bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
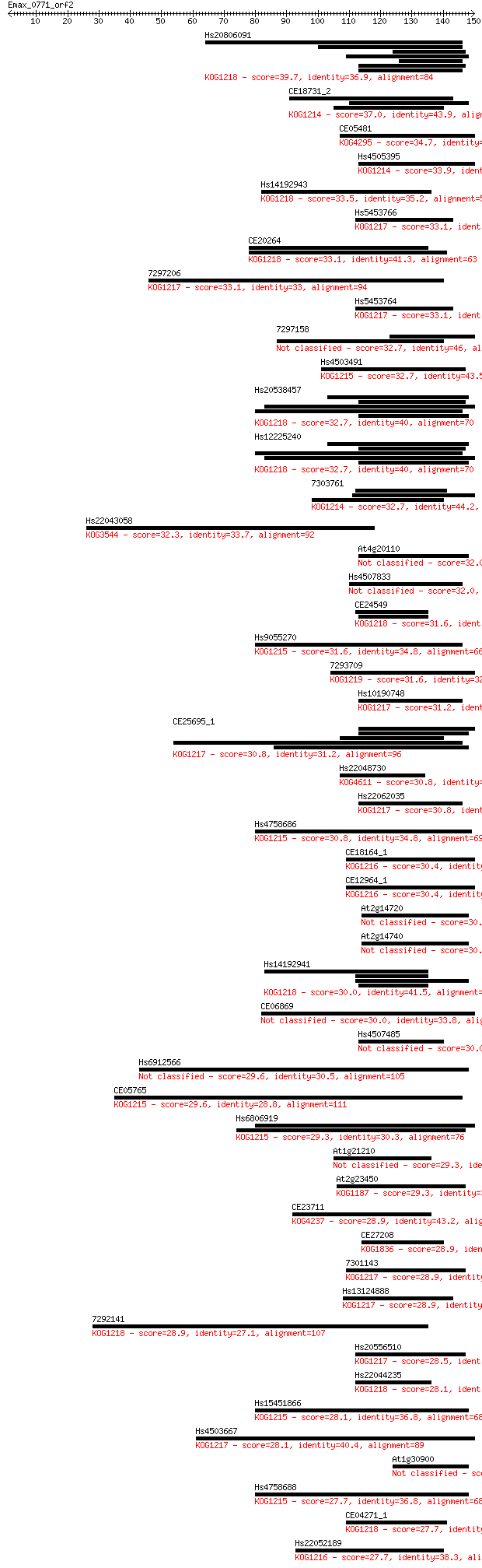
Query= Emax_0771_orf2
Length=149
Score E
Sequences producing significant alignments: (Bits) Value
Hs20806091 39.7 0.002
CE18731_2 37.0 0.013
CE05481 34.7 0.063
Hs4505395 33.9 0.11
Hs14192943 33.5 0.15
Hs5453766 33.1 0.18
CE20264 33.1 0.18
7297206 33.1 0.21
Hs5453764 33.1 0.21
7297158 32.7 0.24
Hs4503491 32.7 0.28
Hs20538457 32.7 0.28
Hs12225240 32.7 0.28
7303761 32.7 0.30
Hs22043058 32.3 0.30
At4g20110 32.0 0.45
Hs4507833 32.0 0.48
CE24549 31.6 0.54
Hs9055270 31.6 0.61
7293709 31.6 0.61
Hs10190748 31.2 0.86
CE25695_1 30.8 0.91
Hs22048730 30.8 1.0
Hs22062035 30.8 1.0
Hs4758686 30.8 1.0
CE18164_1 30.4 1.3
CE12964_1 30.4 1.3
At2g14720 30.4 1.3
At2g14740 30.4 1.3
Hs14192941 30.0 1.5
CE06869 30.0 1.8
Hs4507485 30.0 1.9
Hs6912566 29.6 2.1
CE05765 29.6 2.2
Hs6806919 29.3 2.7
At1g21210 29.3 2.7
At2g23450 29.3 3.0
CE23711 28.9 3.5
CE27208 28.9 3.7
7301143 28.9 3.7
Hs13124888 28.9 3.8
7292141 28.9 4.2
Hs20556510 28.5 5.4
Hs22044235 28.1 5.8
Hs15451866 28.1 6.1
Hs4503667 28.1 6.8
At1g30900 27.7 7.8
Hs4758688 27.7 8.1
CE04271_1 27.7 9.1
Hs22052189 27.7 9.2
> Hs20806091
Length=2551
Score = 39.7 bits (91), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 29/92 (31%), Positives = 39/92 (42%), Gaps = 10/92 (10%)
Query 64 AEIEKCGEFVEWDPPMNGDCVRGGTHTR-------YRQNCPDRKEVRVCGAFDCSSCSVN 116
+E++ C N +C++ GT T + N D E+ C C N
Sbjct 866 SEMDPCTGLTPGGCSRNAECIKTGTGTHTCVCQQGWTGNGRDCSEINNCLLPSAGGCHDN 925
Query 117 ATCDPIGAS---CECKPGFRGNGKTCEAFNPC 145
A+C +G CECK GFRGNG CE C
Sbjct 926 ASCLYVGPGQNECECKKGFRGNGIDCEPITSC 957
Score = 33.9 bits (76), Expect = 0.11, Method: Compositional matrix adjust.
Identities = 14/23 (60%), Positives = 16/23 (69%), Gaps = 0/23 (0%)
Query 124 ASCECKPGFRGNGKTCEAFNPCE 146
ASC+C GF+GNG C A N CE
Sbjct 1454 ASCKCAAGFQGNGTICTAINACE 1476
Score = 33.9 bits (76), Expect = 0.13, Method: Compositional matrix adjust.
