bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_3096_orf2
Length=156
Score E
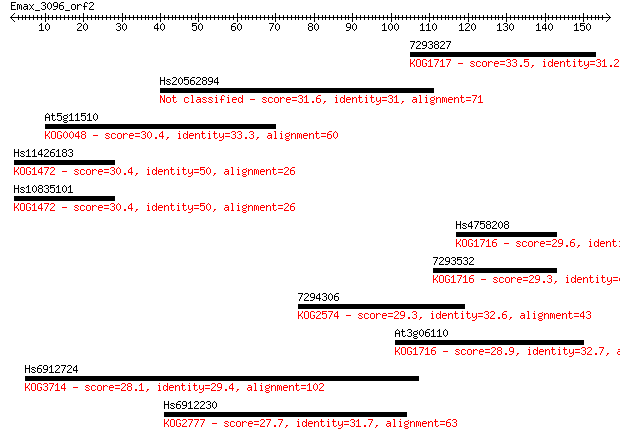
Sequences producing significant alignments: (Bits) Value
7293827 33.5 0.18
Hs20562894 31.6 0.63
At5g11510 30.4 1.4
Hs11426183 30.4 1.6
Hs10835101 30.4 1.6
Hs4758208 29.6 2.2
7293532 29.3 3.4
7294306 29.3 3.6
At3g06110 28.9 4.6
Hs6912724 28.1 6.7
Hs6912230 27.7 8.5
> 7293827
Length=348
Score = 33.5 bits (75), Expect = 0.18, Method: Compositional matrix adjust.
Identities = 15/48 (31%), Positives = 26/48 (54%), Gaps = 1/48 (2%)
Query 105 NFGSVISDVYEGRLFVSGAAAACNLECLQRAGITHVVNTIGDICPNVF 152
N+ ++ G LF+ A +C+ E L++ I +V+N D+ PN F
Sbjct 84 NYNEAPVEIIPGLLFLGNATHSCDSEALKKYNIKYVLNVTPDL-PNKF 130
> Hs20562894
Length=171
Score = 31.6 bits (70), Expect = 0.63, Method: Compositional matrix adjust.
Identities = 22/71 (30%), Positives = 33/71 (46%), Gaps = 9/71 (12%)
Query 40 PARSPGRLHPSGLLFSPAASPGGLQCHSSEGSKAGSSRESFGSRRRRTEEEGMGLYSRTR 99
P R PGRLHP +P A+ GL S+ SR S+ RT E + ++ R
Sbjct 84 PLRMPGRLHP-----TPGANETGLDPRPSDAMSDTESRTSYP----RTIESEILVFIRAT 134
Query 100 QQQLVNFGSVI 110
++F ++I
Sbjct 135 AAITIHFLALI 145
> At5g11510
Length=952
Score = 30.4 bits (67), Expect = 1.4, Method: Composition-based stats.
Identities = 20/63 (31%), Positives = 29/63 (46%), Gaps = 8/63 (12%)
Query 10 SSSSLDLAAAGPSPNQQQQTVSAGARLSGVPARSPGRLHPSGLLFSP---AASPGGLQCH 66
S+S ++ A SP Q S+ +R + A G L P+ L+ SP + GL CH
Sbjct 417 SASDRQISEATKSPTQ-----SSSSRFTATAASGKGTLRPAPLIISPDKYSKKSSGLICH 471
Query 67 SSE 69
E
Sbjct 472 PFE 474
> Hs11426183
Length=837
Score = 30.4 bits (67), Expect = 1.6, Method: Composition-based stats.
Identities = 13/26 (50%), Positives = 17/26 (65%), Gaps = 0/26 (0%)
Query 2 PRAGGGRDSSSSLDLAAAGPSPNQQQ 27
P GGG +SS SLD A A P P +++
Sbjct 409 PSMGGGSNSSLSLDSAGAEPMPGEKR 434
> Hs10835101
Length=837
Score = 30.4 bits (67), Expect = 1.6, Method: Composition-based stats.
