), Gaps = 2/126 (1%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDL 66
P I ++ LI PTN L + +L + GVT LVR C TYD+ PV GI V D
Sbjct 6 PVEISYENMRFLITHNPTNATLNKFTEELKKYGVTTLVRVCDATYDKAPVEKEGIHVLDW 65
Query 67 TFPDGEAPPAEVIARWRALAAQAKAE--GGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
F DG PP +++ W L E G +AVHCVAGLGR PVLVA++LI+ G + E+
Sbjct 66 PFDDGAPPPNQIVDDWLNLLKTKFREEPGCCVAVHCVAGLGRAPVLVALALIECGMKYED 125
Query 125 AVNFIR 130
AV FIR
Sbjct 126 AVQFIR 131
> Hs14589856
Length=173
Score = 107 bits (268), Expect = 4e-24, Method: Compositional matrix adjust.
Identities = 56/126 (44%), Positives = 77/126 (61%), Gaps = 2/126 (1%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDL 66
P + + ++ LI PTN L ++ L + G T +VR C TYD+ P+ GI V D
Sbjct 9 PVEVSYKHMRFLITHNPTNATLSTFIEDLKKYGATTVVRVCEVTYDKTPLEKDGITVVDW 68
Query 67 TFPDGEAPPAEVIARWRAL--AAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
F DG PP +V+ W +L A +A G +AVHCVAGLGR PVLVA++LI+SG + E+
Sbjct 69 PFDDGAPPPGKVVEDWLSLVKAKFCEAPGSCVAVHCVAGLGRAPVLVALALIESGMKYED 128
Query 125 AVNFIR 130
A+ FIR
Sbjct 129 AIQFIR 134
> CE07515
Length=190
Score = 107 bits (267), Expect = 6e-24, Method: Compositional matrix adjust.
Identities = 57/126 (45%), Positives = 76/126 (60%), Gaps = 2/126 (1%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDL 66
P+ I ++ LI D P N ++Q+Y+ +L + G +VR C PTYD + AGI V D
Sbjct 24 PSEIAWGKMRFLITDRPNNSSIQSYIEELEKHGARAVVRVCEPTYDTLALKEAGIDVLDW 83
Query 67 TFPDGEAPPAEVIARWRALAAQAKAE--GGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
F DG PP EVI W L + E +AVHCVAGLGR PVLVA++LI++G + E+
Sbjct 84 QFSDGSPPPPEVIKSWFQLCMTSFKEHPDKSIAVHCVAGLGRAPVLVAIALIEAGMKYED 143
Query 125 AVNFIR 130
AV IR
Sbjct 144 AVEMIR 149
> 7298276
Length=178
Score = 103 bits (258), Expect = 7e-23, Method: Compositional matrix adjust.
Identities = 54/128 (42%), Positives = 74/128 (57%), Gaps = 4/128 (3%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDL 66
P LIE +G+K LI D P++ + Y+ +L + V +VR C P+Y+ + GI V DL
Sbjct 14 PALIEYKGMKFLITDRPSDITINHYIMELKKNNVNTVVRVCEPSYNTDELETQGITVKDL 73
Query 67 TFPDGEAPPAEVIARWR----ALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEA 122
F DG PP +V+ W L + +AVHCVAGLGR PVLVA++LI+ G +
Sbjct 74 AFEDGTFPPQQVVDEWFEFFVVLYRYQQNPEACVAVHCVAGLGRAPVLVALALIELGLKY 133
Query 123 EEAVNFIR 130
E AV IR
Sbjct 134 EAAVEMIR 141
> Hs4506283
Length=173
Score = 101 bits (252), Expect = 3e-22, Method: Compositional matrix adjust.
Identities = 54/126 (42%), Positives = 73/126 (57%), Gaps = 2/126 (1%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDL 66
P + + ++ LI PTN L ++ +L + GVT +VR C TYD V GI V D
Sbjct 9 PVEVTYKNMRFLITHNPTNATLNKFIEELKKYGVTTIVRVCEATYDTTLVEKEGIHVLDW 68
Query 67 TFPDGEAPPAEVIARWRALAAQAKAE--GGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
F DG P +++ W +L E G +AVHCVAGLGR PVLVA++LI+ G + E+
Sbjct 69 PFDDGAPPSNQIVDDWLSLVKIKFREEPGCCIAVHCVAGLGRAPVLVALALIEGGMKYED 128
Query 125 AVNFIR 130
AV FIR
Sbjct 129 AVQFIR 134
> Hs14589854
Length=148
Score = 78.6 bits (192), Expect = 3e-15, Method: Compositional matrix adjust.