Identities = 18/48 (37%), Positives = 26/48 (54%), Gaps = 3/48 (6%)
Query 100 KEVRVCGAFDCSSCSVNATCDPIG--ASCECKPGFRGNGKTCEAFNPC 145
K+ CG + C ++ATC+ ASC CK G+ G+G C +PC
Sbjct 825 KQTSACGPY-VQFCHIHATCEYSNGTASCICKAGYEGDGTLCSEMDPC 871
Score = 30.4 bits (67), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 11/20 (55%), Positives = 15/20 (75%), Gaps = 0/20 (0%)
Query 126 CECKPGFRGNGKTCEAFNPC 145
C+C P +RG+GK C+ NPC
Sbjct 222 CKCLPNYRGDGKYCDPINPC 241
Score = 30.4 bits (67), Expect = 1.2, Method: Compositional matrix adjust.
Identities = 15/41 (36%), Positives = 22/41 (53%), Gaps = 2/41 (4%)
Query 109 DCSSCSVNATCDPIG--ASCECKPGFRGNGKTCEAFNPCED 147
D C+ A C G SC C+ G++G+G +C +PC D
Sbjct 2089 DNGGCAKVARCSQKGTKVSCSCQKGYKGDGHSCTEIDPCAD 2129
Score = 29.6 bits (65), Expect = 2.1, Method: Compositional matrix adjust.
Identities = 14/37 (37%), Positives = 22/37 (59%), Gaps = 3/37 (8%)
Query 113 CSVNATCDPIG---ASCECKPGFRGNGKTCEAFNPCE 146
C +A C +G SC C+ G+RG+G+ C +PC+
Sbjct 246 CHPHAHCTYLGPNRHSCTCQEGYRGDGQVCLPVDPCQ 282
Score = 29.6 bits (65), Expect = 2.3, Method: Compositional matrix adjust.
Identities = 15/36 (41%), Positives = 18/36 (50%), Gaps = 3/36 (8%)
Query 113 CSVNATCDPIG---ASCECKPGFRGNGKTCEAFNPC 145
C NA C G A+C C P + G+GK C N C
Sbjct 1524 CDKNAECTQTGPNQAACNCLPAYTGDGKVCTLINVC 1559
> CE18731_2
Length=1256
Score = 37.0 bits (84), Expect = 0.013, Method: Compositional matrix adjust.
Identities = 23/55 (41%), Positives = 30/55 (54%), Gaps = 7/55 (12%)
Query 91 RYRQNCPDRKEVRVCGAFDCSSCSVNATCDP---IGASCECKPGFRGNGKTCEAF 142
++ N P R + CG++ C VNA C P G+ C CK GF GNG TCE+
Sbjct 495 KHNANIPQRGG-QACGSY---VCDVNAECMPEPSGGSECVCKAGFSGNGVTCESL 545
Score = 31.2 bits (69), Expect = 0.83, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 25/42 (59%), Gaps = 4/42 (9%)
Query 110 CSSCSVNATC--DPIGAS--CECKPGFRGNGKTCEAFNPCED 147
C++CS++A C +P + C+C G+ GNG C + + C D
Sbjct 875 CTNCSIHAYCAQNPTSGAYQCKCNAGYNGNGHLCVSMSSCLD 916
Score = 27.7 bits (60), Expect = 7.4, Method: Compositional matrix adjust.
Identities = 16/37 (43%), Positives = 17/37 (45%), Gaps = 2/37 (5%)
Query 105 CGAFDCSSCSVNATCDPI--GASCECKPGFRGNGKTC 139
C D SC N+ C G C C GF GNGK C
Sbjct 32 CSQRDDKSCHANSVCQDFEGGFCCNCDTGFYGNGKEC 68
> CE05481
Length=838
Score = 34.7 bits (78), Expect = 0.063, Method: Compositional matrix adjust.
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 2/43 (4%)
Query 107 AFDCSSCSVNATCDPIGASCECKPGFRGNGKTCEAFNPCEDTP 149
+ DC CS++ATC + C+CK G+ G+G C N C P
Sbjct 247 SIDCKDCSMHATC--MNGVCQCKEGYEGDGFNCTDVNECLRRP 287
> Hs4505395
Length=1247
Score = 33.9 bits (76), Expect = 0.11, Method: Compositional matrix adjust.
Identities = 17/40 (42%), Positives = 22/40 (55%), Gaps = 3/40 (7%)
Query 113 CSVNATCDP---IGASCECKPGFRGNGKTCEAFNPCEDTP 149
C NA C P +CEC GFRG+G+TC + C + P
Sbjct 679 CDTNAACRPGPRTQFTCECSIGFRGDGRTCYDIDECSEQP 718
> Hs14192943
Length=1140
Score = 33.5 bits (75), Expect = 0.15, Method: Compositional matrix adjust.
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Query 82 DCVRGGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRGN 135
DC+ G T + CP + + C +C+ N TC+PI SC+C PG+ G+
Sbjct 646 DCLPGFTGALCNEVCPSGRFGKNCAGI--CTCTNNGTCNPIDRSCQCYPGWIGS 697
> Hs5453766
Length=816
Score = 33.1 bits (74), Expect = 0.18, Method: Compositional matrix adjust.
Identities = 17/33 (51%), Positives = 21/33 (63%), Gaps = 2/33 (6%)
Query 112 SCSVNATCDPI--GASCECKPGFRGNGKTCEAF 142
+C NA C G +C CKPG+ GNG TC+AF
Sbjct 492 NCDENALCFNTVGGHNCVCKPGYTGNGTTCKAF 524
> CE20264
Length=1111
Score = 33.1 bits (74), Expect = 0.18, Method: Compositional matrix adjust.
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 3/59 (5%)
Query 78 PMNGDCV-RGGTHTRY-RQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRG 134
P G+C+ G H + + CPD K C A DC C+ +TCD I C C G G
Sbjct 699 PTTGECICEPGYHGKTCSEKCPDGKYGYGC-ALDCPKCASGSTCDHINGLCICPAGLEG 756
Score = 27.7 bits (60), Expect = 9.1, Method: Compositional matrix adjust.
