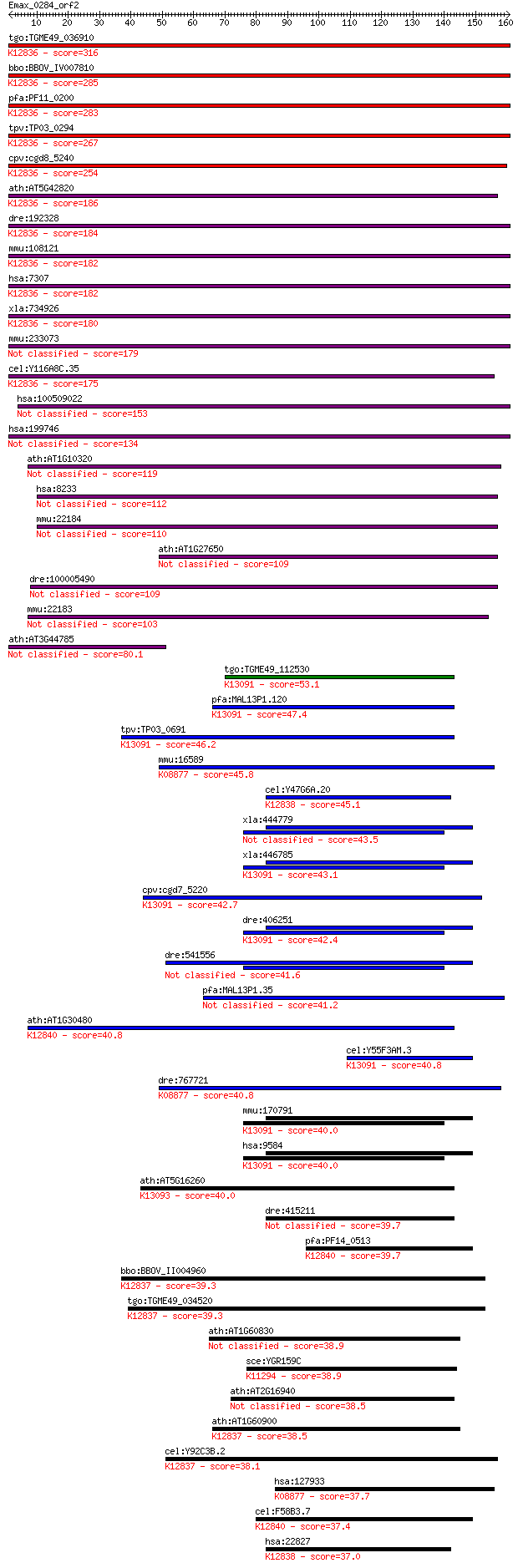
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_865
164,496 sequences; 82,071,388 total letters
Query= Emax_0284_orf2
Length=160
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_036910 U2 snRNP auxiliary factor small subunit, put... 316 2e-86
bbo:BBOV_IV007810 23.m06352; U2 splicing factor subunit; K1283... 285 5e-77
pfa:PF11_0200 U2 snRNP auxiliary factor, small subunit, putati... 283 1e-76
tpv:TP03_0294 U2 small nuclear ribonucleoprotein, auxiliary fa... 267 1e-71
cpv:cgd8_5240 U2AG splicing factor U2AF U2snRNP auxilliary fac... 254 8e-68
ath:AT5G42820 U2AF35B; RNA binding / nucleic acid binding / nu... 186 4e-47
dre:192328 u2af1, CHUNP6860, MGC111896, zgc:111896; U2(RNU2) s... 184 1e-46
mmu:108121 U2af1, 2010107D16Rik, 35kDa; U2 small nuclear ribon... 182 4e-46
hsa:7307 U2AF1, DKFZp313J1712, FP793, RN, RNU2AF1, U2AF35, U2A... 182 4e-46
xla:734926 u2af1, MGC131026; U2 small nuclear RNA auxiliary fa... 180 2e-45
mmu:233073 U2af1l4, AA407033, AF419339, AI451269, AW553050, U2... 179 5e-45
cel:Y116A8C.35 uaf-2; U2AF splicing factor family member (uaf-... 175 6e-44
hsa:100509022 splicing factor U2AF 35 kDa subunit-like 153 2e-37
hsa:199746 U2AF1L4, FLJ35525, MGC33901, U2AF1-RS3, U2AF1L3, U2... 134 1e-31
ath:AT1G10320 U2 snRNP auxiliary factor-related 119 5e-27
hsa:8233 ZRSR2, MGC142014, MGC142040, U2AF1-RS2, U2AF1L2, U2AF... 112 5e-25
mmu:22184 Zrsr2, 35kDa, 5031411E02Rik, A230052C13Rik, C77286, ... 110 1e-24
ath:AT1G27650 ATU2AF35A; RNA binding / nucleic acid binding / ... 109 3e-24
dre:100005490 zrsr2, fb73a09, wu:fb73a09; zinc finger (CCCH ty... 109 4e-24
mmu:22183 Zrsr1, D11Ncvs75, Irlgs2, SP2, U2af1-rs1, U2afbp-rs;... 103 3e-22
ath:AT3G44785 U2AF splicing factor subunit, putative / U2 auxi... 80.1 3e-15
tgo:TGME49_112530 splicing factor protein, putative ; K13091 R... 53.1 4e-07
pfa:MAL13P1.120 splicing factor, putative; K13091 RNA-binding ... 47.4 2e-05
tpv:TP03_0691 splicing factor; K13091 RNA-binding protein 39 46.2
mmu:16589 Uhmk1, 4732477C12Rik, 4930500M09Rik, AA673513, AI449... 45.8 6e-05
cel:Y47G6A.20 rnp-6; RNP (RRM RNA binding domain) containing f... 45.1 9e-05
xla:444779 MGC81970 protein 43.5 3e-04
xla:446785 rbm39, MGC80448, rnpc2; RNA binding motif protein 3... 43.1 4e-04
cpv:cgd7_5220 splicing factor with 3 RRM domains ; K13091 RNA-... 42.7 6e-04
dre:406251 rbm39a, rnpc2, zgc:55780; RNA binding motif protein... 42.4 6e-04
