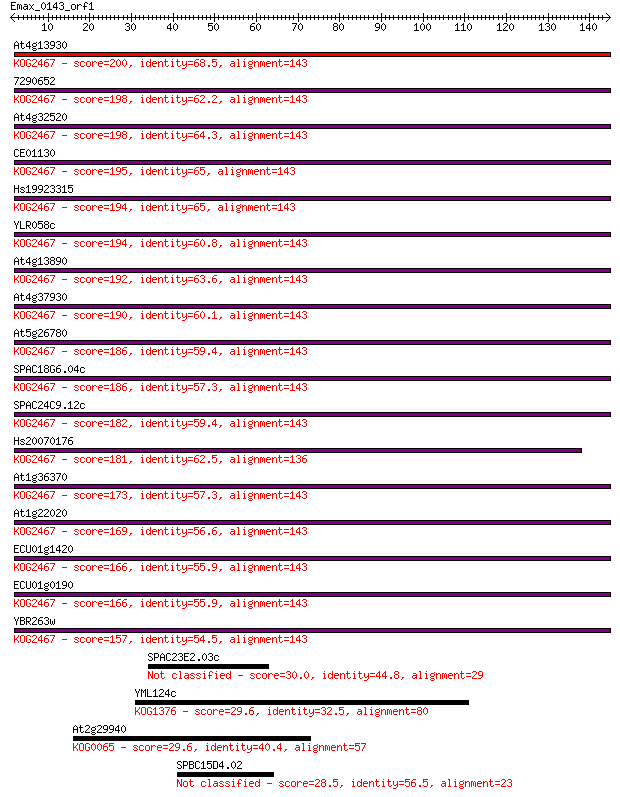
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0143_orf1
Length=144
Score E
Sequences producing significant alignments: (Bits) Value
At4g13930 200 8e-52
7290652 198 3e-51
At4g32520 198 3e-51
CE01130 195 3e-50
Hs19923315 194 3e-50
YLR058c 194 4e-50
At4g13890 192 2e-49
At4g37930 190 9e-49
At5g26780 186 1e-47
SPAC18G6.04c 186 2e-47
SPAC24C9.12c 182 3e-46
Hs20070176 181 4e-46
At1g36370 173 1e-43
At1g22020 169 1e-42
ECU01g1420 166 2e-41
ECU01g0190 166 2e-41
YBR263w 157 8e-39
SPAC23E2.03c 30.0 1.4
YML124c 29.6 2.0
At2g29940 29.6 2.2
SPBC15D4.02 28.5 4.4
> At4g13930
Length=471
Score = 200 bits (508), Expect = 8e-52, Method: Compositional matrix adjust.
Identities = 98/144 (68%), Positives = 113/144 (78%), Gaps = 1/144 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPS-RRISASSLFFESLP 60
VNVQP SGSPANFA + ALLQPHDR MGL L GGHLTHG YT ++ISA+S++FESLP
Sbjct 99 VNVQPYSGSPANFAAYTALLQPHDRIMGLDLPSGGHLTHGYYTSGGKKISATSIYFESLP 158
Query 61 YGLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAH 120
Y +N TG IDY++L+ A FRPKL+ICG SAYPR +Y RFR+IADKVGA L CDMAH
Sbjct 159 YKVNFTTGYIDYDKLEEKALDFRPKLLICGGSAYPRDWDYARFRAIADKVGALLLCDMAH 218
Query 121 FSGFVAAGIFESPFDYCDVVTTTT 144
SG VAA +PF+YCDVVTTTT
Sbjct 219 ISGLVAAQEAANPFEYCDVVTTTT 242
> 7290652
Length=537
Score = 198 bits (504), Expect = 3e-51, Method: Compositional matrix adjust.
Identities = 89/143 (62%), Positives = 113/143 (79%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQP SGSPAN AV+ + +PHDR MGL L DGGHLTHG +TP+++ISA+S+FFES+PY
Sbjct 168 VNVQPYSGSPANLAVYTGVCRPHDRIMGLDLPDGGHLTHGFFTPTKKISATSIFFESMPY 227
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
+NP+TG+IDY++L A FRP++II G S Y RLL+Y RFR I D VGA+L DMAH
Sbjct 228 KVNPETGIIDYDKLAEAAKNFRPQIIIAGISCYSRLLDYARFRQICDDVGAYLMADMAHV 287
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
+G VAAG+ SPF++ D+VTTTT
Sbjct 288 AGIVAAGLIPSPFEWADIVTTTT 310
> At4g32520
Length=462
Score = 198 bits (503), Expect = 3e-51, Method: Compositional matrix adjust.