Identities = 42/102 (41%), Positives = 57/102 (55%), Gaps = 2/102 (1%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDL 66
P + + ++ LI PTN L ++ L + G T +VR C TYD+ P+ GI V D
Sbjct 9 PVEVSYKHMRFLITHNPTNATLSTFIEDLKKYGATTVVRVCEVTYDKTPLEKDGITVVDW 68
Query 67 TFPDGEAPPAEVIARWRAL--AAQAKAEGGVLAVHCVAGLGR 106
F DG PP +V+ W +L A +A G +AVHCVAGLGR
Sbjct 69 PFDDGAPPPGKVVEDWLSLVKAKFCEAPGSCVAVHCVAGLGR 110
> YFR028c
Length=551
Score = 54.3 bits (129), Expect = 6e-08, Method: Composition-based stats.
Identities = 29/72 (40%), Positives = 38/72 (52%), Gaps = 1/72 (1%)
Query 60 GIRVHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDS- 118
GI+ DL F DG P ++ + A GG +AVHC AGLGR L+ LI +
Sbjct 243 GIQHLDLIFEDGTCPDLSIVKNFVGAAETIIKRGGKIAVHCKAGLGRTGCLIGAHLIYTY 302
Query 119 GFEAEEAVNFIR 130
GF A E + F+R
Sbjct 303 GFTANECIGFLR 314
> SPAC1782.09c
Length=537
Score = 53.9 bits (128), Expect = 8e-08, Method: Composition-based stats.
Identities = 32/89 (35%), Positives = 46/89 (51%), Gaps = 2/89 (2%)
Query 43 LVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVHCVA 102
+VR P YD+ GIR ++ F DG P ++ + L + + E GV+AVHC A
Sbjct 230 IVRLNGPLYDKKTFENVGIRHKEMYFEDGTVPELSLVKEFIDLTEEVE-EDGVIAVHCKA 288
Query 103 GLGRGPVLVAVSLI-DSGFEAEEAVNFIR 130
GLGR L+ LI F A E + ++R
Sbjct 289 GLGRTGCLIGAYLIYKHCFTANEVIAYMR 317
> Hs15451933
Length=383
Score = 53.9 bits (128), Expect = 8e-08, Method: Compositional matrix adjust.
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+AY + VT +VR Y+ AG +DL F DG P ++ R+ +
Sbjct 210 EAYFPYFKKHNVTAVVRLNKKIYEAKRFTDAGFEHYDLFFIDGSTPSDNIVRRFLNICEN 269
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
+ G +AVHC AGLGR L+A ++ F E + +IR
Sbjct 270 TE---GAIAVHCKAGLGRTGTLIACYVMKHYRFTHAEIIAWIR 309
> Hs22046043
Length=447
Score = 52.8 bits (125), Expect = 1e-07, Method: Composition-based stats.
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+ Y+ VT ++R YD AG HDL F DG P ++ R+ +
Sbjct 216 ETYIQYFKNHNVTTIIRLNKRMYDAKRFTDAGFDHHDLFFADGSTPTDAIVKRFLDICEN 275
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
A+ G +AVHC AGLGR L+A ++ A E + ++R
Sbjct 276 AE---GAIAVHCKAGLGRTGTLIACYIMKHYRMTAAETIAWVR 315
> Hs22046041
Length=400
Score = 50.8 bits (120), Expect = 6e-07, Method: Composition-based stats.
Identities = 31/103 (30%), Positives = 48/103 (46%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+ Y+ VT ++R +D AG HDL F DG P ++ R+ +
Sbjct 169 ETYIQYFKNHNVTTIIRLNKRIHDAKRFTDAGFDHHDLFFADGSTPTDAIVKRFLDICEN 228
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
A+ G +AVHC AGLGR L+A ++ A E + ++R
Sbjct 229 AE---GAIAVHCKAGLGRTGTLIACYIMKHYRMTAAETIAWVR 268
> Hs4502699
Length=459
Score = 50.4 bits (119), Expect = 7e-07, Method: Composition-based stats.