Identities = 22/66 (33%), Positives = 28/66 (42%), Gaps = 7/66 (10%)
Query 78 PMNGDCVR---GGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRG 134
P NG C+ G + NCP C SC+ CDP C C+PG+
Sbjct 655 PKNGVCLSCPPGSSGIHCEHNCPAGSYGDGCQQV--CSCADGHGCDPTTGECICEPGY-- 710
Query 135 NGKTCE 140
+GKTC
Sbjct 711 HGKTCS 716
> 7297206
Length=3396
Score = 33.1 bits (74), Expect = 0.21, Method: Compositional matrix adjust.
Identities = 31/116 (26%), Positives = 44/116 (37%), Gaps = 25/116 (21%)
Query 46 RYNSKCGPVEVRECNMDDAEIEKCGEFVEWDPPMNGDCVRGGTHTRYRQN----CPDRKE 101
R K G V EC + + G ++ M +C +G + +Q CP
Sbjct 3097 RTTPKVGASSVEECTL---PVCSAGTYLNATQNMCIECRKGYYQSESQQTSCLQCPPNHS 3153
Query 102 VRVCGAFDCSSCS--------------VNATCDPIGAS----CECKPGFRGNGKTC 139
++ GA S C+ VNA C + + CECKPGF G G C
Sbjct 3154 TKITGATSKSECTNPCEHIAEGKPHCDVNAYCIMVPETSDFKCECKPGFNGTGMAC 3209
> Hs5453764
Length=810
Score = 33.1 bits (74), Expect = 0.21, Method: Compositional matrix adjust.
Identities = 17/33 (51%), Positives = 19/33 (57%), Gaps = 2/33 (6%)
Query 112 SCSVNATCDPI--GASCECKPGFRGNGKTCEAF 142
+C NA C G SC CKPG+ GNG C AF
Sbjct 486 NCDENAICTNTVQGHSCTCKPGYVGNGTICRAF 518
> 7297158
Length=1024
Score = 32.7 bits (73), Expect = 0.24, Method: Compositional matrix adjust.
Identities = 15/28 (53%), Positives = 19/28 (67%), Gaps = 1/28 (3%)
Query 123 GASCE-CKPGFRGNGKTCEAFNPCEDTP 149
GA C+ C G+ G+G+TC NPC DTP
Sbjct 324 GAQCDSCPAGYEGDGRTCSLRNPCLDTP 351
Score = 28.5 bits (62), Expect = 4.8, Method: Compositional matrix adjust.
Identities = 21/56 (37%), Positives = 25/56 (44%), Gaps = 5/56 (8%)
Query 87 GTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATC---DPIGASCECKPGFRGNGKTC 139
G T +C DR V +C D + C NA C D I C C G+ GNG C
Sbjct 472 GYVTNGTYSCLDRSSVFMCP--DGTVCDRNAVCLRMDNIRHKCHCNVGWAGNGLIC 525
> Hs4503491
Length=1207
Score = 32.7 bits (73), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 20/50 (40%), Positives = 27/50 (54%), Gaps = 4/50 (8%)
Query 101 EVRVCGAFDCS--SCSVNATCDPIG--ASCECKPGFRGNGKTCEAFNPCE 146
E+ V DC+ CS+ A C G A+C+C GF G+GK C + CE
Sbjct 826 EIMVSDQDDCAPVGCSMYARCISEGEDATCQCLKGFAGDGKLCSDIDECE 875
> Hs20538457
Length=2570
Score = 32.7 bits (73), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 20/52 (38%), Positives = 26/52 (50%), Gaps = 7/52 (13%)
Query 103 RVCGAFDC-----SSCSVNATCDPIG--ASCECKPGFRGNGKTCEAFNPCED 147
RVC D CS +A C +G +C C P + G+G +C A NPC D
Sbjct 2087 RVCTVADLCQDGHGGCSEHANCSQVGTMVTCTCLPDYEGDGWSCRARNPCTD 2138
Score = 32.3 bits (72), Expect = 0.33, Method: Compositional matrix adjust.
Identities = 16/38 (42%), Positives = 23/38 (60%), Gaps = 4/38 (10%)
Query 113 CSVNATCDPIG---ASCECKPGFRGNG-KTCEAFNPCE 146
C ++A C P G SC C+ G+ G+G +TCE +PC
Sbjct 1508 CHIHAECIPTGPQQVSCSCREGYSGDGIRTCELLDPCS 1545
Score = 30.4 bits (67), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 22/67 (32%), Positives = 26/67 (38%), Gaps = 9/67 (13%)
Query 83 CVRGGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRGNGKTCEAF 142
C G T Q P +E+R C N C SC C PG+ G C A
Sbjct 183 CFAGYTGPHCDQELPVCQELR---------CPQNTQCSAEAPSCRCLPGYTQQGSECRAP 233
Query 143 NPCEDTP 149
NPC +P
Sbjct 234 NPCWPSP 240
Score = 30.0 bits (66), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 26/79 (32%), Positives = 34/79 (43%), Gaps = 16/79 (20%)
Query 80 NGDCVRGGTHTRYRQNCPDRK----EVRVCGAFD------CSSCSVNATCDPIG---ASC 126
N +CV G T + C K + RVC A D C +A C +G + C
Sbjct 876 NAECVPGSLGTHH---CTCHKGWSGDGRVCVAIDECELDMRGGCHTDALCSYVGPGQSRC 932
Query 127 ECKPGFRGNGKTCEAFNPC 145
CK GF G+G C +PC
Sbjct 933 TCKLGFAGDGYQCSPIDPC 951
Score = 28.5 bits (62), Expect = 5.0, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 20/35 (57%), Gaps = 1/35 (2%)
Query 113 CSVNATCDPIGASCECKPGFRGNGKTCEAFNPCED 147
C+ A C G SCEC G+ G+G+ C + C+D
Sbjct 2064 CAPEAVCR-AGNSCECSLGYEGDGRVCTVADLCQD 2097
> Hs12225240
Length=2570
Score = 32.7 bits (73), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 20/52 (38%), Positives = 26/52 (50%), Gaps = 7/52 (13%)
Query 103 RVCGAFDC-----SSCSVNATCDPIG--ASCECKPGFRGNGKTCEAFNPCED 147
RVC D CS +A C +G +C C P + G+G +C A NPC D
Sbjct 2087 RVCTVADLCQDGHGGCSEHANCSQVGTMVTCTCLPDYEGDGWSCRARNPCTD 2138
Score = 32.3 bits (72), Expect = 0.33, Method: Compositional matrix adjust.