Identities = 92/143 (64%), Positives = 110/143 (76%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQPLSGSPANFAV+ A+L PHDR MGL L GGHL+HG T RR+S +S++FES+PY
Sbjct 103 VNVQPLSGSPANFAVYTAILSPHDRIMGLDLPHGGHLSHGFMTAKRRVSGTSIYFESMPY 162
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
L+ TG++DY+ L++ ATLFRPKLII G SAY R +Y R R IAD VGAFL DMAH
Sbjct 163 RLDESTGIVDYDMLEKTATLFRPKLIIAGASAYSRDFDYPRMRKIADSVGAFLMMDMAHI 222
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG VAA + PF+YCD+VTTTT
Sbjct 223 SGLVAASVVADPFEYCDIVTTTT 245
> CE01130
Length=484
Score = 195 bits (495), Expect = 3e-50, Method: Compositional matrix adjust.
Identities = 93/143 (65%), Positives = 114/143 (79%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQPLSGSPANFAV+ A++ + R MGL L DGGHLTHG +TP+R++SA+S FF+SLPY
Sbjct 116 VNVQPLSGSPANFAVYTAIVGSNGRIMGLDLPDGGHLTHGFFTPARKVSATSEFFQSLPY 175
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
++P TGLIDY++L++ A LFRPK II G S Y R L+Y+RFR IA K GA+L DMAH
Sbjct 176 KVDPTTGLIDYDKLEQNAMLFRPKAIIAGVSCYARHLDYERFRKIATKAGAYLMSDMAHI 235
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG VAAG+ SPF+Y DVVTTTT
Sbjct 236 SGLVAAGLIPSPFEYSDVVTTTT 258
> Hs19923315
Length=504
Score = 194 bits (494), Expect = 3e-50, Method: Compositional matrix adjust.
Identities = 93/143 (65%), Positives = 109/143 (76%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQP SGSPAN AV+ ALLQPHDR MGL L DGGHLTHG + +RISA+S+FFES+PY
Sbjct 136 VNVQPYSGSPANLAVYTALLQPHDRIMGLDLPDGGHLTHGYMSDVKRISATSIFFESMPY 195
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
LNP+TGLIDY +L A LFRP+LII G SAY RL++Y R R + D+V A L DMAH
Sbjct 196 KLNPKTGLIDYNQLALTARLFRPRLIIAGTSAYARLIDYARMREVCDEVKAHLLADMAHI 255
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG VAA + SPF + D+VTTTT
Sbjct 256 SGLVAAKVIPSPFKHADIVTTTT 278
> YLR058c
Length=469
Score = 194 bits (494), Expect = 4e-50, Method: Compositional matrix adjust.
Identities = 87/143 (60%), Positives = 113/143 (79%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQ LSGSPAN V+ A+++PH+R MGL L DGGHL+HG T +R+ISA S +FES PY
Sbjct 104 VNVQTLSGSPANLQVYQAIMKPHERLMGLYLPDGGHLSHGYATENRKISAVSTYFESFPY 163
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
+NP+TG+IDY+ L++ A L+RPK+++ G SAY RL++YKR R IADK GA+L DMAH
Sbjct 164 RVNPETGIIDYDTLEKNAILYRPKVLVAGTSAYCRLIDYKRMREIADKCGAYLMVDMAHI 223
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG +AAG+ SPF+Y D+VTTTT
Sbjct 224 SGLIAAGVIPSPFEYADIVTTTT 246
> At4g13890
Length=470
Score = 192 bits (488), Expect = 2e-49, Method: Compositional matrix adjust.
