bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_1745_orf1
Length=111
Score E
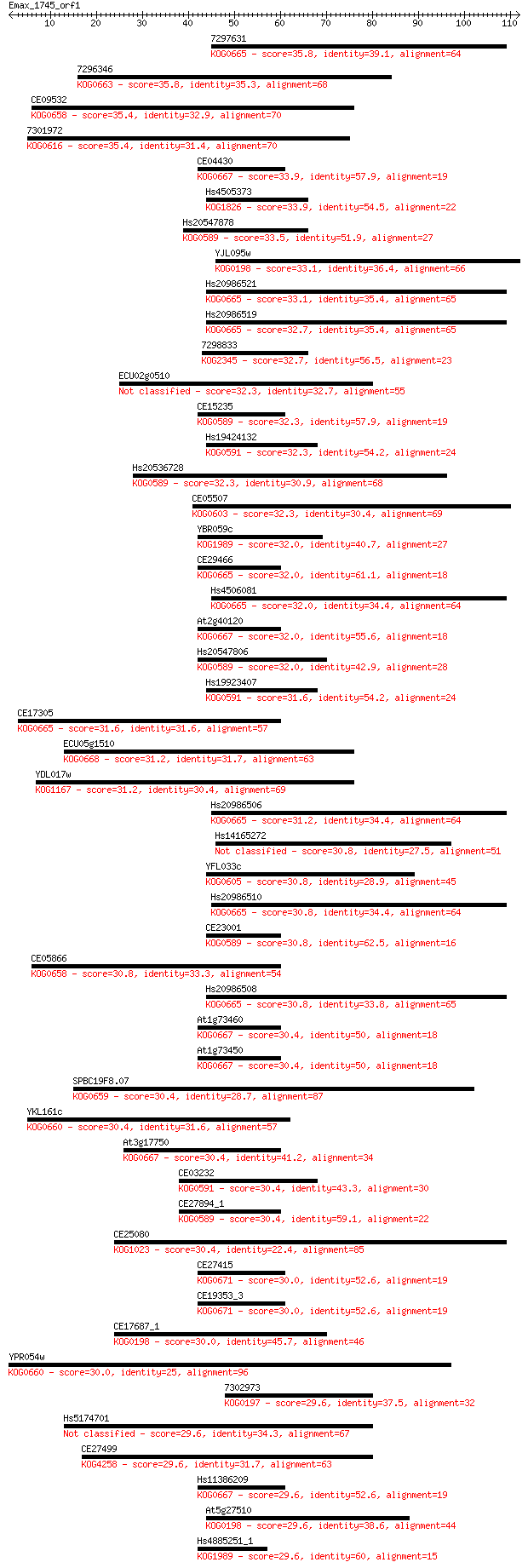
Sequences producing significant alignments: (Bits) Value
7297631 35.8 0.019
7296346 35.8 0.020
CE09532 35.4 0.027
7301972 35.4 0.029
CE04430 33.9 0.077
Hs4505373 33.9 0.078
Hs20547878 33.5 0.10
YJL095w 33.1 0.11
Hs20986521 33.1 0.14
Hs20986519 32.7 0.17
7298833 32.7 0.19
ECU02g0510 32.3 0.21
CE15235 32.3 0.22
Hs19424132 32.3 0.22
Hs20536728 32.3 0.23
CE05507 32.3 0.24
YBR059c 32.0 0.28
CE29466 32.0 0.30
Hs4506081 32.0 0.31
At2g40120 32.0 0.32
Hs20547806 32.0 0.32
Hs19923407 31.6 0.34
CE17305 31.6 0.35
ECU05g1510 31.2 0.52
YDL017w 31.2 0.53
Hs20986506 31.2 0.56
Hs14165272 30.8 0.57
YFL033c 30.8 0.60
Hs20986510 30.8 0.60
CE23001 30.8 0.62
CE05866 30.8 0.68
Hs20986508 30.8 0.73
At1g73460 30.4 0.78
At1g73450 30.4 0.78
SPBC19F8.07 30.4 0.80
YKL161c 30.4 0.88
At3g17750 30.4 0.89
CE03232 30.4 0.90
CE27894_1 30.4 0.94
CE25080 30.4 0.94
CE27415 30.0 1.1
CE19353_3 30.0 1.1
CE17687_1 30.0 1.1
YPR054w 30.0 1.2
7302973 29.6 1.3
Hs5174701 29.6 1.3
CE27499 29.6 1.3
Hs11386209 29.6 1.4
At5g27510 29.6 1.4
Hs4885251_1 29.6 1.4
> 7297631
Length=372
Score = 35.8 bits (81), Expect = 0.019, Method: Composition-based stats.