dre:541556 rbm39b, rnpc2l, wu:fa97g07, wu:fb09c08, zgc:112139,... 41.6 0.001
pfa:MAL13P1.35 U1 small nuclear ribonucleoprotein A, putative 41.2 0.002
ath:AT1G30480 DRT111; DRT111; nucleic acid binding / nucleotid... 40.8 0.002
cel:Y55F3AM.3 hypothetical protein; K13091 RNA-binding protein 39 40.8
dre:767721 uhmk1, MGC153241, zgc:153241; U2AF homology motif (... 40.8 0.002
mmu:170791 Rbm39, 1500012C14Rik, 2310040E03Rik, B330012G18Rik,... 40.0 0.003
hsa:9584 RBM39, CAPER, CAPERalpha, CC1.3, DKFZp781C0423, FLJ44... 40.0 0.003
ath:AT5G16260 RNA recognition motif (RRM)-containing protein; ... 40.0 0.003
dre:415211 puf60b, SIAHBP1, fe37c05, repressor, si:ch211-12p8.... 39.7 0.004
pfa:PF14_0513 RNA binding protein, putative; K12840 splicing f... 39.7 0.005
bbo:BBOV_II004960 18.m06413; RNA recognition motif (RRM)-conta... 39.3 0.005
tgo:TGME49_034520 U2 snRNP auxiliary factor or splicing factor... 39.3 0.006
ath:AT1G60830 U2 snRNP auxiliary factor large subunit, putative 38.9 0.007
sce:YGR159C NSR1, SHE5; Nsr1p; K11294 nucleolin 38.9 0.007
ath:AT2G16940 RNA recognition motif (RRM)-containing protein 38.5 0.010
ath:AT1G60900 U2 snRNP auxiliary factor large subunit, putativ... 38.5 0.010
cel:Y92C3B.2 uaf-1; U2AF splicing factor family member (uaf-1)... 38.1 0.011
hsa:127933 UHMK1, DKFZp434C1613, FLJ23015, KIS, KIST; U2AF hom... 37.7 0.015
cel:F58B3.7 hypothetical protein; K12840 splicing factor 45 37.4
hsa:22827 PUF60, FIR, FLJ31379, RoBPI, SIAHBP1; poly-U binding... 37.0 0.027
> tgo:TGME49_036910 U2 snRNP auxiliary factor small subunit, putative
; K12836 splicing factor U2AF 35 kDa subunit
Length=254
Score = 316 bits (809), Expect = 2e-86, Method: Compositional matrix adjust.
Identities = 147/160 (91%), Positives = 156/160 (97%), Gaps = 0/160 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKP+SSPTIVLRHMYPNPPVA+AIA
Sbjct 3 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPTSSPTIVLRHMYPNPPVAVAIA 62
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDA 120
EGQNVSDELLD+AADHFEAFFSEVFEEL KYGEVEDMVVCDNIGDHIIGNVYVKY+D++A
Sbjct 63 EGQNVSDELLDQAADHFEAFFSEVFEELAKYGEVEDMVVCDNIGDHIIGNVYVKYTDEEA 122
Query 121 AKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
A KAL+ALQGR+Y+GK I AEFTPVTDFREARCRQFVDGQ
Sbjct 123 ANKALAALQGRFYSGKQIHAEFTPVTDFREARCRQFVDGQ 162
> bbo:BBOV_IV007810 23.m06352; U2 splicing factor subunit; K12836
splicing factor U2AF 35 kDa subunit
Length=251
Score = 285 bits (729), Expect = 5e-77, Method: Compositional matrix adjust.