Identities = 31/103 (30%), Positives = 47/103 (45%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+ Y+ VT ++R YD AG HDL F DG P ++ + +
Sbjct 246 ETYIQYFKNHNVTTIIRLNKRMYDAKRFTDAGFDHHDLFFADGSTPTDAIVKEFLDICEN 305
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
A+ G +AVHC AGLGR L+A ++ A E + ++R
Sbjct 306 AE---GAIAVHCKAGLGRTGTLIACYIMKHYRMTAAETIAWVR 345
> Hs15451938
Length=471
Score = 50.4 bits (119), Expect = 7e-07, Method: Composition-based stats.
Identities = 31/103 (30%), Positives = 47/103 (45%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+ Y+ VT ++R YD AG HDL F DG P ++ + +
Sbjct 246 ETYIQYFKNHNVTTIIRLNKRMYDAKRFTDAGFDHHDLFFADGSTPTDAIVKEFLDICEN 305
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
A+ G +AVHC AGLGR L+A ++ A E + ++R
Sbjct 306 AE---GAIAVHCKAGLGRTGTLIACYIMKHYRMTAAETIAWVR 345
> Hs15451936
Length=498
Score = 50.4 bits (119), Expect = 7e-07, Method: Composition-based stats.
Identities = 31/103 (30%), Positives = 47/103 (45%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+ Y+ VT ++R YD AG HDL F DG P ++ + +
Sbjct 246 ETYIQYFKNHNVTTIIRLNKRMYDAKRFTDAGFDHHDLFFADGSTPTDAIVKEFLDICEN 305
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
A+ G +AVHC AGLGR L+A ++ A E + ++R
Sbjct 306 AE---GAIAVHCKAGLGRTGTLIACYIMKHYRMTAAETIAWVR 345
> 7297299
Length=553
Score = 49.3 bits (116), Expect = 2e-06, Method: Composition-based stats.
Identities = 32/103 (31%), Positives = 49/103 (47%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+ Y + VT ++R Y AG DL F DG P ++ ++ ++
Sbjct 215 ERYFSYFRDNNVTTVIRLNAKVYHASSFENAGFDHKDLFFIDGSTPSDAIMKKFLSICET 274
Query 89 AKAEGGVLAVHCVAGLGR-GPVLVAVSLIDSGFEAEEAVNFIR 130
K G +AVHC AGLGR G ++ A + GF A EA+ ++R
Sbjct 275 TK---GAIAVHCKAGLGRTGSLIGAYIMKHYGFTALEAIAWLR 314
> CE16860
Length=708
Score = 49.3 bits (116), Expect = 2e-06, Method: Composition-based stats.
Identities = 33/92 (35%), Positives = 47/92 (51%), Gaps = 4/92 (4%)
Query 40 VTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVH 99
V+ +VR YD AG DL F DG P E++ ++ + K GGV AVH
Sbjct 238 VSTIVRLNAKNYDASKFTKAGFDHVDLFFIDGSTPSDEIMLKFIKVVDNTK--GGV-AVH 294
Query 100 CVAGLGRGPVLVAVSLI-DSGFEAEEAVNFIR 130
C AGLGR L+A ++ + G A E + ++R
Sbjct 295 CKAGLGRTGTLIACWMMKEYGLTAGECMGWLR 326
> CE08288
Length=681
Score = 49.3 bits (116), Expect = 2e-06, Method: Composition-based stats.
Identities = 33/92 (35%), Positives = 47/92 (51%), Gaps = 4/92 (4%)
Query 40 VTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVH 99
V+ +VR YD AG DL F DG P E++ ++ + K GGV AVH
Sbjct 238 VSTIVRLNAKNYDASKFTKAGFDHVDLFFIDGSTPSDEIMLKFIKVVDNTK--GGV-AVH 294
Query 100 CVAGLGRGPVLVAVSLI-DSGFEAEEAVNFIR 130
C AGLGR L+A ++ + G A E + ++R
Sbjct 295 CKAGLGRTGTLIACWMMKEYGLTAGECMGWLR 326
> CE08287_1
Length=753
Score = 49.3 bits (116), Expect = 2e-06, Method: Composition-based stats.