Identities = 16/38 (42%), Positives = 23/38 (60%), Gaps = 4/38 (10%)
Query 113 CSVNATCDPIG---ASCECKPGFRGNG-KTCEAFNPCE 146
C ++A C P G SC C+ G+ G+G +TCE +PC
Sbjct 1508 CHIHAECIPTGPQQVSCSCREGYSGDGIRTCELLDPCS 1545
Score = 30.8 bits (68), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 26/79 (32%), Positives = 34/79 (43%), Gaps = 16/79 (20%)
Query 80 NGDCVRGGTHTRYRQNCPDRK----EVRVCGAFD------CSSCSVNATCDPIG---ASC 126
N +CV G T + C K + RVC A D C +A C +G + C
Sbjct 876 NAECVPGSLGTHH---CTCHKGWSGDGRVCVAIDECELDVGGGCHTDALCSYVGPGQSRC 932
Query 127 ECKPGFRGNGKTCEAFNPC 145
CK GF G+G C +PC
Sbjct 933 TCKLGFAGDGYQCSPIDPC 951
Score = 30.4 bits (67), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 22/67 (32%), Positives = 26/67 (38%), Gaps = 9/67 (13%)
Query 83 CVRGGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRGNGKTCEAF 142
C G T Q P +E+R C N C SC C PG+ G C A
Sbjct 183 CFAGYTGPHCDQELPVCQELR---------CPQNTQCSAEAPSCRCLPGYTQQGSECRAP 233
Query 143 NPCEDTP 149
NPC +P
Sbjct 234 NPCWPSP 240
Score = 28.5 bits (62), Expect = 5.1, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 20/35 (57%), Gaps = 1/35 (2%)
Query 113 CSVNATCDPIGASCECKPGFRGNGKTCEAFNPCED 147
C+ A C G SCEC G+ G+G+ C + C+D
Sbjct 2064 CAPEAVCR-AGNSCECSLGYEGDGRVCTVADLCQD 2097
> 7303761
Length=1352
Score = 32.7 bits (73), Expect = 0.30, Method: Compositional matrix adjust.
Identities = 15/33 (45%), Positives = 21/33 (63%), Gaps = 4/33 (12%)
Query 112 SCSVNATCDPIGAS----CECKPGFRGNGKTCE 140
+C +NATC+ G C C+PGFRG+G C+
Sbjct 797 NCHINATCNWYGQELRHICTCQPGFRGDGYNCD 829
Score = 30.8 bits (68), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 15/43 (34%), Positives = 23/43 (53%), Gaps = 4/43 (9%)
Query 111 SSCSVNATC----DPIGASCECKPGFRGNGKTCEAFNPCEDTP 149
++C ++ATC DP C+C GF+G+G C C + P
Sbjct 963 NNCGIHATCEPTEDPANYECQCIAGFKGDGYVCIEEQNCLNNP 1005
Score = 27.7 bits (60), Expect = 9.5, Method: Compositional matrix adjust.
Identities = 20/49 (40%), Positives = 24/49 (48%), Gaps = 7/49 (14%)
Query 98 DRKEVRVCGAFD-CSS----CSVNATCDPI--GASCECKPGFRGNGKTC 139
D + V VC D C++ C NA CD G +C C GF GNG C
Sbjct 582 DERGVEVCLDIDECATGSHVCDENAVCDNTEGGFNCYCTEGFEGNGYRC 630
> Hs22043058
Length=3631
Score = 32.3 bits (72), Expect = 0.30, Method: Composition-based stats.
Identities = 31/93 (33%), Positives = 40/93 (43%), Gaps = 29/93 (31%)
Query 26 IECNDQWTPWTMCDTNRVQERYNSKCGPVEVRECNMDDAEIEKCGEFVEWDPPMNGDCVR 85
IE N+Q TP +T R + ++ GP C+MD PM G+C
Sbjct 739 IEENEQETPAKQKET-RKEINADTTYGP-----CSMD---------------PMEGEC-- 775
Query 86 GGTHT-RYRQNCPDRKEVRVCGAFDCSSCSVNA 117
HT ++ N KE RVC F C SC NA
Sbjct 776 -QDHTLKWHYN----KEERVCQQFWCGSCGGNA 803
> At4g20110
Length=625
Score = 32.0 bits (71), Expect = 0.45, Method: Compositional matrix adjust.
Identities = 13/35 (37%), Positives = 21/35 (60%), Gaps = 0/35 (0%)
Query 113 CSVNATCDPIGASCECKPGFRGNGKTCEAFNPCED 147
+ +A D + C+C GF+G+G TCE N C++
Sbjct 486 LTFSACSDSVSTGCKCPEGFQGDGLTCEDINECKE 520
> Hs4507833
Length=640
Score = 32.0 bits (71), Expect = 0.48, Method: Compositional matrix adjust.