Identities = 25/66 (37%), Positives = 31/66 (46%), Gaps = 11/66 (16%)
Query 45 EKWDIWSMGCVLQEMF--GFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQG 102
E DIWS+GC++ EM G L P HI D N I E+L GT P +Q
Sbjct 202 ENVDIWSVGCIMGEMIRGGVLFPGTDHI---DQWNKIIEQL------GTPSPSFMQRLQP 252
Query 103 IARNII 108
RN +
Sbjct 253 TVRNYV 258
> 7296346
Length=952
Score = 35.8 bits (81), Expect = 0.020, Method: Composition-based stats.
Identities = 24/68 (35%), Positives = 33/68 (48%), Gaps = 15/68 (22%)
Query 16 WFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPMHVHISNRDS 75
W+ AP ELL CSPV P+ D+WS+GC+ E L P+ S D
Sbjct 725 WYRAP-ELLLCSPVYST---------PI----DVWSVGCIFAEFLQML-PLFPGKSEIDE 769
Query 76 PNVIYEKL 83
N I+++L
Sbjct 770 LNRIFKEL 777
> CE09532
Length=440
Score = 35.4 bits (80), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 23/70 (32%), Positives = 31/70 (44%), Gaps = 17/70 (24%)
Query 6 GRVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSP 65
G+ + T FY P ELL ++ M +TV D+WS GC++ EM
Sbjct 173 GKANNFYQVTRFYRPPELL----MKAMEYNSTV---------DVWSAGCIMAEMVK---- 215
Query 66 MHVHISNRDS 75
HV RDS
Sbjct 216 RHVVFPGRDS 225
> 7301972
Length=393
Score = 35.4 bits (80), Expect = 0.029, Method: Composition-based stats.
Identities = 22/70 (31%), Positives = 29/70 (41%), Gaps = 17/70 (24%)
Query 5 DGRVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLS 64
DGR S + G + AP VQL P N+ D W+ G ++ E S
Sbjct 194 DGRTSTLCGTPEYLAPE---------------IVQLRPYNKSVDWWAFGILVYEFVAGRS 238
Query 65 PMHVHISNRD 74
P +H NRD
Sbjct 239 PFAIH--NRD 246
> CE04430
Length=821
Score = 33.9 bits (76), Expect = 0.077, Method: Compositional matrix adjust.
Identities = 11/19 (57%), Positives = 15/19 (78%), Gaps = 0/19 (0%)
Query 42 PLNEKWDIWSMGCVLQEMF 60
P NE D+WS+GCV+ E+F
Sbjct 300 PFNESIDMWSLGCVIAELF 318
> Hs4505373
Length=445
Score = 33.9 bits (76), Expect = 0.078, Method: Composition-based stats.
Identities = 12/22 (54%), Positives = 16/22 (72%), Gaps = 0/22 (0%)
Query 44 NEKWDIWSMGCVLQEMFGFLSP 65
NEK DIWS+GC+L E+ + P
Sbjct 194 NEKSDIWSLGCLLYELCALMPP 215
> Hs20547878
Length=758
Score = 33.5 bits (75), Expect = 0.10, Method: Composition-based stats.
Identities = 14/27 (51%), Positives = 16/27 (59%), Gaps = 0/27 (0%)
Query 39 QLPPLNEKWDIWSMGCVLQEMFGFLSP 65
Q P N K DIWS+GCVL E+ P
Sbjct 153 QNKPYNNKTDIWSLGCVLYELCTLKHP 179
> YJL095w
Length=1478
Score = 33.1 bits (74), Expect = 0.11, Method: Composition-based stats.
Identities = 24/67 (35%), Positives = 33/67 (49%), Gaps = 7/67 (10%)
Query 46 KWDIWSMGCVLQEMFGFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVP-DIASGIQGIA 104
K DIWS+GC++ EMF P SN + +++ KSK S +P D I I
Sbjct 1360 KVDIWSLGCIVLEMFAGKRPW----SNLEVVAAMFKIGKSK--SAPPIPEDTLPLISQIG 1413
Query 105 RNIIARC 111
RN + C
Sbjct 1414 RNFLDAC 1420
> Hs20986521
Length=427
Score = 33.1 bits (74), Expect = 0.14, Method: Compositional matrix adjust.
