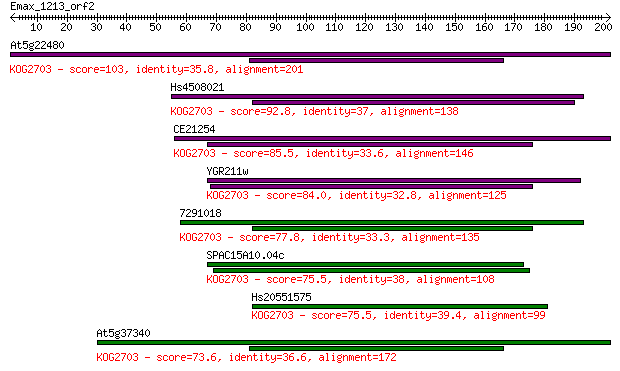
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_1213_orf2
Length=201
Score E
Sequences producing significant alignments: (Bits) Value
At5g22480 103 2e-22
Hs4508021 92.8 4e-19
CE21254 85.5 6e-17
YGR211w 84.0 2e-16
7291018 77.8 1e-14
SPAC15A10.04c 75.5 7e-14
Hs20551575 75.5 7e-14
At5g37340 73.6 3e-13
> At5g22480
Length=493
Score = 103 bits (257), Expect = 2e-22, Method: Compositional matrix adjust.
Identities = 72/208 (34%), Positives = 102/208 (49%), Gaps = 12/208 (5%)
Query 1 GESASSGESGESASGDTQAAGAAGAAGAAGAAGAAAGAAAGAAGGEEEDCSSSLLR---- 56
G A+ ++G+S G A A G GA AG A A D S +L R
Sbjct 219 GYVANPSQAGQS-EGSLGAPSTKTAYVPNGTIGATAGHRA-IAQSNSTDISDNLFRYSAP 276
Query 57 EEALLLPVERPHCGTQTHNKVCQLF---IPGFRECLLFAFSCSICGAKNSVIKTAGAYGQ 113
EE + P CG T ++F IP F+E ++ A +C CG +NS +K GA +
Sbjct 277 EEVMTFPST---CGACTEPCETRMFVTKIPYFQEVIVMASTCDSCGYRNSELKPGGAIPE 333
Query 114 TGRRWILTVEGEEDLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLA 173
G++ L+V DL+RDV+K DT+ +IIP LD + GG G +TT+EGL+ +RE LA
Sbjct 334 KGKKITLSVRNITDLSRDVIKSDTAGVIIPELDLELAGGTLGGMVTTVEGLVTQIRESLA 393
Query 174 ESVPFAVGDSAAAASRERMGCIGTYLQR 201
F GDS + + G L +
Sbjct 394 RVHGFTFGDSMEESKLNKWREFGARLTK 421
Score = 63.2 bits (152), Expect = 3e-10, Method: Compositional matrix adjust.
Identities = 33/87 (37%), Positives = 52/87 (59%), Gaps = 2/87 (2%)
Query 81 FIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEG--EEDLNRDVLKGDTS 138
IP FR+ L+ AF C CG +N+ ++ AG G + L V + +R V+K +++
Sbjct 50 LIPHFRKVLISAFECPHCGERNNEVQFAGEIQPRGCCYNLEVLAGDVKIFDRQVVKSESA 109
Query 139 TIIIPSLDFSMEGGAQAGTLTTIEGLI 165
TI IP LDF + AQ G+L+T+EG++
Sbjct 110 TIKIPELDFEIPPEAQRGSLSTVEGIL 136
> Hs4508021
Length=459
Score = 92.8 bits (229), Expect = 4e-19, Method: Compositional matrix adjust.