Identities = 91/144 (63%), Positives = 112/144 (77%), Gaps = 1/144 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPS-RRISASSLFFESLP 60
VNVQP SGSPANFA + ALLQPHDR MGL L GGH+THG Y+ + ISA+S++FE+LP
Sbjct 99 VNVQPYSGSPANFAAYTALLQPHDRIMGLDLPSGGHITHGYYSSGGKNISATSIYFENLP 158
Query 61 YGLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAH 120
Y ++ +TG IDY++L+ A FRPKLIICG ++YPR +Y RFR++ADKVGAFL CDMAH
Sbjct 159 YKVDSKTGYIDYDKLEEKAMDFRPKLIICGGTSYPREWDYARFRAVADKVGAFLLCDMAH 218
Query 121 FSGFVAAGIFESPFDYCDVVTTTT 144
S VAA PF+YCDVVTT+T
Sbjct 219 NSALVAAQEAADPFEYCDVVTTST 242
> At4g37930
Length=517
Score = 190 bits (482), Expect = 9e-49, Method: Composition-based stats.
Identities = 86/143 (60%), Positives = 108/143 (75%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQPLSGSPANF V+ ALL+PH+R M L L GGHL+HG T +++ISA S+FFE++PY
Sbjct 142 VNVQPLSGSPANFHVYTALLKPHERIMALDLPHGGHLSHGYQTDTKKISAVSIFFETMPY 201
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
L+ TG IDY+++++ ATLFRPKLI+ G SAY RL +Y R R + +K A + DMAH
Sbjct 202 RLDESTGYIDYDQMEKSATLFRPKLIVAGASAYARLYDYARIRKVCNKQKAVMLADMAHI 261
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG VAA + SPFDY DVVTTTT
Sbjct 262 SGLVAANVIPSPFDYADVVTTTT 284
> At5g26780
Length=532
Score = 186 bits (473), Expect = 1e-47, Method: Composition-based stats.
Identities = 85/143 (59%), Positives = 107/143 (74%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQ LSGSPANF V+ ALL+PH+R M L L GGHL+HG T +++ISA S+FFE++PY
Sbjct 134 VNVQSLSGSPANFQVYTALLKPHERIMALDLPHGGHLSHGYQTDTKKISAVSIFFETMPY 193
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
L+ TG IDY++L++ A LFRPKLI+ G SAY RL +Y R R + +K A + DMAH
Sbjct 194 RLDENTGYIDYDQLEKSAVLFRPKLIVAGASAYARLYDYARIRKVCNKQKAVMLADMAHI 253
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG VAAG+ SPF+Y DVVTTTT
Sbjct 254 SGLVAAGVIPSPFEYADVVTTTT 276
> SPAC18G6.04c
Length=472
Score = 186 bits (471), Expect = 2e-47, Method: Compositional matrix adjust.
Identities = 82/143 (57%), Positives = 106/143 (74%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQP SGSPAN + A+++PHDR MGL L GGHL+HG TP + ISA S +F ++PY
Sbjct 105 VNVQPHSGSPANLQAYQAVMKPHDRLMGLDLPHGGHLSHGFSTPQKAISAVSTYFSTMPY 164
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
+N +TG+IDY+ L++ A FRPK+I+ G SAY RL++YKR R I + A+L CDMAH
Sbjct 165 NVNKETGIIDYDSLEKAAIQFRPKVIVAGASAYARLVDYKRMRKITEMCNAYLLCDMAHI 224
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG VAAG+ SPF+Y D+VTTTT
Sbjct 225 SGLVAAGVIPSPFEYADIVTTTT 247
> SPAC24C9.12c
Length=467
Score = 182 bits (461), Expect = 3e-46, Method: Compositional matrix adjust.
Identities = 85/143 (59%), Positives = 107/143 (74%), Gaps = 0/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQ LSGSPAN V+ A++ PH R MGL L GGHL+HG T +++ISA S +FES+PY
Sbjct 99 VNVQCLSGSPANMQVYQAIMPPHGRLMGLDLPSGGHLSHGYQTDTKKISAVSTYFESMPY 158
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
++P TGLIDY+ L+ A LFRPK+++ G SAY RL++Y R R IAD V A+L DMAH
Sbjct 159 RVDPNTGLIDYDMLEHDAQLFRPKILVAGTSAYCRLIDYARMRQIADSVNAYLVVDMAHI 218
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
SG V+AG+ SPF+Y DVVTTTT
Sbjct 219 SGLVSAGVIPSPFEYADVVTTTT 241
> Hs20070176
Length=483
Score = 181 bits (460), Expect = 4e-46, Method: Compositional matrix adjust.