Identities = 23/67 (34%), Positives = 31/67 (46%), Gaps = 11/67 (16%)
Query 44 NEKWDIWSMGCVLQEMF--GFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQ 101
E DIWS+GC++ EM G L P HI D N + E+L GT P+ +Q
Sbjct 203 KENVDIWSVGCIMGEMIKGGVLFPGTDHI---DQWNKVIEQL------GTPCPEFMKKLQ 253
Query 102 GIARNII 108
R +
Sbjct 254 PTVRTYV 260
> Hs20986519
Length=384
Score = 32.7 bits (73), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 23/67 (34%), Positives = 31/67 (46%), Gaps = 11/67 (16%)
Query 44 NEKWDIWSMGCVLQEMF--GFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQ 101
E DIWS+GC++ EM G L P HI D N + E+L GT P+ +Q
Sbjct 203 KENVDIWSVGCIMGEMIKGGVLFPGTDHI---DQWNKVIEQL------GTPCPEFMKKLQ 253
Query 102 GIARNII 108
R +
Sbjct 254 PTVRTYV 260
> 7298833
Length=320
Score = 32.7 bits (73), Expect = 0.19, Method: Compositional matrix adjust.
Identities = 13/23 (56%), Positives = 17/23 (73%), Gaps = 0/23 (0%)
Query 43 LNEKWDIWSMGCVLQEMFGFLSP 65
++E+ DIWS+GCVL M F SP
Sbjct 230 IDERTDIWSLGCVLYAMCYFNSP 252
> ECU02g0510
Length=586
Score = 32.3 bits (72), Expect = 0.21, Method: Composition-based stats.
Identities = 18/56 (32%), Positives = 26/56 (46%), Gaps = 7/56 (12%)
Query 25 RCSPVECMASRTTVQ-LPPLNEKWDIWSMGCVLQEMFGFLSPMHVHISNRDSPNVI 79
+C V M+ Q + DIWS+GC+L EM P+H + PN+I
Sbjct 443 QCGTVNYMSPEAVTQNKSKVARSSDIWSLGCILYEMVHSNPPLH------EYPNLI 492
> CE15235
Length=357
Score = 32.3 bits (72), Expect = 0.22, Method: Composition-based stats.
Identities = 11/19 (57%), Positives = 15/19 (78%), Gaps = 0/19 (0%)
Query 42 PLNEKWDIWSMGCVLQEMF 60
P N+K D+WS+GCVL E+
Sbjct 188 PYNQKSDMWSLGCVLYELL 206
> Hs19424132
Length=302
Score = 32.3 bits (72), Expect = 0.22, Method: Compositional matrix adjust.
Identities = 13/24 (54%), Positives = 16/24 (66%), Gaps = 0/24 (0%)
Query 44 NEKWDIWSMGCVLQEMFGFLSPMH 67
N K DIWS+GC+L EM SP +
Sbjct 214 NFKSDIWSLGCLLYEMAALQSPFY 237
> Hs20536728
Length=484
Score = 32.3 bits (72), Expect = 0.23, Method: Compositional matrix adjust.
Identities = 21/79 (26%), Positives = 32/79 (40%), Gaps = 11/79 (13%)
Query 28 PVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPMH-----------VHISNRDSP 76
PV+ ++ P EK D+W++GC+L +M P + V P
Sbjct 3 PVQPAEPPEVLKSEPYGEKADVWAVGCILYQMATLSPPFYSTNMLSLATKIVEAVYEPVP 62
Query 77 NVIYEKLKSKAISGTLVPD 95
IY + + IS L PD
Sbjct 63 EGIYSEKVTDTISRCLTPD 81
> CE05507
Length=785
Score = 32.3 bits (72), Expect = 0.24, Method: Compositional matrix adjust.