Identities = 33/92 (35%), Positives = 47/92 (51%), Gaps = 4/92 (4%)
Query 40 VTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVH 99
V+ +VR YD AG DL F DG P E++ ++ + K GGV AVH
Sbjct 238 VSTIVRLNAKNYDASKFTKAGFDHVDLFFIDGSTPSDEIMLKFIKVVDNTK--GGV-AVH 294
Query 100 CVAGLGRGPVLVAVSLI-DSGFEAEEAVNFIR 130
C AGLGR L+A ++ + G A E + ++R
Sbjct 295 CKAGLGRTGTLIACWMMKEYGLTAGECMGWLR 326
> Hs15451931
Length=623
Score = 49.3 bits (116), Expect = 2e-06, Method: Composition-based stats.
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+AY + VT +VR Y+ AG +DL F DG P ++ R+ +
Sbjct 210 EAYFPYFKKHNVTAVVRLNKKIYEAKRFTDAGFEHYDLFFIDGSTPSDNIVRRFLNICEN 269
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
+ G +AVHC AGLGR L+A ++ F E + +IR
Sbjct 270 TE---GAIAVHCKAGLGRTGTLIACYVMKHYRFTHAEIIAWIR 309
> Hs15451929
Length=594
Score = 49.3 bits (116), Expect = 2e-06, Method: Composition-based stats.
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 4/103 (3%)
Query 29 QAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ 88
+AY + VT +VR Y+ AG +DL F DG P ++ R+ +
Sbjct 210 EAYFPYFKKHNVTAVVRLNKKIYEAKRFTDAGFEHYDLFFIDGSTPSDNIVRRFLNICEN 269
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS-GFEAEEAVNFIR 130
+ G +AVHC AGLGR L+A ++ F E + +IR
Sbjct 270 TE---GAIAVHCKAGLGRTGTLIACYVMKHYRFTHAEIIAWIR 309
> Hs18104973
Length=82
Score = 48.9 bits (115), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/57 (42%), Positives = 30/57 (52%), Gaps = 0/57 (0%)
Query 7 PTLIESRGVKCLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRV 63
P I ++ LI PTN L + +L + GVT LVR C TYD+ PV GI V
Sbjct 6 PVEISYENMRFLITHNPTNATLNKFTEELKKYGVTTLVRVCDATYDKAPVEKEGIHV 62
> Hs8923413
Length=150
Score = 47.0 bits (110), Expect = 9e-06, Method: Compositional matrix adjust.
Identities = 35/108 (32%), Positives = 51/108 (47%), Gaps = 8/108 (7%)
Query 27 NLQAYLAQLVQVGVTDLVRTC---PPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWR 83
L A+ L+ +GV LV PP D P G+ +H L PD P + I R+
Sbjct 23 RLPAHYQFLLDLGVRHLVSLTERGPPHSDSCP----GLTLHRLRIPDFCPPAPDQIDRFV 78
Query 84 ALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLI-DSGFEAEEAVNFIR 130
+ +A A G + VHC G GR ++A L+ + G A +A+ IR
Sbjct 79 QIVDEANARGEAVGVHCALGFGRTGTMLACYLVKERGLAAGDAIAEIR 126
> 7294551
Length=1428
Score = 37.4 bits (85), Expect = 0.007, Method: Composition-based stats.
Identities = 23/66 (34%), Positives = 35/66 (53%), Gaps = 2/66 (3%)
Query 64 HDLTFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFE 121
H LT+ D AP P +I R + + + G + VHC AG+GR LVA+ + E
Sbjct 1025 HYLTWKDFMAPEHPHGIIKFIRQINSVYSLQRGPILVHCSAGVGRTGTLVALDSLIQQLE 1084
Query 122 AEEAVN 127
E++V+
Sbjct 1085 EEDSVS 1090
> Hs4506313
Length=1118
Score = 36.6 bits (83), Expect = 0.011, Method: Composition-based stats.
Identities = 22/66 (33%), Positives = 33/66 (50%), Gaps = 4/66 (6%)
Query 62 RVHDLTFPDGEAP--PAEVIARWRALAA--QAKAEGGVLAVHCVAGLGRGPVLVAVSLID 117
+ H +PD P P ++A WR L EGG VHC AG+GR L+A+ ++
Sbjct 980 QFHYQAWPDHGVPSSPDTLLAFWRMLRQWLDQTMEGGPPIVHCSAGVGRTGTLIALDVLL 1039
Query 118 SGFEAE 123
++E
Sbjct 1040 RQLQSE 1045
> Hs4506249
Length=403
Score = 34.3 bits (77), Expect = 0.058, Method: Composition-based stats.