Identities = 85/136 (62%), Positives = 103/136 (75%), Gaps = 0/136 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQP SGSPANFAV+ AL++PH R MGL L DGGHLTHG T ++ISA+S+FFES+PY
Sbjct 113 VNVQPYSGSPANFAVYTALVEPHGRIMGLDLPDGGHLTHGFMTDKKKISATSIFFESMPY 172
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
+NP TG I+Y++L+ A LF PKLII G S Y R L Y R R IAD+ GA+L DMAH
Sbjct 173 KVNPDTGYINYDQLEENARLFHPKLIIAGTSCYSRNLEYARLRKIADENGAYLMADMAHI 232
Query 122 SGFVAAGIFESPFDYC 137
SG VAAG+ SPF++C
Sbjct 233 SGLVAAGVVPSPFEHC 248
> At1g36370
Length=578
Score = 173 bits (438), Expect = 1e-43, Method: Compositional matrix adjust.
Identities = 82/144 (56%), Positives = 105/144 (72%), Gaps = 1/144 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPS-RRISASSLFFESLP 60
VNVQP S + ANFAV+ LL P +R MGL GGH++HG TP ++ISA+S+FFES P
Sbjct 205 VNVQPYSCTSANFAVYTGLLLPGERIMGLDSPSGGHMSHGYCTPGGKKISAASIFFESFP 264
Query 61 YGLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAH 120
Y +NPQTG IDY++L+ A +RPK++ICG S+YPR ++ R R IADK GA L CDMAH
Sbjct 265 YKVNPQTGYIDYDKLEDKALDYRPKILICGGSSYPRDWDFARVRQIADKCGAVLMCDMAH 324
Query 121 FSGFVAAGIFESPFDYCDVVTTTT 144
SG VA +PFD+CD+VT+TT
Sbjct 325 ISGLVATKECSNPFDHCDIVTSTT 348
> At1g22020
Length=599
Score = 169 bits (429), Expect = 1e-42, Method: Compositional matrix adjust.
Identities = 81/144 (56%), Positives = 104/144 (72%), Gaps = 1/144 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPS-RRISASSLFFESLP 60
VNVQP S + ANFAVF LL P +R MGL GGH++HG YTP +++S +S+FFES P
Sbjct 229 VNVQPYSCTSANFAVFTGLLMPGERIMGLDSPSGGHMSHGYYTPGGKKVSGASIFFESFP 288
Query 61 YGLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAH 120
Y ++P+TG IDY++L+ A +RPK++ICG S+YPR + RFR IADK GA L DMA
Sbjct 289 YKVDPRTGYIDYDKLEEKALDYRPKILICGGSSYPRDWEFPRFRHIADKCGAVLMFDMAQ 348
Query 121 FSGFVAAGIFESPFDYCDVVTTTT 144
SG VAA +PFDYCD+VT+TT
Sbjct 349 ISGLVAAKESPNPFDYCDIVTSTT 372
> ECU01g1420
Length=460
Score = 166 bits (419), Expect = 2e-41, Method: Compositional matrix adjust.
Identities = 80/143 (55%), Positives = 107/143 (74%), Gaps = 1/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQP SGSPANFA++ A++ P R MGL L GGHLTHG T +R+ISASS++F+S PY
Sbjct 101 VNVQPYSGSPANFAIYTAVVPPGGRIMGLDLPSGGHLTHGYKTKTRKISASSVYFDSRPY 160
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
+ GLIDYE L++ T F P ++ICG+SAY R ++YKR +SIA + GAFL+ D++H
Sbjct 161 TVG-SNGLIDYEGLEKTFTDFLPHILICGYSAYSRDIDYKRLQSIAGRNGAFLFADISHI 219
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
S VA+G+ SPF++CD+V TTT
Sbjct 220 SPLVASGLMNSPFEHCDIVMTTT 242
> ECU01g0190
Length=460
Score = 166 bits (419), Expect = 2e-41, Method: Compositional matrix adjust.