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query 41 PPLNEKWDIWSMGCVLQEMFGFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGI 100
P NE+ D+WS+G VL M P H S ++S I +++ S T D + +
Sbjct 579 PEYNEQCDLWSLGVVLFTMLSGQVPFHAR-SRQESATEIMQRICRAEFSFT--GDAWTNV 635
Query 101 QGIARNIIA 109
A+N+I
Sbjct 636 SADAKNLIT 644
> YBR059c
Length=1108
Score = 32.0 bits (71), Expect = 0.28, Method: Composition-based stats.
Identities = 11/27 (40%), Positives = 18/27 (66%), Gaps = 0/27 (0%)
Query 42 PLNEKWDIWSMGCVLQEMFGFLSPMHV 68
P+NEK DIW++G L ++ F +P +
Sbjct 244 PINEKSDIWALGVFLYKLLFFTTPFEM 270
> CE29466
Length=390
Score = 32.0 bits (71), Expect = 0.30, Method: Composition-based stats.
Identities = 11/18 (61%), Positives = 14/18 (77%), Gaps = 0/18 (0%)
Query 42 PLNEKWDIWSMGCVLQEM 59
P +EK DIWS+GC+ EM
Sbjct 216 PYSEKVDIWSVGCIFAEM 233
> Hs4506081
Length=422
Score = 32.0 bits (71), Expect = 0.31, Method: Composition-based stats.
Identities = 22/66 (33%), Positives = 31/66 (46%), Gaps = 11/66 (16%)
Query 45 EKWDIWSMGCVLQEMF--GFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQG 102
E DIWS+GC++ EM L P +I D N + E+L GT P+ +Q
Sbjct 242 ENVDIWSVGCIMGEMVRHKILFPGRDYI---DQWNKVIEQL------GTPCPEFMKKLQP 292
Query 103 IARNII 108
RN +
Sbjct 293 TVRNYV 298
> At2g40120
Length=570
Score = 32.0 bits (71), Expect = 0.32, Method: Composition-based stats.
Identities = 10/18 (55%), Positives = 15/18 (83%), Gaps = 0/18 (0%)
Query 42 PLNEKWDIWSMGCVLQEM 59
P +EK D+WS+GC+L E+
Sbjct 441 PYDEKIDLWSLGCILAEL 458
> Hs20547806
Length=489
Score = 32.0 bits (71), Expect = 0.32, Method: Compositional matrix adjust.
Identities = 12/28 (42%), Positives = 16/28 (57%), Gaps = 0/28 (0%)
Query 42 PLNEKWDIWSMGCVLQEMFGFLSPMHVH 69
P N K DIWS+GC+L E+ P +
Sbjct 178 PYNNKSDIWSLGCILYELCTLKHPFQAN 205
> Hs19923407
Length=313
Score = 31.6 bits (70), Expect = 0.34, Method: Compositional matrix adjust.
Identities = 13/24 (54%), Positives = 16/24 (66%), Gaps = 0/24 (0%)
Query 44 NEKWDIWSMGCVLQEMFGFLSPMH 67
N K DIWS+GC+L EM SP +
Sbjct 225 NFKSDIWSLGCLLYEMAALQSPFY 248
> CE17305
Length=375
Score = 31.6 bits (70), Expect = 0.35, Method: Composition-based stats.
Identities = 18/57 (31%), Positives = 27/57 (47%), Gaps = 15/57 (26%)
Query 3 SADGRVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEM 59
+ D R+S G ++ AP +L + +EK DIWS+GC+L EM
Sbjct 181 NVDMRMSDYVGTRYYRAPEVILGLN---------------YSEKVDIWSVGCILAEM 222
> ECU05g1510
Length=319
Score = 31.2 bits (69), Expect = 0.52, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 25/65 (38%), Gaps = 2/65 (3%)
Query 13 GATWFYAPAEL--LRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPMHVHI 70
G FY P + +R + V P + DIWS GCVL E+ P
Sbjct 172 GLAEFYHPKKEYSVRVASRYYKGPELLVDYPYYDYSLDIWSFGCVLAELVFKKRPFFHGE 231
Query 71 SNRDS 75
SN D
Sbjct 232 SNSDQ 236
> YDL017w
Length=507
Score = 31.2 bits (69), Expect = 0.53, Method: Composition-based stats.