Identities = 23/71 (32%), Positives = 36/71 (50%), Gaps = 4/71 (5%)
Query 62 RVHDLTFPDGEAPPAEVIARWRALAAQ--AKAEGGVLAVHCVAGLGRGPVLVAVSLIDSG 119
RV F D P E+I + Q ++ + V A+HC AG GR V++ L+ G
Sbjct 84 RVAQYPFEDHNPPQLELIKPFCEDLDQWLSEDDNHVAAIHCKAGKGRTGVMICAYLLHRG 143
Query 120 --FEAEEAVNF 128
+A+EA++F
Sbjct 144 KFLKAQEALDF 154
> Hs12707566
Length=384
Score = 33.9 bits (76), Expect = 0.076, Method: Compositional matrix adjust.
Identities = 14/42 (33%), Positives = 25/42 (59%), Gaps = 1/42 (2%)
Query 90 KAEGGVLAVHCVAGLGRGPVLVAVSLIDSG-FEAEEAVNFIR 130
+ +GG + VHC AG+ R P + L+ + F +EA ++I+
Sbjct 253 REKGGKVLVHCEAGISRSPTICMAYLMKTKQFRLKEAFDYIK 294
> Hs18641364
Length=1256
Score = 33.9 bits (76), Expect = 0.084, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 2/63 (3%)
Query 67 TFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
++PD P P ++ R + A + G + VHC AG+GR + + + G EAE
Sbjct 768 SWPDHGVPEDPHLLLKLRRRVNAFSNFFSGPIVVHCSAGVGRTGTYIGIDAMLEGLEAEN 827
Query 125 AVN 127
V+
Sbjct 828 KVD 830
> Hs18641362
Length=1143
Score = 33.9 bits (76), Expect = 0.084, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 2/63 (3%)
Query 67 TFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
++PD P P ++ R + A + G + VHC AG+GR + + + G EAE
Sbjct 655 SWPDHGVPEDPHLLLKLRRRVNAFSNFFSGPIVVHCSAGVGRTGTYIGIDAMLEGLEAEN 714
Query 125 AVN 127
V+
Sbjct 715 KVD 717
> Hs18641347
Length=1304
Score = 33.9 bits (76), Expect = 0.084, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 2/63 (3%)
Query 67 TFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEE 124
++PD P P ++ R + A + G + VHC AG+GR + + + G EAE
Sbjct 816 SWPDHGVPEDPHLLLKLRRRVNAFSNFFSGPIVVHCSAGVGRTGTYIGIDAMLEGLEAEN 875
Query 125 AVN 127
V+
Sbjct 876 KVD 878
> 7292674
Length=1647
Score = 33.5 bits (75), Expect = 0.12, Method: Composition-based stats.
Identities = 19/55 (34%), Positives = 28/55 (50%), Gaps = 2/55 (3%)
Query 64 HDLTFPDGEAP-PAEVIARW-RALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLI 116
H T+PD P P + + R+ RA + AE + VHC AG+GR + + I
Sbjct 1446 HFTTWPDFGVPNPPQTLVRFVRAFRDRIGAEQRPIVVHCSAGVGRSGTFITLDRI 1500
> Hs18543654
Length=1688
Score = 33.1 bits (74), Expect = 0.13, Method: Composition-based stats.
Identities = 18/48 (37%), Positives = 21/48 (43%), Gaps = 1/48 (2%)
Query 64 HDLTFPDGEAPPAEVIARWR-ALAAQAKAEGGVLAVHCVAGLGRGPVL 110
H + P E PPA + W ALA + G A GRGPVL
Sbjct 370 HSRSLPPSEGPPASPLMAWMAALAVGLRTTAGSTAHRSSEARGRGPVL 417
> 7301780
Length=1285
Score = 32.7 bits (73), Expect = 0.17, Method: Composition-based stats.