Identities = 51/140 (36%), Positives = 75/140 (53%), Gaps = 5/140 (3%)
Query 55 LREEALLLPVERPHCGT--QTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYG 112
LR E L P C QT+ K+ Q IP F+E ++ A +C CG + + +K+ GA
Sbjct 248 LRNEVLQFSTNCPECNAPAQTNMKLVQ--IPHFKEVIIMATNCENCGHRTNEVKSGGAVE 305
Query 113 QTGRRWILTVEGEEDLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGL 172
G R L + D+ RD+LK +T ++ IP L+F + G TT+EGL+K +RE L
Sbjct 306 PLGTRITLHITDASDMTRDLLKSETCSVEIPELEFELGMAVLGGKFTTLEGLLKDIRE-L 364
Query 173 AESVPFAVGDSAAAASRERM 192
PF +GDS+ ER+
Sbjct 365 VTKNPFTLGDSSNPGQTERL 384
Score = 90.1 bits (222), Expect = 3e-18, Method: Compositional matrix adjust.
Identities = 45/108 (41%), Positives = 63/108 (58%), Gaps = 0/108 (0%)
Query 82 IPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEEDLNRDVLKGDTSTII 141
IP FRE ++ +FSC CG N+ I++AG G R+ L+V ED+NR+V+K D++
Sbjct 67 IPFFREIIVSSFSCEHCGWNNTEIQSAGRIQDQGVRYTLSVRALEDMNREVVKTDSAATR 126
Query 142 IPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAESVPFAVGDSAAAASR 189
IP LDF + +Q G LTT+EGLI GL + P + A A R
Sbjct 127 IPELDFEIPAFSQKGALTTVEGLITRAISGLEQDQPARRANKDATAER 174
> CE21254
Length=455
Score = 85.5 bits (210), Expect = 6e-17, Method: Compositional matrix adjust.
Identities = 49/146 (33%), Positives = 78/146 (53%), Gaps = 3/146 (2%)
Query 56 REEALLLPVERPHCGTQTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTG 115
++E L + P+C T K+ IP F+ ++ + +C CG K++ +K+ GA G
Sbjct 237 KQEVLHFATDCPNCHGPTEVKMKPTDIPFFQTVIIMSLACDRCGYKSNEVKSGGAIRDQG 296
Query 116 RRWILTVEGEEDLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAES 175
R + +E + DL RDVLK DT + IP +D + G A G TTIEGL+ + +E L
Sbjct 297 CRMSVKLEKDLDLARDVLKTDTCALSIPEIDLEVGGNALCGRFTTIEGLLTATKEQLDAQ 356
Query 176 VPFAVGDSAAAASRERMGCIGTYLQR 201
F +GDSA + + T+L++
Sbjct 357 SSFFMGDSAQTGEK---SAVTTFLEK 379
Score = 72.4 bits (176), Expect = 6e-13, Method: Compositional matrix adjust.
Identities = 33/109 (30%), Positives = 60/109 (55%), Gaps = 0/109 (0%)
Query 67 PHCGTQTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEE 126
P C ++ IP +R +L +F C CG KN+ I++ A + G +L V+ E
Sbjct 29 PVCEEDGETRIMCTSIPYYRAVILMSFECPHCGHKNNEIQSGEAVQEHGTLIVLRVQKPE 88
Query 127 DLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAES 175
DL R ++K + ++I +P L + +Q G +TT+EG+++ + GL++
Sbjct 89 DLRRQLVKSEYASIEVPELQLEIPHKSQPGEVTTVEGVLERVHRGLSQD 137
> YGR211w
Length=486
Score = 84.0 bits (206), Expect = 2e-16, Method: Compositional matrix adjust.
Identities = 41/125 (32%), Positives = 67/125 (53%), Gaps = 0/125 (0%)
Query 67 PHCGTQTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEE 126
P C + + + IP F+E ++ + C CG K++ +KT GA GRR L +
Sbjct 296 PSCTQECETHMKPVNIPHFKEVIIMSTVCDHCGYKSNEVKTGGAIPDKGRRITLYCDDAA 355
Query 127 DLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAESVPFAVGDSAAA 186
DL+RD+LK +T +++IP L ++ G G TT+EGL++ + E L + DS
Sbjct 356 DLSRDILKSETCSMVIPELHLDIQEGTLGGRFTTLEGLLRQVYEELESRIFTQTSDSMDE 415
Query 187 ASRER 191
A++ R
Sbjct 416 ATKAR 420
Score = 78.6 bits (192), Expect = 9e-15, Method: Compositional matrix adjust.