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 15/69 (21%)
Query 7 RVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPM 66
R +R +G F AP L++C +++T K DIWS+G +L + G PM
Sbjct 275 RANR-AGTRGFRAPEVLMKC------GAQST--------KIDIWSVGVILLSLLGRRFPM 319
Query 67 HVHISNRDS 75
+ + DS
Sbjct 320 FQSLDDADS 328
> Hs20986506
Length=426
Score = 31.2 bits (69), Expect = 0.56, Method: Compositional matrix adjust.
Identities = 22/66 (33%), Positives = 31/66 (46%), Gaps = 11/66 (16%)
Query 45 EKWDIWSMGCVLQEM--FGFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQG 102
E DIWS+GC++ EM L P +I D N + E+L GT P+ +Q
Sbjct 204 ENVDIWSVGCIMGEMVRHKILFPGRDYI---DQWNKVIEQL------GTPCPEFMKKLQP 254
Query 103 IARNII 108
RN +
Sbjct 255 TVRNYV 260
> Hs14165272
Length=581
Score = 30.8 bits (68), Expect = 0.57, Method: Compositional matrix adjust.
Identities = 14/55 (25%), Positives = 30/55 (54%), Gaps = 9/55 (16%)
Query 46 KWDIWSMGCVLQEMFGFLSPMH----VHISNRDSPNVIYEKLKSKAISGTLVPDI 96
K D W++G + E+FG ++P + H+ +R Y++ + A+ ++ PD+
Sbjct 433 KADAWAVGAIAYEIFGLVNPFYGQGKAHLESRS-----YQEAQLPALPESVPPDV 482
> YFL033c
Length=1770
Score = 30.8 bits (68), Expect = 0.60, Method: Composition-based stats.
Identities = 13/45 (28%), Positives = 25/45 (55%), Gaps = 5/45 (11%)
Query 44 NEKWDIWSMGCVLQEMFGFLSPMHVHISNRDSPNVIYEKLKSKAI 88
N++ D WS+GC+ E+ P H ++P+ +++K+ S I
Sbjct 1166 NKQCDWWSVGCIFFELLLGYPPFHA-----ETPDAVFKKILSGVI 1205
> Hs20986510
Length=464
Score = 30.8 bits (68), Expect = 0.60, Method: Compositional matrix adjust.
Identities = 22/66 (33%), Positives = 31/66 (46%), Gaps = 11/66 (16%)
Query 45 EKWDIWSMGCVLQEM--FGFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQG 102
E DIWS+GC++ EM L P +I D N + E+L GT P+ +Q
Sbjct 242 ENVDIWSVGCIMGEMVRHKILFPGRDYI---DQWNKVIEQL------GTPCPEFMKKLQP 292
Query 103 IARNII 108
RN +
Sbjct 293 TVRNYV 298
> CE23001
Length=579
Score = 30.8 bits (68), Expect = 0.62, Method: Compositional matrix adjust.
Identities = 10/16 (62%), Positives = 14/16 (87%), Gaps = 0/16 (0%)
Query 44 NEKWDIWSMGCVLQEM 59
NEK D+W++GC+L EM
Sbjct 349 NEKSDMWALGCILYEM 364
> CE05866
Length=399
Score = 30.8 bits (68), Expect = 0.68, Method: Composition-based stats.
Identities = 18/54 (33%), Positives = 25/54 (46%), Gaps = 13/54 (24%)
Query 6 GRVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEM 59
G+ + T FY P ELL+ + M TV D+WS GC++ EM
Sbjct 173 GKANNHYQVTRFYRPPELLQ----KAMEYNCTV---------DVWSAGCIMAEM 213
> Hs20986508
Length=277
Score = 30.8 bits (68), Expect = 0.73, Method: Compositional matrix adjust.
Identities = 22/67 (32%), Positives = 31/67 (46%), Gaps = 11/67 (16%)
Query 44 NEKWDIWSMGCVLQEM--FGFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDIASGIQ 101
E DIWS+GC++ EM L P +I D N + E+L GT P+ +Q
Sbjct 96 KENVDIWSVGCIMGEMVRHKILFPGRDYI---DQWNKVIEQL------GTPCPEFMKKLQ 146
Query 102 GIARNII 108
RN +
Sbjct 147 PTVRNYV 153
> At1g73460
Length=1157
Score = 30.4 bits (67), Expect = 0.78, Method: Composition-based stats.