Identities = 21/72 (29%), Positives = 33/72 (45%), Gaps = 5/72 (6%)
Query 64 HDLTFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFE 121
H +PD P P V+ + +A AE G + VHC AG+GR + + + +
Sbjct 632 HYTNWPDHGTPDHPLPVLNFVKKSSAANPAEAGPIVVHCSAGVGRTGTYIVLDAMLKQIQ 691
Query 122 AEEAVN---FIR 130
+ VN F+R
Sbjct 692 QKNIVNVFGFLR 703
> 7297635
Length=514
Score = 32.7 bits (73), Expect = 0.18, Method: Composition-based stats.
Identities = 24/69 (34%), Positives = 34/69 (49%), Gaps = 4/69 (5%)
Query 62 RVHDLTFPDGEAPPAEVIARWRALAAQAKAE--GGVLAVHCVAGLGRGPVLVAVSLIDSG 119
RV F D P E+I R+ + E V+AVHC AG GR ++ L+ SG
Sbjct 92 RVAVYPFDDHNPPTIELIQRFCSDVDMWLKEDSSNVVAVHCKAGKGRTGTMICAYLVFSG 151
Query 120 FE--AEEAV 126
+ A+EA+
Sbjct 152 IKKSADEAL 160
> Hs4506311
Length=1897
Score = 32.3 bits (72), Expect = 0.25, Method: Composition-based stats.
Identities = 17/64 (26%), Positives = 31/64 (48%), Gaps = 2/64 (3%)
Query 66 LTFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAE 123
+ +PD P P ++A R + A + G + VHC AG+GR + + + + E
Sbjct 1502 MAWPDHGVPEYPTPILAFLRRVKACNPLDAGPMVVHCSAGVGRTGCFIVIDAMLERMKHE 1561
Query 124 EAVN 127
+ V+
Sbjct 1562 KTVD 1565
Score = 27.7 bits (60), Expect = 5.1, Method: Composition-based stats.
Identities = 12/40 (30%), Positives = 21/40 (52%), Gaps = 0/40 (0%)
Query 91 AEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEEAVNFIR 130
+ G + VHC AG+GR V + +S++ E V+ +
Sbjct 1820 GQDGPITVHCSAGVGRTGVFITLSIVLERMRYEGVVDMFQ 1859
> Hs6005956
Length=340
Score = 32.3 bits (72), Expect = 0.26, Method: Compositional matrix adjust.
Identities = 26/90 (28%), Positives = 44/90 (48%), Gaps = 8/90 (8%)
Query 34 QLVQVGVTDL--VRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIA---RWRALAAQ 88
L + G+T + V + P++ GP V R L P + P ++++ R A Q
Sbjct 47 HLREAGITAVLTVDSEEPSFKAGPGVEDLWR---LFVPALDKPETDLLSHLDRCVAFIGQ 103
Query 89 AKAEGGVLAVHCVAGLGRGPVLVAVSLIDS 118
A+AEG + VHC AG+ R ++ L+ +
Sbjct 104 ARAEGRAVLVHCHAGVSRSVAIITAFLMKT 133
> Hs18860896
Length=1888
Score = 32.3 bits (72), Expect = 0.26, Method: Composition-based stats.
Identities = 17/64 (26%), Positives = 31/64 (48%), Gaps = 2/64 (3%)
Query 66 LTFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAE 123
+ +PD P P ++A R + A + G + VHC AG+GR + + + + E
Sbjct 1493 MAWPDHGVPEYPTPILAFLRRVKACNPLDAGPMVVHCSAGVGRTGCFIVIDAMLERMKHE 1552
Query 124 EAVN 127
+ V+
Sbjct 1553 KTVD 1556
Score = 27.7 bits (60), Expect = 5.2, Method: Composition-based stats.
Identities = 12/40 (30%), Positives = 21/40 (52%), Gaps = 0/40 (0%)
Query 91 AEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEEAVNFIR 130
+ G + VHC AG+GR V + +S++ E V+ +
Sbjct 1811 GQDGPITVHCSAGVGRTGVFITLSIVLERMRYEGVVDMFQ 1850
> At3g52180_1
Length=248
Score = 32.0 bits (71), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 17/43 (39%), Positives = 21/43 (48%), Gaps = 1/43 (2%)
Query 84 ALAAQAKAEGGVLAVHCVAGLGRGP-VLVAVSLIDSGFEAEEA 125
L K GGV VHC AG+GR P V + G++ EA
Sbjct 182 TLYKAVKRNGGVTYVHCTAGMGRAPAVALTYMFWVQGYKLMEA 224
> CE24794
Length=2234
Score = 32.0 bits (71), Expect = 0.34, Method: Composition-based stats.