Identities = 129/160 (80%), Positives = 146/160 (91%), Gaps = 0/160 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LARIIGTEEDRVNCPFYWKIGACRHGDQCSR+HYKPS++ T+V+RHMY NPPVAIAIA
Sbjct 3 ENLARIIGTEEDRVNCPFYWKIGACRHGDQCSRAHYKPSAAQTLVIRHMYQNPPVAIAIA 62
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDA 120
EGQ +SDELLD+AADHFE F+ EVF EL KYGE+EDMVVCDNIGDHIIGNVYVKY D+++
Sbjct 63 EGQMISDELLDKAADHFEEFYEEVFLELMKYGEIEDMVVCDNIGDHIIGNVYVKYRDENS 122
Query 121 AKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
A A+S L GR+Y GKPIQ E+TPVTDFREARCRQFV+GQ
Sbjct 123 AAHAISMLSGRFYGGKPIQCEYTPVTDFREARCRQFVEGQ 162
> pfa:PF11_0200 U2 snRNP auxiliary factor, small subunit, putative;
K12836 splicing factor U2AF 35 kDa subunit
Length=294
Score = 283 bits (725), Expect = 1e-76, Method: Compositional matrix adjust.
Identities = 125/160 (78%), Positives = 147/160 (91%), Gaps = 0/160 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
EHLARIIGTEEDRVNCPF+WKIGACRHGDQCSRSHYKP+ + T+V+RHMY NPP+A+AIA
Sbjct 3 EHLARIIGTEEDRVNCPFFWKIGACRHGDQCSRSHYKPNCAQTLVIRHMYDNPPIAVAIA 62
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDA 120
EGQ V DE+LD+AADHFE F+ EVF+EL KYGE+EDMVVCDNIGDHIIGNVY+KY+ +D
Sbjct 63 EGQMVEDEVLDKAADHFEEFYEEVFDELMKYGEIEDMVVCDNIGDHIIGNVYIKYTHEDY 122
Query 121 AKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
A+KA++ L GR+YAGKP+Q E+TPVTDFREARCRQFV+GQ
Sbjct 123 AEKAVNELNGRFYAGKPLQIEYTPVTDFREARCRQFVEGQ 162
> tpv:TP03_0294 U2 small nuclear ribonucleoprotein, auxiliary
factor, small subunit; K12836 splicing factor U2AF 35 kDa subunit
Length=235
Score = 267 bits (682), Expect = 1e-71, Method: Compositional matrix adjust.
Identities = 129/160 (80%), Positives = 148/160 (92%), Gaps = 0/160 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
EHLARIIGTEEDRVNCPF+WKIGACRHGDQCSR+HYKPS++ T+V+RHMY NPPVAIAIA
Sbjct 3 EHLARIIGTEEDRVNCPFFWKIGACRHGDQCSRTHYKPSAAQTLVIRHMYQNPPVAIAIA 62
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDA 120
EGQ +SDELLD+AADHFE FF EVF EL KYGE+EDM+VCDNIGDHIIGNVY+KYSD+ A
Sbjct 63 EGQMISDELLDKAADHFEEFFEEVFLELMKYGEIEDMIVCDNIGDHIIGNVYIKYSDEAA 122
Query 121 AKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
A +A+++L GRYY G+PIQ E+TPVTDFREARCRQFV+GQ
Sbjct 123 ACRAVTSLSGRYYGGRPIQCEYTPVTDFREARCRQFVEGQ 162
> cpv:cgd8_5240 U2AG splicing factor U2AF U2snRNP auxilliary factor
small subunit CCCh+RRM+CCCh-like ; K12836 splicing factor
U2AF 35 kDa subunit
Length=256
Score = 254 bits (649), Expect = 8e-68, Method: Compositional matrix adjust.
Identities = 108/159 (67%), Positives = 140/159 (88%), Gaps = 0/159 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
EHLARI+GTEEDRVNCPFYWKIGACRHGDQCSR+HYKP+SSPT+++RH+Y N PVA+AIA
Sbjct 10 EHLARILGTEEDRVNCPFYWKIGACRHGDQCSRNHYKPTSSPTVIIRHIYENSPVALAIA 69
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDA 120
EGQ VSD+L DE +D E F+ E+F+EL KYGE+ ++++CDNIGDH+IGNVY+++S ++
Sbjct 70 EGQEVSDKLADEESDKVEVFYEEMFKELSKYGEILELLICDNIGDHMIGNVYIRFSTEEY 129
Query 121 AKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDG 159
AK AL+ L+G+ YAGKPI E +PV+DF+EARCRQ++DG
Sbjct 130 AKTALANLRGKMYAGKPINIELSPVSDFKEARCRQYIDG 168
> ath:AT5G42820 U2AF35B; RNA binding / nucleic acid binding /
nucleotide binding / zinc ion binding; K12836 splicing factor
U2AF 35 kDa subunit
Length=283
Score = 186 bits (471), Expect = 4e-47, Method: Compositional matrix adjust.