Identities = 40/108 (37%), Positives = 61/108 (56%), Gaps = 2/108 (1%)
Query 68 HCGTQTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEED 127
+CG ++ IP FRE ++ +F C CG KN I+ A + G R++L VE ED
Sbjct 56 NCGKNGTTRLLLTSIPYFREIIIMSFDCPHCGFKNCEIQPASQIQEKGSRYVLKVECRED 115
Query 128 LNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAES 175
NR V+K +T+T LD +E A+ G LTT+EGL+ + + L++
Sbjct 116 FNRQVIKSETATCKFVELD--IEIPAKRGQLTTVEGLLSEMIDDLSQD 161
> 7291018
Length=457
Score = 77.8 bits (190), Expect = 1e-14, Method: Compositional matrix adjust.
Identities = 45/135 (33%), Positives = 70/135 (51%), Gaps = 3/135 (2%)
Query 58 EALLLPVERPHCGTQTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRR 117
E L P P C + IP F+E ++ A C CG K + +K+ G G R
Sbjct 253 EVLQFPTNCPSCQAPCETNMKLTNIPHFKEVVIMATVCGACGHKTNEVKSGGGVEAQGVR 312
Query 118 WILTVEGEEDLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAESVP 177
+ + + EDL RDVLK +T ++ IP LD + A G TT+EGL+ ++R+ L ++
Sbjct 313 FRVQIASREDLTRDVLKSETCSMSIPELDLEVGPHALCGRFTTVEGLLVAMRDQLDGTL- 371
Query 178 FAVGDSAAAASRERM 192
DSA A++++M
Sbjct 372 --FHDSADDATKQQM 384
Score = 74.7 bits (182), Expect = 1e-13, Method: Compositional matrix adjust.
Identities = 35/94 (37%), Positives = 57/94 (60%), Gaps = 0/94 (0%)
Query 82 IPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEEDLNRDVLKGDTSTII 141
IP FRE +L +F C CG N+ +++A ++G R L V+ DLNR V++ D S+I
Sbjct 59 IPFFREVVLMSFKCDHCGHINNEMQSASEIQKSGIRIELRVQSVADLNRRVVRSDNSSIS 118
Query 142 IPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAES 175
IP ++ + +Q G +TT+EG+I+ GL++
Sbjct 119 IPEIELEIPVQSQKGEVTTVEGIIERTIAGLSQD 152
> SPAC15A10.04c
Length=459
Score = 75.5 bits (184), Expect = 7e-14, Method: Compositional matrix adjust.
Identities = 41/108 (37%), Positives = 61/108 (56%), Gaps = 4/108 (3%)
Query 67 PHCGTQ--THNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEG 124
P C Q TH K+ L IP F+E ++ + C CG +++ +KT G GR+ L V
Sbjct 260 PSCSHQCDTHMKL--LDIPHFKEVIIMSTVCDRCGYRSNEVKTGGEIPPKGRKITLKVMD 317
Query 125 EEDLNRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGL 172
EDL+RD+LK +T+++ IP L + G G TTIEGL+ + + L
Sbjct 318 AEDLSRDILKSETASLKIPELGLDLFPGTLGGRFTTIEGLLAQVYDEL 365
Score = 72.4 bits (176), Expect = 5e-13, Method: Compositional matrix adjust.