Identities = 14/34 (41%), Positives = 20/34 (58%), Gaps = 0/34 (0%)
Query 94 GVLAVHCVAGLGRGPVLVAVSLIDSGFEAEEAVN 127
G + VHC +G GR V +A+S+I AE V+
Sbjct 2161 GPITVHCCSGAGRTAVFIALSIILDRMRAEHVVD 2194
> CE10628
Length=374
Score = 31.6 bits (70), Expect = 0.36, Method: Compositional matrix adjust.
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 2/50 (4%)
Query 67 TFPDGEAPPAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLI 116
+PD APP + A L + + VHC AG+GR +V +SLI
Sbjct 234 NWPDHGAPPINMGAI--NLIEAVNYDTNPVVVHCSAGVGRSGTIVGISLI 281
> At3g50110
Length=628
Score = 31.6 bits (70), Expect = 0.42, Method: Composition-based stats.
Identities = 21/71 (29%), Positives = 34/71 (47%), Gaps = 4/71 (5%)
Query 62 RVHDLTFPDGEAPPAEVIARWRALAAQAKAEG--GVLAVHCVAGLGRGPVLVAVSLIDSG 119
+V F D PP ++I + A E V+ VHC AG+ R +++ L+
Sbjct 267 KVASFPFDDHNCPPIQLIPSFCQSAYTWLKEDIQNVVVVHCKAGMARTGLMICCLLLYLK 326
Query 120 F--EAEEAVNF 128
F AEEA+++
Sbjct 327 FFPTAEEAIDY 337
> CE15349
Length=344
Score = 31.2 bits (69), Expect = 0.53, Method: Compositional matrix adjust.
Identities = 22/66 (33%), Positives = 32/66 (48%), Gaps = 4/66 (6%)
Query 53 EGPVVLAG--IRVHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVL 110
EGP LA + H + +PD P A++ L A+ + G VHC AG+GR +
Sbjct 210 EGPGGLAQKVTQYHWIDWPDRGVPTADMAIV--ELLAKTRPSKGPTVVHCSAGIGRTGSV 267
Query 111 VAVSLI 116
V + I
Sbjct 268 VMIEYI 273
> CE27280
Length=352
Score = 31.2 bits (69), Expect = 0.57, Method: Compositional matrix adjust.
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 5/80 (6%)
Query 52 DEGPVVLAGIRVHDL---TFPDGEAPPAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGP 108
DE P L +RV+ + +PD P + + L AQ + G VHC AG+GR
Sbjct 215 DEQPANLKELRVNLIKWPNWPDRGVPDEKCHTVPQRLLAQVR--HGPCVVHCSAGIGRTG 272
Query 109 VLVAVSLIDSGFEAEEAVNF 128
+VA+ + + V+F
Sbjct 273 CVVALEFAYNKLDRGLKVDF 292
> Hs5729993
Length=700
Score = 31.2 bits (69), Expect = 0.58, Method: Composition-based stats.
Identities = 16/68 (23%), Positives = 32/68 (47%), Gaps = 2/68 (2%)
Query 62 RVHDLTFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSG 119
++H ++PD P P ++ + + G + VHC AG+GR + + + +
Sbjct 295 QLHFTSWPDFGVPFTPIGMLKFLKKVKTLNPVHAGPIVVHCSAGVGRTGTFIVIDAMMAM 354
Query 120 FEAEEAVN 127
AE+ V+
Sbjct 355 MHAEQKVD 362
> Hs18860859
Length=642
Score = 31.2 bits (69), Expect = 0.58, Method: Composition-based stats.
Identities = 16/68 (23%), Positives = 32/68 (47%), Gaps = 2/68 (2%)
Query 62 RVHDLTFPDGEAP--PAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSG 119
++H ++PD P P ++ + + G + VHC AG+GR + + + +
Sbjct 237 QLHFTSWPDFGVPFTPIGMLKFLKKVKTLNPVHAGPIVVHCSAGVGRTGTFIVIDAMMAM 296
Query 120 FEAEEAVN 127
AE+ V+
Sbjct 297 MHAEQKVD 304
> SPBC17A3.06
Length=330
Score = 30.8 bits (68), Expect = 0.63, Method: Compositional matrix adjust.