Identities = 13/26 (50%), Positives = 17/26 (65%), Gaps = 0/26 (0%)
Query 2 PRAGGGRDSSSSLDLAAAGPSPNQQQ 27
P GGG +SS SLD A A P P +++
Sbjct 409 PSMGGGSNSSLSLDSAGAEPMPGEKR 434
> Hs4758208
Length=185
Score = 29.6 bits (65), Expect = 2.2, Method: Compositional matrix adjust.
Identities = 12/26 (46%), Positives = 19/26 (73%), Gaps = 0/26 (0%)
Query 117 RLFVSGAAAACNLECLQRAGITHVVN 142
R++V A+ A ++ LQ+ GITHV+N
Sbjct 36 RIYVGNASVAQDIPKLQKLGITHVLN 61
> 7293532
Length=245
Score = 29.3 bits (64), Expect = 3.4, Method: Compositional matrix adjust.
Identities = 15/32 (46%), Positives = 21/32 (65%), Gaps = 1/32 (3%)
Query 111 SDVYEGRLFVSGAAAACNLECLQRAGITHVVN 142
+VY G +++ AAAA N L+ GITHV+N
Sbjct 56 DEVYPG-IYIGDAAAAKNKTYLRLMGITHVLN 86
> 7294306
Length=501
Score = 29.3 bits (64), Expect = 3.6, Method: Composition-based stats.
Identities = 14/43 (32%), Positives = 22/43 (51%), Gaps = 0/43 (0%)
Query 76 SRESFGSRRRRTEEEGMGLYSRTRQQQLVNFGSVISDVYEGRL 118
S++ G +R R +E L +Q +NFG + D Y+G L
Sbjct 358 SKKKRGGKRVRKMKERYALTEFRKQANRMNFGDIEEDAYQGDL 400
> At3g06110
Length=167
Score = 28.9 bits (63), Expect = 4.6, Method: Compositional matrix adjust.
Identities = 16/49 (32%), Positives = 28/49 (57%), Gaps = 1/49 (2%)
Query 101 QQLVNFGSVISDVYEGRLFVSGAAAACNLECLQRAGITHVVNTIGDICP 149
Q+L+ G +S++ +G LF+ A A N + L+ + ITHV+ + P
Sbjct 16 QKLLEGGKDLSEIQQG-LFIGSVAEANNKDFLKSSNITHVLTVAVALAP 63
> Hs6912724
Length=1015
Score = 28.1 bits (61), Expect = 6.7, Method: Composition-based stats.
Identities = 30/121 (24%), Positives = 46/121 (38%), Gaps = 21/121 (17%)
Query 5 GGGRDSSSSLDLAAAGPSPNQQQQTVSAGARLSGVPAR-------SPGRLHPSGLLFSPA 57
G S+ L+ A+ SP+ + G + +G R SPG LH + FSP
Sbjct 89 GATGHSTGGLEEQASESSPDTT--AMDTGTKEAGKDGRENTTLLHSPGTLHAAAKTFSPR 146
Query 58 AS-----------PGGLQCHSSEGSKAGSSRESFGSRRRRTEEEG-MGLYSRTRQQQLVN 105
PGG+ + G+ GS R F R E+ + RT ++ +
Sbjct 147 VRRATTSRTERIWPGGVIPYVIGGNFTGSQRAIFKQAMRHWEKHTCVTFIERTDEESFIV 206
Query 106 F 106
F
Sbjct 207 F 207
> Hs6912230
Length=502
Score = 27.7 bits (60), Expect = 8.5, Method: Compositional matrix adjust.
Identities = 20/63 (31%), Positives = 35/63 (55%), Gaps = 1/63 (1%)
Query 41 ARSPGRLHPSGLLFSPAASPGGLQCHSSEGSKAGSSRESFGSRRRRTEEEGMGLYSRTRQ 100
A SPGRL P G S +A P ++ G G+++++ GS + R++ + L+ R+ Q
Sbjct 385 ADSPGRLVPCGAAISWSAVPEQPLDVTANGFPQGTTKKTIGSLQARSQISKVELF-RSFQ 443
Query 101 QQL 103
+ L
Sbjct 444 KLL 446
Lambda K H
0.314 0.131 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2070320142
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40