Identities = 93/158 (58%), Positives = 121/158 (76%), Gaps = 5/158 (3%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
EHLA I GTE+DRVNCPFY+KIGACRHGD+CSR H +P+ SPT++L +MY P + I
Sbjct 3 EHLASIFGTEKDRVNCPFYFKIGACRHGDRCSRLHNRPTISPTLLLSNMYQRPDM---IT 59
Query 61 EGQNVSDELLDEAA--DHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDD 118
G + + LD + DHFE F+ ++FEEL K+GEVE + VCDN+ DH+IGNVYV + ++
Sbjct 60 PGVDPQGQPLDPSKIQDHFEDFYEDIFEELNKFGEVESLNVCDNLADHMIGNVYVLFKEE 119
Query 119 DAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQF 156
D A AL ALQGR+Y+G+PI A+F+PVTDFREA CRQ+
Sbjct 120 DHAAAALQALQGRFYSGRPIIADFSPVTDFREATCRQY 157
> dre:192328 u2af1, CHUNP6860, MGC111896, zgc:111896; U2(RNU2)
small nuclear RNA auxiliary factor 1; K12836 splicing factor
U2AF 35 kDa subunit
Length=249
Score = 184 bits (466), Expect = 1e-46, Method: Compositional matrix adjust.
Identities = 88/161 (54%), Positives = 118/161 (73%), Gaps = 3/161 (1%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC FY+KIGACRHGD+CSR H KP+ S TI L ++Y NP A
Sbjct 3 EYLASIFGTEKDKVNCSFYFKIGACRHGDRCSRLHNKPTFSQTIALLNIYRNPQNTAQSA 62
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELY-KYGEVEDMVVCDNIGDHIIGNVYVKYSDDD 119
+G N ++ E +H++ FF EVF E+ KYGEVE+M VCDN+GDH++GNVYVK+ ++
Sbjct 63 DGLNAVSDV--EMQEHYDEFFEEVFTEMEEKYGEVEEMNVCDNLGDHLVGNVYVKFRREE 120
Query 120 AAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
A+KA+ L R++ G+PI AE +PVTDFREA CRQ+ G+
Sbjct 121 DAEKAVINLNNRWFNGQPIHAELSPVTDFREACCRQYEMGE 161
> mmu:108121 U2af1, 2010107D16Rik, 35kDa; U2 small nuclear ribonucleoprotein
auxiliary factor (U2AF) 1; K12836 splicing factor
U2AF 35 kDa subunit
Length=239
Score = 182 bits (462), Expect = 4e-46, Method: Compositional matrix adjust.
Identities = 89/163 (54%), Positives = 122/163 (74%), Gaps = 6/163 (3%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC FY+KIGACRHGD+CSR H KP+ S TI+++++Y NP + A
Sbjct 3 EYLASIFGTEKDKVNCSFYFKIGACRHGDRCSRLHNKPTFSQTILIQNIYRNPQNSAQTA 62
Query 61 EGQN--VSDELLDEAADHFEAFFSEVFEEL-YKYGEVEDMVVCDNIGDHIIGNVYVKYSD 117
+G + VSD E +H++ FF EVF E+ KYGEVE+M VCDN+GDH++GNVYVK+
Sbjct 63 DGSHCAVSDV---EMQEHYDEFFEEVFTEMEEKYGEVEEMNVCDNLGDHLVGNVYVKFRR 119
Query 118 DDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
++ A+KA+ L R++ G+PI AE +PVTDFREA CRQ+ G+
Sbjct 120 EEDAEKAVIDLNNRWFNGQPIHAELSPVTDFREACCRQYEMGE 162
> hsa:7307 U2AF1, DKFZp313J1712, FP793, RN, RNU2AF1, U2AF35, U2AFBP;
U2 small nuclear RNA auxiliary factor 1; K12836 splicing
factor U2AF 35 kDa subunit
Length=240
Score = 182 bits (462), Expect = 4e-46, Method: Compositional matrix adjust.
Identities = 89/163 (54%), Positives = 122/163 (74%), Gaps = 6/163 (3%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC FY+KIGACRHGD+CSR H KP+ S TI+++++Y NP + A
Sbjct 3 EYLASIFGTEKDKVNCSFYFKIGACRHGDRCSRLHNKPTFSQTILIQNIYRNPQNSAQTA 62
Query 61 EGQN--VSDELLDEAADHFEAFFSEVFEEL-YKYGEVEDMVVCDNIGDHIIGNVYVKYSD 117
+G + VSD E +H++ FF EVF E+ KYGEVE+M VCDN+GDH++GNVYVK+
Sbjct 63 DGSHCAVSDV---EMQEHYDEFFEEVFTEMEEKYGEVEEMNVCDNLGDHLVGNVYVKFRR 119
Query 118 DDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
++ A+KA+ L R++ G+PI AE +PVTDFREA CRQ+ G+
Sbjct 120 EEDAEKAVIDLNNRWFNGQPIHAELSPVTDFREACCRQYEMGE 162
> xla:734926 u2af1, MGC131026; U2 small nuclear RNA auxiliary
factor 1; K12836 splicing factor U2AF 35 kDa subunit
Length=245
Score = 180 bits (456), Expect = 2e-45, Method: Compositional matrix adjust.