Identities = 17/38 (44%), Positives = 21/38 (55%), Gaps = 2/38 (5%)
Query 110 CSSCSVNATC--DPIGASCECKPGFRGNGKTCEAFNPC 145
CS C NATC D +C C+ GF G+G TC + C
Sbjct 32 CSECHSNATCTEDEAVTTCTCQEGFTGDGLTCVDLDEC 69
> CE24549
Length=1664
Score = 31.6 bits (70), Expect = 0.54, Method: Compositional matrix adjust.
Identities = 14/23 (60%), Positives = 17/23 (73%), Gaps = 0/23 (0%)
Query 112 SCSVNATCDPIGASCECKPGFRG 134
SC ATCD + SCEC+PG+RG
Sbjct 990 SCQNGATCDSVTGSCECRPGWRG 1012
Score = 28.9 bits (63), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 12/22 (54%), Positives = 14/22 (63%), Gaps = 0/22 (0%)
Query 113 CSVNATCDPIGASCECKPGFRG 134
C A CDPI C C+PG+RG
Sbjct 1216 CENGALCDPISGHCSCQPGWRG 1237
> Hs9055270
Length=4599
Score = 31.6 bits (70), Expect = 0.61, Method: Compositional matrix adjust.
Identities = 23/73 (31%), Positives = 34/73 (46%), Gaps = 7/73 (9%)
Query 80 NGDCVRGGTHTRYRQNCPDRKEVRVCGAFDC-----SSCSVNATCDPIGASCECKPGF-- 132
NG C+ G + +C D + R C +C S CS + P+ C+C PGF
Sbjct 2900 NGRCIPSGGLCDNKDDCGDGSDERNCHINECLSKKVSGCSQDCQDLPVSYKCKCWPGFQL 2959
Query 133 RGNGKTCEAFNPC 145
+ +GKTC + C
Sbjct 2960 KDDGKTCVDIDEC 2972
> 7293709
Length=4643
Score = 31.6 bits (70), Expect = 0.61, Method: Composition-based stats.
Identities = 15/47 (31%), Positives = 23/47 (48%), Gaps = 3/47 (6%)
Query 104 VCGAFDCSSCSVNATCDPIGASCECKPGFRGNGKTCEA-FNPCEDTP 149
+CG+ C++ + D C C+P R +GK CE +PC P
Sbjct 4132 ICGSQPCANSGICKELDTDVFECACQP--RYSGKHCEIDLDPCSSGP 4176
> Hs10190748
Length=999
Score = 31.2 bits (69), Expect = 0.86, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 19/35 (54%), Gaps = 2/35 (5%)
Query 113 CSVNATCD--PIGASCECKPGFRGNGKTCEAFNPC 145
C +A C P C CKPG++G G+ CE + C
Sbjct 56 CHADALCQNTPTSYKCSCKPGYQGEGRQCEDIDEC 90
> CE25695_1
Length=2353
Score = 30.8 bits (68), Expect = 0.91, Method: Compositional matrix adjust.
Identities = 16/39 (41%), Positives = 23/39 (58%), Gaps = 2/39 (5%)
Query 113 CSVNATCDPIGASCECKPGFRGNGKTCEAFNPCE--DTP 149
C +A+C + C+CK G+ G+G TC N C+ DTP
Sbjct 314 CDRHASCHIVLDICDCKTGYTGDGITCHDINECDAKDTP 352
Score = 30.8 bits (68), Expect = 0.91, Method: Compositional matrix adjust.
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query 113 CSVNATCDPIGA--SCECKPGFRGNGKTCEAFNPCED 147
CS+NA C + SC CK G+RG+G C N C++
Sbjct 1595 CSLNANCVNMNGTFSCSCKQGYRGDGFMCTDINECDE 1631
Score = 30.4 bits (67), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 16/33 (48%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Query 107 AFDCSSCSVNATCDPIGASCECKPGFRGNGKTC 139
A +CS C NA C G +C+C PG+ GNG C
Sbjct 2145 ASNCSQCDANAHC-VGGTTCKCNPGYFGNGLCC 2176
Score = 29.6 bits (65), Expect = 2.3, Method: Compositional matrix adjust.
Identities = 23/95 (24%), Positives = 40/95 (42%), Gaps = 18/95 (18%)
Query 54 VEVRECNMDDAEIEKCGEFVEWDPPMNGDCVRGGTHTRYRQNCPDRKEVRVCGAFDCSSC 113
+E ++C +D+ E+ +CG + P+NG C + C + + C
Sbjct 1823 LEKKQCTVDEEEVPQCGACLPGHHPINGTC----QSLQISGLCAQKND-----------C 1867
Query 114 SVNATC---DPIGASCECKPGFRGNGKTCEAFNPC 145
+ +A C P C C GF G+G C+ + C
Sbjct 1868 NKHAECIDIHPDSHFCSCPDGFIGDGMICDDVDEC 1902
Score = 28.9 bits (63), Expect = 3.5, Method: Compositional matrix adjust.
Identities = 23/63 (36%), Positives = 29/63 (46%), Gaps = 8/63 (12%)
Query 86 GGTHTRYRQNCPDRKEVRVCGAFDCSS-CSVNATCDPIGASCECKPGFRGNGKTCEAFNP 144
GG+ T R P E+ G C+S C N+ C +G CEC G+ GN A
Sbjct 492 GGSITVTRGLIPKDVELTTSGRLACTSYCPPNSEC--VGGYCECVSGYGGN-----ALVG 544
Query 145 CED 147
CED
Sbjct 545 CED 547
> Hs22048730
Length=995
Score = 30.8 bits (68), Expect = 1.0, Method: Composition-based stats.