Identities = 37/106 (34%), Positives = 58/106 (54%), Gaps = 2/106 (1%)
Query 69 CGTQTHNKVCQLFIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEEDL 128
CG K+ IP FRE +L +F C CG KN+ ++ A G + VE +EDL
Sbjct 41 CGKNGTTKLLLTVIPYFREVVLMSFECPHCGFKNAQVQHAETIQPEGSKITFHVEDKEDL 100
Query 129 NRDVLKGDTSTIIIPSLDFSMEGGAQAGTLTTIEGLIKSLREGLAE 174
NR V+K + + IP + + G + G LTTIEG++ ++ + L++
Sbjct 101 NRTVVKSQEAIVSIPEIQLEIPG--RLGQLTTIEGILSNVVDDLSK 144
> Hs20551575
Length=363
Score = 75.5 bits (184), Expect = 7e-14, Method: Compositional matrix adjust.
Identities = 39/110 (35%), Positives = 59/110 (53%), Gaps = 11/110 (10%)
Query 82 IPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEEDLNRDVLKGDTSTII 141
IP FRE ++ +FSC CG N+ I++AG G R+ L++ ED+NR+V+K D++T
Sbjct 67 IPFFREIIVSSFSCHHCGWNNTEIQSAGRIQDQGVRYTLSIRALEDMNREVVKTDSATTR 126
Query 142 IPSLDFSMEGGAQAG-----------TLTTIEGLIKSLREGLAESVPFAV 180
IP LDF + +Q G T I+ I L+E + PF +
Sbjct 127 IPELDFQIPAFSQKGGLACAKGNKDATAERIDEFIVKLKELKQSASPFTL 176
> At5g37340
Length=600
Score = 73.6 bits (179), Expect = 3e-13, Method: Compositional matrix adjust.
Identities = 63/220 (28%), Positives = 92/220 (41%), Gaps = 54/220 (24%)
Query 30 GAAGAAAGAAAGAAGGEEEDCSSSLLR----EEALLLPVERPHCGTQTHNKVCQ--LFIP 83
G GA AG A A D S +L R EE + P CG T K+C+ +F+
Sbjct 315 GTIGATAGHRA-IAQSNSTDISDNLFRYSAPEEVMTFPST---CGACT--KLCETRMFVT 368
Query 84 GFR--------ECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEGEEDLNRDVLK- 134
E ++ A +C CG +NS +K GA + G++ L+V+ DL+RDV+K
Sbjct 369 SILSKLCSRSLEVIVMASTCDDCGYRNSELKPGGAIPEKGKKITLSVKNITDLSRDVIKV 428
Query 135 ---------------------------------GDTSTIIIPSLDFSMEGGAQAGTLTTI 161
DT+ + IP LD + GG G +TT+
Sbjct 429 SSYRSMYLKVAENVTLRLKYMPLQNVYSLSLVKSDTAGVKIPELDLELAGGTLGGMVTTV 488
Query 162 EGLIKSLREGLAESVPFAVGDSAAAASRERMGCIGTYLQR 201
EGL+ +RE LA F GDS + + G+ L +
Sbjct 489 EGLVTQIRESLARVHGFTFGDSLEQSKINKWKEFGSRLTK 528
Score = 56.2 bits (134), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 33/103 (32%), Positives = 53/103 (51%), Gaps = 18/103 (17%)
Query 81 FIPGFRECLLFAFSCSICGAKNSVIKTAGAYGQTGRRWILTVEG--EEDLNRDVLKGDTS 138
IP FR+ L+ AF C CG +N+ ++ AG G + L V + +R V+K +++
Sbjct 102 LIPHFRKVLISAFECPHCGERNNEVQFAGEIQPRGCSYHLEVSAGDVKTFDRQVVKSESA 161
Query 139 TII----------------IPSLDFSMEGGAQAGTLTTIEGLI 165
TI IP LDF + AQ+G+L+T+EG++
Sbjct 162 TIKVSSLTDSGKFNASSDSIPELDFEIPPEAQSGSLSTVEGIL 204
Lambda K H
0.313 0.130 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3515233148
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40