Identities = 9/18 (50%), Positives = 15/18 (83%), Gaps = 0/18 (0%)
Query 42 PLNEKWDIWSMGCVLQEM 59
P ++K D+WS+GC+L E+
Sbjct 1026 PYDKKIDVWSLGCILAEL 1043
> At1g73450
Length=1155
Score = 30.4 bits (67), Expect = 0.78, Method: Composition-based stats.
Identities = 9/18 (50%), Positives = 15/18 (83%), Gaps = 0/18 (0%)
Query 42 PLNEKWDIWSMGCVLQEM 59
P ++K D+WS+GC+L E+
Sbjct 1009 PYDKKIDVWSLGCILAEL 1026
> SPBC19F8.07
Length=335
Score = 30.4 bits (67), Expect = 0.80, Method: Composition-based stats.
Identities = 25/87 (28%), Positives = 38/87 (43%), Gaps = 20/87 (22%)
Query 15 TWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPMHVHISNRD 74
T +Y P EL + C + T V D+WS+GC+ E+ +P S+ D
Sbjct 170 TRWYRPPELF----MGCRSYGTGV---------DMWSVGCIFAELM-LRTPYLPGESDLD 215
Query 75 SPNVIYEKLKSKAISGTLVPDIASGIQ 101
NVI+ L GT P++ +Q
Sbjct 216 QLNVIFRAL------GTPEPEVIKSMQ 236
> YKL161c
Length=433
Score = 30.4 bits (67), Expect = 0.88, Method: Composition-based stats.
Identities = 18/57 (31%), Positives = 24/57 (42%), Gaps = 14/57 (24%)
Query 5 DGRVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFG 61
DG + + W+ AP LL EC + DIWS GC+L E+ G
Sbjct 186 DGFIKGYITSIWYKAPEILLNYQ--ECTKAV------------DIWSTGCILAELLG 228
> At3g17750
Length=1138
Score = 30.4 bits (67), Expect = 0.89, Method: Compositional matrix adjust.
Identities = 14/36 (38%), Positives = 22/36 (61%), Gaps = 2/36 (5%)
Query 26 CSPVECMASRT--TVQLPPLNEKWDIWSMGCVLQEM 59
CS V+ + R + P ++K DIWS+GC+L E+
Sbjct 989 CSYVQSRSYRAPEVILGLPYDKKIDIWSLGCILAEL 1024
> CE03232
Length=294
Score = 30.4 bits (67), Expect = 0.90, Method: Compositional matrix adjust.
Identities = 13/30 (43%), Positives = 17/30 (56%), Gaps = 0/30 (0%)
Query 38 VQLPPLNEKWDIWSMGCVLQEMFGFLSPMH 67
+Q N K D+WS GC+L EM SP +
Sbjct 197 IQESGYNFKSDLWSTGCLLYEMAALQSPFY 226
> CE27894_1
Length=1011
Score = 30.4 bits (67), Expect = 0.94, Method: Compositional matrix adjust.
Identities = 13/22 (59%), Positives = 14/22 (63%), Gaps = 0/22 (0%)
Query 38 VQLPPLNEKWDIWSMGCVLQEM 59
VQ P EK DIWS GC + EM
Sbjct 614 VQNLPYGEKADIWSFGCCIYEM 635
> CE25080
Length=1118
Score = 30.4 bits (67), Expect = 0.94, Method: Compositional matrix adjust.
Identities = 19/86 (22%), Positives = 38/86 (44%), Gaps = 20/86 (23%)
Query 24 LRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPMHVHISNRDSPN-VIYEK 82
+ CS VECM + TT+ MG +++F + V ++N +S + +Y+
Sbjct 350 MNCSTVECMTANTTI-------------MGGYARQLFDVVYLYGVALTNTNSTDPAVYDD 396
Query 83 LKSKAISGTLVPDIASGIQGIARNII 108
+ +VP + QG+ ++
Sbjct 397 VD------VIVPQFVTSFQGMTGKVV 416
> CE27415
Length=302
Score = 30.0 bits (66), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 10/19 (52%), Positives = 14/19 (73%), Gaps = 0/19 (0%)
Query 42 PLNEKWDIWSMGCVLQEMF 60
P + K DIWS GC+L E++
Sbjct 159 PFSVKSDIWSFGCILSELY 177
> CE19353_3
Length=343
Score = 30.0 bits (66), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 10/19 (52%), Positives = 14/19 (73%), Gaps = 0/19 (0%)
Query 42 PLNEKWDIWSMGCVLQEMF 60
P + K DIWS GC+L E++
Sbjct 214 PFSVKSDIWSFGCILSELY 232
> CE17687_1
Length=511
Score = 30.0 bits (66), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 21/51 (41%), Positives = 23/51 (45%), Gaps = 7/51 (13%)
Query 24 LRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSP-----MHVH 69
L SP E A R+ P L DIWS+GCVL M P MH H
Sbjct 134 LLASPPE--AFRSLESYPCLTPSSDIWSVGCVLVTMLTRYPPFLEHYMHFH 182
> YPR054w
Length=388
Score = 30.0 bits (66), Expect = 1.2, Method: Compositional matrix adjust.