Identities = 12/27 (44%), Positives = 13/27 (48%), Gaps = 0/27 (0%)
Query 107 AFDCSSCSVNATCDPIGASCECKPGFR 133
A C C N D G SC C PGF+
Sbjct 59 ALSCVPCGANQRQDARGTSCVCLPGFQ 85
> Hs22062035
Length=179
Score = 30.8 bits (68), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 19/35 (54%), Gaps = 2/35 (5%)
Query 113 CSVNATCD--PIGASCECKPGFRGNGKTCEAFNPC 145
C +A C P C CKPG++G G+ CE + C
Sbjct 19 CHADALCQNTPTSYKCSCKPGYQGEGRQCEDIDEC 53
> Hs4758686
Length=4544
Score = 30.8 bits (68), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 34/76 (44%), Gaps = 7/76 (9%)
Query 80 NGDCVRGGTHTRYRQNCPDRKEVRVCGAFDC-----SSCSVNATCDPIGASCECKPGFR- 133
+G CV + +C D + R C +C S CS + IG C C+PGFR
Sbjct 2914 SGRCVAEALLCNGQDDCGDSSDERGCHINECLSRKLSGCSQDCEDLKIGFKCRCRPGFRL 2973
Query 134 -GNGKTCEAFNPCEDT 148
+G+TC + C T
Sbjct 2974 KDDGRTCADVDECSTT 2989
> CE18164_1
Length=654
Score = 30.4 bits (67), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 4/41 (9%)
Query 109 DCSSCSVNATCDPIGASCECKPGFRGNGKTCEAFNPCEDTP 149
D SC +N+ I C+C G+RG+G C N C +TP
Sbjct 587 DDGSCVLNS----IDMQCKCNNGYRGDGYNCTDINECVETP 623
> CE12964_1
Length=679
Score = 30.4 bits (67), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 4/41 (9%)
Query 109 DCSSCSVNATCDPIGASCECKPGFRGNGKTCEAFNPCEDTP 149
D SC +N+ I C+C G+RG+G C N C +TP
Sbjct 599 DDGSCVLNS----IDMQCKCNNGYRGDGYNCTDINECVETP 635
> At2g14720
Length=628
Score = 30.4 bits (67), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 15/35 (42%), Positives = 21/35 (60%), Gaps = 1/35 (2%)
Query 114 SVNATCDPIGASCECKPGFRGNG-KTCEAFNPCED 147
+ +A D CEC PGF+G+G K CE N C++
Sbjct 489 AFSACVDKDSVKCECPPGFKGDGVKKCEDINECKE 523
> At2g14740
Length=628
Score = 30.4 bits (67), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 15/35 (42%), Positives = 21/35 (60%), Gaps = 1/35 (2%)
Query 114 SVNATCDPIGASCECKPGFRGNG-KTCEAFNPCED 147
+ +A D CEC PGF+G+G K CE N C++
Sbjct 489 AFSACVDKDSVKCECPPGFKGDGTKKCEDINECKE 523
> Hs14192941
Length=969
Score = 30.0 bits (66), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 21/52 (40%), Positives = 25/52 (48%), Gaps = 2/52 (3%)
Query 83 CVRGGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRG 134
C+ G T R CP + C + CS C +C P SCEC PGFRG
Sbjct 478 CLAGWTGIRCDSTCPPGRWGPNC-SVSCS-CENGGSCSPEDGSCECAPGFRG 527
Score = 28.9 bits (63), Expect = 3.5, Method: Compositional matrix adjust.
Identities = 13/23 (56%), Positives = 16/23 (69%), Gaps = 0/23 (0%)
Query 112 SCSVNATCDPIGASCECKPGFRG 134
SC+ N TC PI SC+C PG+ G
Sbjct 593 SCANNGTCSPIDGSCQCFPGWIG 615
Score = 28.5 bits (62), Expect = 5.6, Method: Compositional matrix adjust.
Identities = 17/36 (47%), Positives = 21/36 (58%), Gaps = 4/36 (11%)
Query 112 SCSVNATCDPIGASCECKPGFRGNGKTCEAFNPCED 147
+C+ A C PI SC C PG+ G+ TCE PC D
Sbjct 419 TCANGAACSPIDGSCSCTPGWLGD--TCEL--PCPD 450
Score = 27.7 bits (60), Expect = 8.1, Method: Compositional matrix adjust.
Identities = 11/22 (50%), Positives = 15/22 (68%), Gaps = 0/22 (0%)
Query 113 CSVNATCDPIGASCECKPGFRG 134
C N+TCD + +C C PGF+G
Sbjct 723 CMNNSTCDHVTGTCYCSPGFKG 744
> CE06869
Length=145
Score = 30.0 bits (66), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 23/78 (29%), Positives = 32/78 (41%), Gaps = 15/78 (19%)
Query 82 DCVRGGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCE----------CKPG 131
DCVR T CP+ + + CG +C DP SCE C PG
Sbjct 64 DCVRRLECTAETSRCPEDEVFQTCGTLCQPTCD-----DPYPTSCEHDRCIRNVCRCLPG 118
Query 132 FRGNGKTCEAFNPCEDTP 149
N TC + + C+++P
Sbjct 119 LVRNSGTCTSLDECDNSP 136
> Hs4507485
Length=1170
Score = 30.0 bits (66), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 6/33 (18%)
Query 113 CSVNATCDPIGAS------CECKPGFRGNGKTC 139
C+ NA C+ +G CECKPG+ GNG C
Sbjct 657 CNKNAKCNYLGHYSDPMYRCECKPGYAGNGIIC 689
> Hs6912566
Length=392
Score = 29.6 bits (65), Expect = 2.1, Method: Compositional matrix adjust.
Identities = 32/121 (26%), Positives = 49/121 (40%), Gaps = 19/121 (15%)
Query 43 VQERYN-SKCGP-----VEVRECNMDDAEIEKCGEFVEWDPPMNGDCVRG-----GTHTR 91
VQ+ Y S+ GP +V+ + ++ E+V+ PP G TH++
Sbjct 51 VQKSYQESETGPESSIITKVKGITTSEHKVWDVEEYVK--PPEGGSVFSIITRVEATHSQ 108
Query 92 YRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCE---CKPGFRGNGKTCEAFN--PCE 146
+ CP+ V + C V D +G C P ++G KTCE F P E
Sbjct 109 TQGTCPESIRVHNATCLSDADC-VAGELDMLGNGLRTGRCVPYYQGPSKTCEVFGWCPVE 167
Query 147 D 147
D
Sbjct 168 D 168
> CE05765
Length=4753
Score = 29.6 bits (65), Expect = 2.2, Method: Compositional matrix adjust.