Identities = 89/163 (54%), Positives = 119/163 (73%), Gaps = 6/163 (3%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC FY+KIGACRHGD+CSR H KP+ S TI L ++Y NP + A
Sbjct 3 EYLASIFGTEKDKVNCSFYFKIGACRHGDRCSRLHNKPTFSQTIALLNIYRNPQNSSQSA 62
Query 61 EGQN--VSDELLDEAADHFEAFFSEVFEEL-YKYGEVEDMVVCDNIGDHIIGNVYVKYSD 117
+G VSD E +H++ FF EVF E+ KYGE+E+M VCDN+GDH++GNVYVK+
Sbjct 63 DGLRCAVSDV---EMQEHYDEFFEEVFTEMEEKYGEIEEMNVCDNLGDHLVGNVYVKFRR 119
Query 118 DDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
++ A+KA+ L R++ G+PI AE +PVTDFREA CRQ+ G+
Sbjct 120 EEDAEKAVKDLNNRWFNGQPIHAELSPVTDFREACCRQYEMGE 162
> mmu:233073 U2af1l4, AA407033, AF419339, AI451269, AW553050,
U2af26; U2 small nuclear RNA auxiliary factor 1-like 4
Length=220
Score = 179 bits (453), Expect = 5e-45, Method: Compositional matrix adjust.
Identities = 87/163 (53%), Positives = 120/163 (73%), Gaps = 6/163 (3%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC FY+KIGACRHGD+CSR H KP+ S TIVL ++Y NP A
Sbjct 3 EYLASIFGTEKDKVNCSFYFKIGACRHGDRCSRLHNKPTFSQTIVLLNLYRNPQNTAQTA 62
Query 61 EGQN--VSDELLDEAADHFEAFFSEVFEELY-KYGEVEDMVVCDNIGDHIIGNVYVKYSD 117
+G + VSD E +H++ FF EVF EL KYGE+E+M VCDN+GDH++GNVYVK+
Sbjct 63 DGSHCHVSDV---EVQEHYDNFFEEVFTELQEKYGEIEEMNVCDNLGDHLVGNVYVKFRR 119
Query 118 DDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
++ A++A++ L R++ G+ + AE +PVTDFRE+ CRQ+ G+
Sbjct 120 EEDAERAVAELNNRWFNGQAVHAELSPVTDFRESCCRQYEMGE 162
> cel:Y116A8C.35 uaf-2; U2AF splicing factor family member (uaf-2);
K12836 splicing factor U2AF 35 kDa subunit
Length=285
Score = 175 bits (443), Expect = 6e-44, Method: Compositional matrix adjust.
Identities = 81/156 (51%), Positives = 117/156 (75%), Gaps = 1/156 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC F++K GACRHGD+CSR+H+ P+ SPT+VL++ Y NP V + A
Sbjct 11 EYLASIYGTEKDKVNCSFFFKTGACRHGDKCSRAHHTPTFSPTVVLKNFYHNPVVDVRQA 70
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEEL-YKYGEVEDMVVCDNIGDHIIGNVYVKYSDDD 119
+ + + D+ +F+ F+ EVF E+ KYGEVE++ VC+NIG+H++GNVYVK+ ++
Sbjct 71 DAFDKVGKRNDQEQRYFDDFYEEVFVEMERKYGEVEEINVCENIGEHMVGNVYVKFMKEE 130
Query 120 AAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQ 155
A+KA + L R++ G+PI AE PVTDFRE+RCRQ
Sbjct 131 DAEKAKNDLNNRWFNGQPIYAELCPVTDFRESRCRQ 166
> hsa:100509022 splicing factor U2AF 35 kDa subunit-like
Length=219
Score = 153 bits (387), Expect = 2e-37, Method: Compositional matrix adjust.
Identities = 83/159 (52%), Positives = 109/159 (68%), Gaps = 6/159 (3%)
Query 4 ARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIAEGQ 63
A I GTE +VNC F +KIGAC HGD+CS H+KP+ S TI L ++Y NP A A+G
Sbjct 6 ASIFGTE--KVNCSFDFKIGACLHGDRCSWLHHKPTFSQTIALLNIYCNPQNASQSADGL 63
Query 64 NVSDELLD-EAADHFEAFFSEVFEEL-YKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAA 121
+ L D E +H++ FF EVF EL KYGEVE+M VCDN+GDH++GNVY K ++ A
Sbjct 64 RCA--LSDVEVQEHYDEFFEEVFIELGEKYGEVEEMNVCDNLGDHLVGNVYFKLPREEDA 121
Query 122 KKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
+KA+ L R++ G+PI AE +PVTDFR A CRQ+ G+
Sbjct 122 EKAVIDLNNRWFNGQPIHAELSPVTDFRGACCRQYEMGE 160
> hsa:199746 U2AF1L4, FLJ35525, MGC33901, U2AF1-RS3, U2AF1L3,
U2AF1L3V1, U2AF1RS3, U2af26; U2 small nuclear RNA auxiliary
factor 1-like 4
Length=181
Score = 134 bits (336), Expect = 1e-31, Method: Compositional matrix adjust.