Identities = 16/44 (36%), Positives = 26/44 (59%), Gaps = 1/44 (2%)
Query 87 AQAKAEGGVLAVHCVAGLGRGPVLVAVSLI-DSGFEAEEAVNFI 129
A A ++ + VHC AG+ R LVA L+ ++ + EEA++ I
Sbjct 118 AFALSKNAKVLVHCFAGISRSVTLVAAYLMKENNWNTEEALSHI 161
> CE09923
Length=530
Score = 30.8 bits (68), Expect = 0.69, Method: Composition-based stats.
Identities = 17/38 (44%), Positives = 24/38 (63%), Gaps = 1/38 (2%)
Query 20 LDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVV 57
L++PTNDN+ +A + Q+G TD+V YD GP V
Sbjct 157 LESPTNDNVVINVALVDQIGKTDIVIKLIYVYD-GPNV 193
> CE19681
Length=272
Score = 30.8 bits (68), Expect = 0.70, Method: Compositional matrix adjust.
Identities = 23/72 (31%), Positives = 33/72 (45%), Gaps = 11/72 (15%)
Query 48 PPTYDEGPVVLAGIRVHDLTFPDGEAPPAEVIARWRALAAQ---AKAEGGVLAVHCVAGL 104
P TY +G V IR+ D P A + + +A + K GG VHC+AG+
Sbjct 47 PSTYMQG-VDTMKIRIED-------HPYARLNEHFDVVADKIRNVKERGGKTLVHCMAGV 98
Query 105 GRGPVLVAVSLI 116
R LV + L+
Sbjct 99 SRSASLVMIYLV 110
> Hs18576899
Length=96
Score = 30.4 bits (67), Expect = 0.77, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 27/65 (41%), Gaps = 17/65 (26%)
Query 17 CLILDAPTNDNLQAYLAQLVQVGVTDLVRTCPPTYDEGPVVLAGIRVHDLTFPDGEAPPA 76
C +LDAP+ L C P D GP++ IR+ +LT P G P
Sbjct 44 CSLLDAPSTVRL-----------------LCSPGKDNGPLLSDAIRLLELTVPSGHWYPP 86
Query 77 EVIAR 81
E + R
Sbjct 87 EGLVR 91
> Hs18860894
Length=1501
Score = 30.4 bits (67), Expect = 0.86, Method: Composition-based stats.
Identities = 25/102 (24%), Positives = 41/102 (40%), Gaps = 6/102 (5%)
Query 32 LAQLVQVGVTDLVRTCPPT---YDEGPVVLAGIRVHDLT-FPDGEAP--PAEVIARWRAL 85
L Q+ + +L C T Y G +R T +PD P P +A R +
Sbjct 1068 LVQVTLLDTVELATYCVRTFALYKNGSSEKREVRQFQFTAWPDHGVPEHPTPFLAFLRRV 1127
Query 86 AAQAKAEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEEAVN 127
+ G + VHC AG+GR + + + + E+ V+
Sbjct 1128 KTCNPPDAGPMVVHCSAGVGRTGCFIVIDAMLERIKHEKTVD 1169
Score = 28.5 bits (62), Expect = 2.9, Method: Composition-based stats.
Identities = 12/40 (30%), Positives = 22/40 (55%), Gaps = 0/40 (0%)
Query 91 AEGGVLAVHCVAGLGRGPVLVAVSLIDSGFEAEEAVNFIR 130
+ G ++VHC AG+GR V + +S++ E V+ +
Sbjct 1424 GQDGPISVHCSAGVGRTGVFITLSIVLERMRYEGVVDIFQ 1463
> CE18278
Length=591
Score = 30.4 bits (67), Expect = 0.88, Method: Composition-based stats.
Identities = 19/51 (37%), Positives = 25/51 (49%), Gaps = 2/51 (3%)
Query 63 VHDLTFPDGEAPPAEVIARWRALAAQAKAEGGVLAVHCVAGLGRGPVLVAV 113
+H ++PD P +A R L A G V VHC AG+GR VA+
Sbjct 283 LHTKSWPD-RCVPNSTMALLRMLYIVRTASGPV-TVHCSAGIGRTGTFVAI 331
Lambda K H
0.321 0.139 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1246445644
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40