Identities = 32/125 (25%), Positives = 54/125 (43%), Gaps = 16/125 (12%)
Query 35 WTMCDTNRVQERYNSKCGPVEVRECNM-----DDAEIEKCGEFV----EWDPPMNGDCVR 85
+ C N Q + N +C P + R C+ D+++ ++CGE+ +W+ P G C+
Sbjct 220 FKKCTANEFQCK-NKRCQPRKFR-CDYYDDCGDNSDEDECGEYRCPPGKWNCPGTGHCID 277
Query 86 GGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPI--GASCECKPGFRGN---GKTCE 140
++C D + + C C S A C P G C C G++ + +TC
Sbjct 278 QLKLCDGSKDCADGADEQQCSQNLCPSLGCQAGCHPSPHGGECTCPSGYKLDDRFHRTCS 337
Query 141 AFNPC 145
N C
Sbjct 338 DINEC 342
> Hs6806919
Length=4655
Score = 29.3 bits (64), Expect = 2.7, Method: Compositional matrix adjust.
Identities = 20/77 (25%), Positives = 33/77 (42%), Gaps = 7/77 (9%)
Query 80 NGDCVRGGTHTRYRQNCPDRKEVRVCGAFDC-----SSCSVNATCDPIGASCECKPGFR- 133
NG C+ + +C D + + CG +C S C N T C C+PG++
Sbjct 3084 NGRCIEMMKLCNHLDDCLDNSDEKGCGINECHDPSISGCDHNCTDTLTSFYCSCRPGYKL 3143
Query 134 -GNGKTCEAFNPCEDTP 149
+ +TC + C + P
Sbjct 3144 MSDKRTCVDIDECTEMP 3160
Score = 28.1 bits (61), Expect = 6.7, Method: Compositional matrix adjust.
Identities = 22/84 (26%), Positives = 36/84 (42%), Gaps = 11/84 (13%)
Query 74 EWDPPMNGDCVRGGTHTRYRQNCPDRKEV------RVCGAFDCSSCSVNATCD--PIGAS 125
EW P +G C+ +CP R++ + C CS+ + C P G +
Sbjct 270 EWSCPESGRCISIYKVCDGILDCPGREDENNTSTGKYCSMTLCSALNCQYQCHETPYGGA 329
Query 126 CECKPGF---RGNGKTCEAFNPCE 146
C C PG+ + +TC F+ C+
Sbjct 330 CFCPPGYIINHNDSRTCVEFDDCQ 353
> At1g21210
Length=738
Score = 29.3 bits (64), Expect = 2.7, Method: Compositional matrix adjust.
Identities = 15/35 (42%), Positives = 18/35 (51%), Gaps = 4/35 (11%)
Query 105 CGAFDCSSCSVNATCD----PIGASCECKPGFRGN 135
CG C VN C IG +C+CK GF+GN
Sbjct 236 CGQVGEKKCGVNGICSNSASGIGYTCKCKGGFQGN 270
> At2g23450
Length=694
Score = 29.3 bits (64), Expect = 3.0, Method: Composition-based stats.
Identities = 15/48 (31%), Positives = 21/48 (43%), Gaps = 11/48 (22%)
Query 106 GAFDCSSCSVNATCDPI-------GASCECKPGFRGNGKTCEAFNPCE 146
G + +C+ N C + G C C GF G+G T NPC+
Sbjct 206 GGCESGTCAANTDCTDVETPHGYAGHRCSCLDGFHGDGYT----NPCQ 249
> CE23711
Length=1440
Score = 28.9 bits (63), Expect = 3.5, Method: Composition-based stats.
Identities = 19/52 (36%), Positives = 22/52 (42%), Gaps = 8/52 (15%)
Query 92 YRQNCPDRKEVRVCG------AFDCSSCSVNATCDPIGAS--CECKPGFRGN 135
YR +CP E + C + C N C PI S C C PGF GN
Sbjct 984 YRCDCPMEYEGKHCEDKLEYCTKKLNPCENNGKCIPINGSYSCMCSPGFTGN 1035
> CE27208
Length=2760
Score = 28.9 bits (63), Expect = 3.7, Method: Compositional matrix adjust.
Identities = 12/26 (46%), Positives = 17/26 (65%), Gaps = 2/26 (7%)
Query 114 SVNATCDPIGASCECKPGFRGNGKTC 139
S++ C+P+ CECKPG G+TC
Sbjct 1654 SLHGACNPLSGQCECKPGV--TGRTC 1677
> 7301143
Length=2146
Score = 28.9 bits (63), Expect = 3.7, Method: Compositional matrix adjust.
Identities = 19/45 (42%), Positives = 26/45 (57%), Gaps = 7/45 (15%)
Query 109 DCSSCSVNATC-DPIGA-SCECKPGFRG-----NGKTCEAFNPCE 146
D + CS + C D IG +CEC+PGF G N C+ +NPC+
Sbjct 907 DSNPCSKHGNCNDGIGTYTCECEPGFEGTHCEINIDECDRYNPCQ 951
> Hs13124888
Length=553
Score = 28.9 bits (63), Expect = 3.8, Method: Compositional matrix adjust.
Identities = 15/37 (40%), Positives = 20/37 (54%), Gaps = 2/37 (5%)
Query 108 FDCSSCSVNATCDPIGAS--CECKPGFRGNGKTCEAF 142
D +CS +A C S C+CK G++GNG C A
Sbjct 225 MDSHTCSHHANCFNTQGSFKCKCKQGYKGNGLRCSAI 261
> 7292141
Length=434
Score = 28.9 bits (63), Expect = 4.2, Method: Compositional matrix adjust.