Identities = 68/161 (42%), Positives = 94/161 (58%), Gaps = 41/161 (25%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIA 60
E+LA I GTE+D+VNC FY+KIG CRHGD+CSR H KP+
Sbjct 3 EYLASIFGTEKDKVNCSFYFKIGVCRHGDRCSRLHNKPT--------------------- 41
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELY-KYGEVEDMVVCDNIGDHIIGNVYVKYSDDD 119
F EVF EL KYGE+E+M VCDN+GDH++GNVYVK+ ++
Sbjct 42 -------------------FSQEVFTELQEKYGEIEEMNVCDNLGDHLVGNVYVKFRREE 82
Query 120 AAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFVDGQ 160
++A++ L R++ G+ + E +PVTDFRE+ CRQ+ G+
Sbjct 83 DGERAVAELSNRWFNGQAVHGELSPVTDFRESCCRQYEMGE 123
> ath:AT1G10320 U2 snRNP auxiliary factor-related
Length=757
Score = 119 bits (298), Expect = 5e-27, Method: Compositional matrix adjust.
Identities = 59/151 (39%), Positives = 92/151 (60%), Gaps = 3/151 (1%)
Query 7 IGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIAEGQNVS 66
GTE+D+ +CPF+ K GACR G +CSR H+ P+ S T+++++MY P + EG +
Sbjct 237 FGTEQDKAHCPFHLKTGACRFGQRCSRVHFYPNKSCTLLMKNMYNGPGITWEQDEGLEYT 296
Query 67 DELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALS 126
DE EA +E F+ +V E KYGE+ + VC N H+ GNVYV Y ++A A
Sbjct 297 DE---EAELCYEEFYEDVHTEFLKYGELVNFKVCRNGSFHLKGNVYVHYRSLESAILAYQ 353
Query 127 ALQGRYYAGKPIQAEFTPVTDFREARCRQFV 157
++ GRY+AGK + EF ++ ++ A C +++
Sbjct 354 SINGRYFAGKQVNCEFVNISRWKVAICGEYM 384
> hsa:8233 ZRSR2, MGC142014, MGC142040, U2AF1-RS2, U2AF1L2, U2AF1RS2,
URP; zinc finger (CCCH type), RNA-binding motif and
serine/arginine rich 2
Length=482
Score = 112 bits (280), Expect = 5e-25, Method: Composition-based stats.
Identities = 60/159 (37%), Positives = 87/159 (54%), Gaps = 21/159 (13%)
Query 10 EEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYP------------NPPVAI 57
E+DR NCPFY K GACR GD+CSR H P+SSPT++++ M+ +P ++
Sbjct 166 EKDRANCPFYSKTGACRFGDRCSRKHNFPTSSPTLLIKSMFTTFGMEQCRRDDYDPDASL 225
Query 58 AIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSD 117
+E +E F F+ +V E G+V V N+ H+ GNVYV+Y
Sbjct 226 EYSE---------EETYQQFLDFYEDVLPEFKNVGKVIQFKVSCNLEPHLRGNVYVQYQS 276
Query 118 DDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQF 156
++ + ALS GR+YAG+ +Q EF PVT ++ A C F
Sbjct 277 EEECQAALSLFNGRWYAGRQLQCEFCPVTRWKMAICGLF 315
> mmu:22184 Zrsr2, 35kDa, 5031411E02Rik, A230052C13Rik, C77286,
U2af1-rs2, URP; zinc finger (CCCH type), RNA binding motif
and serine/arginine rich 2
Length=541
Score = 110 bits (276), Expect = 1e-24, Method: Compositional matrix adjust.
Identities = 59/159 (37%), Positives = 86/159 (54%), Gaps = 21/159 (13%)
Query 10 EEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYP------------NPPVAI 57
E+DR NCPFY K GACR GD+CSR H P+SSPT++++ M+ +P ++
Sbjct 170 EKDRANCPFYSKTGACRFGDRCSRKHNFPTSSPTLLIKGMFTTFGMEQCRRDDYDPDSSL 229
Query 58 AIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSD 117
+E +E F F+ +V E G+V V N+ H+ GNVYV+Y
Sbjct 230 EFSE---------EEIYQQFLDFYYDVLPEFKSVGKVIQFKVSCNLEPHLRGNVYVQYQS 280
Query 118 DDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQF 156
++ + A S GR+YAG+ +Q EF PVT ++ A C F
Sbjct 281 EEDCQAAFSVFNGRWYAGRQLQCEFCPVTRWKMAICGLF 319
> ath:AT1G27650 ATU2AF35A; RNA binding / nucleic acid binding
/ nucleotide binding / zinc ion binding
Length=246
Score = 109 bits (273), Expect = 3e-24, Method: Compositional matrix adjust.
Identities = 57/110 (51%), Positives = 80/110 (72%), Gaps = 5/110 (4%)
Query 49 MYPNPPVAIAIAEGQNVSDELLD--EAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDH 106
MY P + I G + + LD + +HFE FF ++FEEL K+GE+E + +CDN+ DH
Sbjct 1 MYQRPDM---ITPGVDAQGQPLDPRKIQEHFEDFFEDLFEELGKFGEIESLNICDNLADH 57
Query 107 IIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQF 156
+IGNVYV++ ++D A AL ALQGR+Y+G+PI A+F+PVTDFREA CRQ+
Sbjct 58 MIGNVYVQFKEEDQAAAALQALQGRFYSGRPIIADFSPVTDFREATCRQY 107
> dre:100005490 zrsr2, fb73a09, wu:fb73a09; zinc finger (CCCH
type), RNA-binding motif and serine/arginine rich 2
Length=635
Score = 109 bits (272), Expect = 4e-24, Method: Compositional matrix adjust.