Identities = 24/96 (25%), Positives = 41/96 (42%), Gaps = 21/96 (21%)
Query 1 EASADGRVSRISGATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMF 60
++ ++ W+ AP LL P ++ DIW++GC+L E +
Sbjct 199 HSTVQPHITNYVATRWYRAPELLLSNQPY--------------SKSVDIWAVGCILAEFY 244
Query 61 GFLSPMHVHISNRDSPNVIYEKLKSKAISGTLVPDI 96
P+ + RDS + I+E +K + GT DI
Sbjct 245 A-RKPVFM---GRDSMHQIFEIIK---VLGTPDKDI 273
> 7302973
Length=939
Score = 29.6 bits (65), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 12/32 (37%), Positives = 19/32 (59%), Gaps = 0/32 (0%)
Query 48 DIWSMGCVLQEMFGFLSPMHVHISNRDSPNVI 79
D+WS G + EMF +P + ISN D+ ++
Sbjct 849 DVWSFGVTIWEMFSLGAPPYGEISNVDAIKLV 880
> Hs5174701
Length=968
Score = 29.6 bits (65), Expect = 1.3, Method: Composition-based stats.
Identities = 23/75 (30%), Positives = 35/75 (46%), Gaps = 18/75 (24%)
Query 13 GATWFYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGF------LSPM 66
G ++ AP E++ C T++ P + K DIWS+G L EM L+PM
Sbjct 194 GTPYWMAP-EVVMCE---------TMKDTPYDYKADIWSLGITLIEMAQIEPPHHELNPM 243
Query 67 HV--HISNRDSPNVI 79
V I+ D P ++
Sbjct 244 RVLLKIAKSDPPTLL 258
> CE27499
Length=1843
Score = 29.6 bits (65), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 20/63 (31%), Positives = 32/63 (50%), Gaps = 2/63 (3%)
Query 17 FYAPAELLRCSPVECMASRTTVQLPPLNEKWDIWSMGCVLQEMFGFLSPMHVHISNRDSP 76
+Y P+ R PV M S +++ + K D+WS G VL EM + ++ +SN +
Sbjct 1415 YYKPSGK-RMMPVRWM-SPESLKDGKFDSKSDVWSFGVVLYEMVTLGAQPYIGLSNDEVL 1472
Query 77 NVI 79
N I
Sbjct 1473 NYI 1475
> Hs11386209
Length=1215
Score = 29.6 bits (65), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 10/19 (52%), Positives = 14/19 (73%), Gaps = 0/19 (0%)
Query 42 PLNEKWDIWSMGCVLQEMF 60
P E D+WS+GCV+ E+F
Sbjct 375 PFCEAIDMWSLGCVIAELF 393
> At5g27510
Length=301
Score = 29.6 bits (65), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 17/51 (33%), Positives = 27/51 (52%), Gaps = 7/51 (13%)
Query 44 NEKWDIWSMGCVLQEMF-------GFLSPMHVHISNRDSPNVIYEKLKSKA 87
N+ D+WS+GC++ EM+ GF+ V+I + +P I E L A
Sbjct 192 NKTLDLWSLGCLVLEMYVCKKPWIGFIPEDFVYILSNGNPPEIPESLPCDA 242
> Hs4885251_1
Length=395
Score = 29.6 bits (65), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 9/15 (60%), Positives = 13/15 (86%), Gaps = 0/15 (0%)
Query 42 PLNEKWDIWSMGCVL 56
P+ EK DIW++GC+L
Sbjct 239 PIGEKQDIWALGCIL 253
Lambda K H
0.321 0.134 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1192473466
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40