Identities = 80/143 (55%), Positives = 107/143 (74%), Gaps = 1/143 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPY 61
VNVQP SGSPANFA++ A++ P R MGL L GGHLTHG T +R+ISASS++F+S PY
Sbjct 101 VNVQPYSGSPANFAIYTAVVPPGGRIMGLDLPSGGHLTHGYKTKTRKISASSVYFDSRPY 160
Query 62 GLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAHF 121
+ GLIDYE L++ T F P ++ICG+SAY R ++YKR +SIA + GAFL+ D++H
Sbjct 161 TVG-SNGLIDYEGLEKTFTDFLPHILICGYSAYSRDIDYKRLQSIAGRNGAFLFADISHI 219
Query 122 SGFVAAGIFESPFDYCDVVTTTT 144
S VA+G+ SPF++CD+V TTT
Sbjct 220 SPLVASGLMNSPFEHCDIVMTTT 242
> YBR263w
Length=565
Score = 157 bits (396), Expect = 8e-39, Method: Compositional matrix adjust.
Identities = 78/144 (54%), Positives = 101/144 (70%), Gaps = 1/144 (0%)
Query 2 VNVQPLSGSPANFAVFMALLQPHDRFMGLGLNDGGHLTHGAYTPSRR-ISASSLFFESLP 60
VNVQPLSG+PAN V+ A++ +R MGL L DGGHL+HG S IS S +F+S+P
Sbjct 195 VNVQPLSGAPANLYVYSAIMNVGERLMGLDLPDGGHLSHGYQLKSGTPISFISKYFQSMP 254
Query 61 YGLNPQTGLIDYEELDRLATLFRPKLIICGHSAYPRLLNYKRFRSIADKVGAFLWCDMAH 120
Y ++ TGLIDY+ L LA FRPK+I+ G SAY RL++Y RF+ I+ GA+L DMAH
Sbjct 255 YHVDHTTGLIDYDNLQVLAKAFRPKVIVAGTSAYSRLIDYARFKEISQGCGAYLMSDMAH 314
Query 121 FSGFVAAGIFESPFDYCDVVTTTT 144
SG VAA + SPF++ D+VTTTT
Sbjct 315 ISGLVAANVVPSPFEHSDIVTTTT 338
> SPAC23E2.03c
Length=569
Score = 30.0 bits (66), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 13/29 (44%), Positives = 19/29 (65%), Gaps = 0/29 (0%)
Query 34 DGGHLTHGAYTPSRRISASSLFFESLPYG 62
D H H A +PSRR S+++L+ +LP G
Sbjct 491 DQVHNNHCAVSPSRRSSSNALYLATLPMG 519
> YML124c
Length=445
Score = 29.6 bits (65), Expect = 2.0, Method: Composition-based stats.
Identities = 26/92 (28%), Positives = 37/92 (40%), Gaps = 13/92 (14%)
Query 31 GLNDGGHLTHGAYTPSRRISASSLFFESLPYG--------LNPQTGLIDYEELDRLATLF 82
G+ + GHL G P S FF YG ++ + +ID R LF
Sbjct 29 GIKEDGHLEDGLSKPKGGEEGFSTFFHETGYGKFVPRAIYVDLEPNVIDEVRTGRFKELF 88
Query 83 RPKLIICGH----SAYPRLLNYKRFRSIADKV 110
P+ +I G + Y R +Y R I D+V
Sbjct 89 HPEQLINGKEDAANNYAR-GHYTVGREIVDEV 119
> At2g29940
Length=1443
Score = 29.6 bits (65), Expect = 2.2, Method: Compositional matrix adjust.
Identities = 23/61 (37%), Positives = 31/61 (50%), Gaps = 9/61 (14%)
Query 16 VFMALLQPH----DRFMGLGLNDGGHLTHGAYTPSRRISASSLFFESLPYGLNPQTGLID 71
V MALLQP D F L L G++ + P + A FFESL + L P+ G+ D
Sbjct 403 VLMALLQPAPETFDLFDDLILLSEGYMVYQG--PREDVIA---FFESLGFRLPPRKGVAD 457
Query 72 Y 72
+
Sbjct 458 F 458
> SPBC15D4.02
Length=547
Score = 28.5 bits (62), Expect = 4.4, Method: Compositional matrix adjust.
Identities = 13/23 (56%), Positives = 15/23 (65%), Gaps = 0/23 (0%)
Query 41 GAYTPSRRISASSLFFESLPYGL 63
G +T SRRI SSL E P+GL
Sbjct 212 GTFTASRRIPVSSLLSEPAPHGL 234
Lambda K H
0.326 0.141 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1675978996
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40