Identities = 59/156 (37%), Positives = 87/156 (55%), Gaps = 11/156 (7%)
Query 8 GTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIAEGQNV-- 65
GTE+D+ NCPF+ K GACR GD+CSR H P+SS T+++R M+ V+ + + +
Sbjct 177 GTEKDKANCPFFLKTGACRFGDRCSRKHDHPASSCTLMVRGMF----VSFGMEQSRRDDY 232
Query 66 -SDELLD----EAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDA 120
+D L+ E F F+ + E G V V N H+ GNVYV+Y ++
Sbjct 233 DTDASLEYSEEELHQQFLDFYEDALPEFKNAGRVVQFKVSCNFEPHLRGNVYVQYETEEQ 292
Query 121 AKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQF 156
K+A GR+YAG+ +Q EF+PVT ++ A C F
Sbjct 293 CKEAFVMFNGRWYAGRQLQCEFSPVTRWKTAICGLF 328
> mmu:22183 Zrsr1, D11Ncvs75, Irlgs2, SP2, U2af1-rs1, U2afbp-rs;
zinc finger (CCCH type), RNA binding motif and serine/arginine
rich 1
Length=428
Score = 103 bits (256), Expect = 3e-22, Method: Compositional matrix adjust.
Identities = 55/150 (36%), Positives = 84/150 (56%), Gaps = 3/150 (2%)
Query 7 IGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPNPPVAIAIAEGQNVS 66
+ E+ R +CPFY K GACR G++CSR H P+SSPT++++ M+ + + +
Sbjct 154 LRLEKYRPSCPFYNKTGACRFGNRCSRKHDFPTSSPTLLVKSMFTTFGMEQCRRDDYDSD 213
Query 67 DELL---DEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKK 123
L +E F F+ +V E G+V V N+ H+ GNVYV+Y ++ +
Sbjct 214 ANLEYSEEETYQQFLDFYHDVLPEFKNVGKVIQFKVSCNLEPHLRGNVYVQYQSEEECQA 273
Query 124 ALSALQGRYYAGKPIQAEFTPVTDFREARC 153
ALS GR+YAG+ +Q EF PVT ++ A C
Sbjct 274 ALSLFNGRWYAGRQLQCEFCPVTRWKVAIC 303
> ath:AT3G44785 U2AF splicing factor subunit, putative / U2 auxiliary
factor 38 kDa subunit, putative
Length=75
Score = 80.1 bits (196), Expect = 3e-15, Method: Compositional matrix adjust.
Identities = 34/50 (68%), Positives = 42/50 (84%), Gaps = 0/50 (0%)
Query 1 EHLARIIGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMY 50
EHLA I GTE+DRVNCPFY+KIG CR+GD+CSR + KPS SPT++L + Y
Sbjct 3 EHLASIYGTEKDRVNCPFYFKIGVCRNGDRCSRLYTKPSISPTLLLSNTY 52
> tgo:TGME49_112530 splicing factor protein, putative ; K13091
RNA-binding protein 39
Length=633
Score = 53.1 bits (126), Expect = 4e-07, Method: Compositional matrix adjust.
Identities = 28/73 (38%), Positives = 43/73 (58%), Gaps = 3/73 (4%)
Query 70 LDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQ 129
L E F +V +E K+G VE + + + ++ GNV+++++ D A+ A AL
Sbjct 553 LKEDPHFFLDLGDDVRDECKKFGSVEKVWIDER---NVDGNVWIRFAHPDQARAAFGALN 609
Query 130 GRYYAGKPIQAEF 142
GRY+AGKPI AEF
Sbjct 610 GRYFAGKPISAEF 622
> pfa:MAL13P1.120 splicing factor, putative; K13091 RNA-binding
protein 39
Length=864
Score = 47.4 bits (111), Expect = 2e-05, Method: Composition-based stats.
Identities = 27/77 (35%), Positives = 42/77 (54%), Gaps = 3/77 (3%)
Query 66 SDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKAL 125
+DE + D F +V EE KYG+V ++ + +I G +Y+KYS++D + K+
Sbjct 778 NDENIGSDPDFFNDILEDVKEECSKYGKVVNIWLDTK---NIDGKIYIKYSNNDESLKSF 834
Query 126 SALQGRYYAGKPIQAEF 142
L GRY+ G I A F
Sbjct 835 QFLNGRYFGGSLINAYF 851
> tpv:TP03_0691 splicing factor; K13091 RNA-binding protein 39
Length=644
Score = 46.2 bits (108), Expect = 4e-05, Method: Composition-based stats.