Identities = 29/111 (26%), Positives = 43/111 (38%), Gaps = 19/111 (17%)
Query 28 CNDQWTPWT----MCDTNRVQERYNSKCGPVEVRECNMDDAEIEKCGEFVEWDPPMNGDC 83
CN WT +C+ N+ N C ++ ++ E C + W + C
Sbjct 192 CNPGWTGAKCAERICEANKYGLDCNRTC-ECDMEHTDLCHPETGNCQCSIGWS---SAQC 247
Query 84 VRGGTHTRYRQNCPDRKEVRVCGAFDCSSCSVNATCDPIGASCECKPGFRG 134
R T RY NC E+ +C A C P+ +C C PG+RG
Sbjct 248 TRPCTFLRYGPNC----ELTC-------NCKNGAKCSPVNGTCLCAPGWRG 287
> Hs20556510
Length=1007
Score = 28.5 bits (62), Expect = 5.4, Method: Compositional matrix adjust.
Identities = 14/37 (37%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Query 112 SCSVNATCD--PIGASCECKPGFRGNGKTCEAFNPCE 146
+C ++A C P C CK G+ G+GK C+ + CE
Sbjct 39 NCHIDAICQNTPRSYKCICKSGYTGDGKHCKDVDECE 75
> Hs22044235
Length=1229
Score = 28.1 bits (61), Expect = 5.8, Method: Compositional matrix adjust.
Identities = 12/24 (50%), Positives = 15/24 (62%), Gaps = 0/24 (0%)
Query 112 SCSVNATCDPIGASCECKPGFRGN 135
SC N+TC+P +C C PGF G
Sbjct 1114 SCHNNSTCEPATGTCRCGPGFYGQ 1137
> Hs15451866
Length=700
Score = 28.1 bits (61), Expect = 6.1, Method: Compositional matrix adjust.
Identities = 25/75 (33%), Positives = 33/75 (44%), Gaps = 7/75 (9%)
Query 80 NGDCVRGGTHTRYRQNCPDRKEVRVC--GAFDC----SSCSVNATCDPIGASCECKPGFR 133
+G CV H Q+CPD + C G +C CS T IG C C GF+
Sbjct 179 DGTCVLAIKHCNQEQDCPDGSDEAGCLQGLNECLHNNGGCSHICTDLKIGFECTCPAGFQ 238
Query 134 -GNGKTCEAFNPCED 147
+ KTC + C+D
Sbjct 239 LLDQKTCGDIDECKD 253
> Hs4503667
Length=2911
Score = 28.1 bits (61), Expect = 6.8, Method: Compositional matrix adjust.
Identities = 36/103 (34%), Positives = 43/103 (41%), Gaps = 20/103 (19%)
Query 61 MDDAEIEKCGEFVEWDPPMNGDC--VRGGTHTRYRQNCPDRKEVRVCG--AFDCSSCSVN 116
+D +I +C ++ D NG C +RG YR NC E G D C VN
Sbjct 763 VDGRDINECA--LDPDICANGICENLRG----SYRCNCNSGYEPDASGRNCIDIDECLVN 816
Query 117 -ATCD-------PIGASCECKPG--FRGNGKTCEAFNPCEDTP 149
CD P SC C PG FR +TCE N CE P
Sbjct 817 RLLCDNGLCRNTPGSYSCTCPPGYVFRTETETCEDINECESNP 859
> At1g30900
Length=631
Score = 27.7 bits (60), Expect = 7.8, Method: Compositional matrix adjust.
Identities = 10/24 (41%), Positives = 16/24 (66%), Gaps = 0/24 (0%)
Query 124 ASCECKPGFRGNGKTCEAFNPCED 147
+ C C PGF+G+G CE + C++
Sbjct 496 SGCRCPPGFKGDGLKCEDIDECKE 519
> Hs4758688
Length=963
Score = 27.7 bits (60), Expect = 8.1, Method: Compositional matrix adjust.
Identities = 25/75 (33%), Positives = 33/75 (44%), Gaps = 7/75 (9%)
Query 80 NGDCVRGGTHTRYRQNCPDRKEVRVC--GAFDC----SSCSVNATCDPIGASCECKPGFR 133
+G CV H Q+CPD + C G +C CS T IG C C GF+
Sbjct 308 DGTCVLAIKHCNQEQDCPDGSDEAGCLQGLNECLHNNGGCSHICTDLKIGFECTCPAGFQ 367
Query 134 -GNGKTCEAFNPCED 147
+ KTC + C+D
Sbjct 368 LLDQKTCGDIDECKD 382
> CE04271_1
Length=2510
Score = 27.7 bits (60), Expect = 9.1, Method: Compositional matrix adjust.
Identities = 16/35 (45%), Positives = 19/35 (54%), Gaps = 5/35 (14%)
Query 109 DCSS---CSVNATCDPIGASCECKPGFRGNGKTCE 140
DC S C VN C + C C PGF+ NG+ CE
Sbjct 826 DCKSDDVCPVNGKC--VNGMCLCLPGFKLNGEVCE 858
> Hs22052189
Length=665
Score = 27.7 bits (60), Expect = 9.2, Method: Compositional matrix adjust.
Identities = 18/51 (35%), Positives = 24/51 (47%), Gaps = 4/51 (7%)
Query 93 RQNCPDRKEVRVCGAFDCSSCSVNATCD----PIGASCECKPGFRGNGKTC 139
+ CP R +CG+ SSC+ A D P C+C PGF +G C
Sbjct 442 KLACPPRSNYELCGSSCPSSCAEPALPDSCLTPCQDGCQCDPGFVLSGMDC 492
Lambda K H
0.320 0.137 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1858150626
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40