Identities = 33/106 (31%), Positives = 51/106 (48%), Gaps = 18/106 (16%)
Query 37 KPSSSPTIVLRHMYPNPPVAIAIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVED 96
+P +S +VL +MY + ++ + F+ +V EE KYG V
Sbjct 544 QPLNSSNLVLSNMYTSAD---------------YEDNREFFDEIEEDVKEECGKYGTVIQ 588
Query 97 MVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
+ V D G VYVK+ ++D A+ A +LQGRY+AG IQ +
Sbjct 589 VFVNKRNPD---GKVYVKFKNNDDAQAANKSLQGRYFAGNTIQVSY 631
> mmu:16589 Uhmk1, 4732477C12Rik, 4930500M09Rik, AA673513, AI449218,
AU021979, KIS, Kist; U2AF homology motif (UHM) kinase
1 (EC:2.7.11.1); K08877 U2AF homology motif (UHM) kinase 1
[EC:2.7.11.1]
Length=419
Score = 45.8 bits (107), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 34/108 (31%), Positives = 53/108 (49%), Gaps = 9/108 (8%)
Query 49 MYPNPPVAIAIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVC-DNIGDHI 107
M P P + + NV D+ E D +E +V EE KYG V ++V +N G
Sbjct 317 MLPTPVLRLL-----NVLDDDYLENEDEYEDVVEDVKEECQKYGPVVSLLVPKENPGR-- 369
Query 108 IGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQ 155
G V+V+Y++ +K A L GR + GK + A F P++ ++ Q
Sbjct 370 -GQVFVEYANAGDSKAAQKLLTGRMFDGKFVVATFYPLSAYKRGYLYQ 416
> cel:Y47G6A.20 rnp-6; RNP (RRM RNA binding domain) containing
family member (rnp-6); K12838 poly(U)-binding-splicing factor
PUF60
Length=749
Score = 45.1 bits (105), Expect = 9e-05, Method: Composition-based stats.
Identities = 23/59 (38%), Positives = 35/59 (59%), Gaps = 1/59 (1%)
Query 83 EVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAE 141
E+ EE KYG V D+V+ N + ++VKYSD +A +AL GR++ G ++AE
Sbjct 678 EIREECGKYGNVIDVVIA-NFASSGLVKIFVKYSDSMQVDRAKAALDGRFFGGNTVKAE 735
> xla:444779 MGC81970 protein
Length=512
Score = 43.5 bits (101), Expect = 3e-04, Method: Compositional matrix adjust.
Identities = 27/66 (40%), Positives = 37/66 (56%), Gaps = 3/66 (4%)
Query 83 EVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
+V EE K+G + V N GNVYVK S +A A++AL GR++AGK I A +
Sbjct 431 DVMEECNKHGGAIHIYVDKNSPQ---GNVYVKCSTITSAIAAVNALHGRWFAGKMITAAY 487
Query 143 TPVTDF 148
PV +
Sbjct 488 VPVPTY 493
Score = 33.1 bits (74), Expect = 0.42, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 35/68 (51%), Gaps = 7/68 (10%)
Query 76 HF---EAFFSEVFEELYKYGEVEDM-VVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGR 131
HF E +FE +G +E + ++ D+ G ++ +SD + AKKAL L G
Sbjct 231 HFNITEDMLRGIFEP---FGRIESIQLMMDSETGRSKGYGFITFSDSECAKKALEQLNGF 287
Query 132 YYAGKPIQ 139
AG+P++
Sbjct 288 ELAGRPMK 295
> xla:446785 rbm39, MGC80448, rnpc2; RNA binding motif protein
39; K13091 RNA-binding protein 39
Length=540
Score = 43.1 bits (100), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 26/66 (39%), Positives = 37/66 (56%), Gaps = 3/66 (4%)
Query 83 EVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
+V EE K+G V + V N GNVYVK +A A++AL GR++AGK I A +
Sbjct 459 DVIEECNKHGGVVHLYVDKNSAQ---GNVYVKCPTIASAIAAVNALHGRWFAGKMITAAY 515
Query 143 TPVTDF 148
P+ +
Sbjct 516 VPLPTY 521
Score = 32.7 bits (73), Expect = 0.59, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 35/68 (51%), Gaps = 7/68 (10%)
Query 76 HF---EAFFSEVFEELYKYGEVEDM-VVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGR 131
HF E +FE +G +E + ++ D+ G ++ +SD + AKKAL L G
Sbjct 257 HFNITEDMLRGIFE---PFGRIESIQLMMDSETGRSKGYGFITFSDSECAKKALEQLNGF 313
Query 132 YYAGKPIQ 139
AG+P++
Sbjct 314 ELAGRPMK 321
> cpv:cgd7_5220 splicing factor with 3 RRM domains ; K13091 RNA-binding
protein 39
Length=563
Score = 42.7 bits (99), Expect = 6e-04, Method: Composition-based stats.
Identities = 30/108 (27%), Positives = 55/108 (50%), Gaps = 11/108 (10%)
Query 44 IVLRHMYPNPPVAIAIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNI 103
++L +M+ + ++ E DE +++ + +A +V EE KYG ++ C
Sbjct 464 LLLSNMFTEQSIKESMEE-----DETIEQILEEIQA---DVEEECGKYGT---LLECFLD 512
Query 104 GDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDFREA 151
+ + GNV+VKYS + A KA GR++AG+ + F +F +A
Sbjct 513 KEKMDGNVWVKYSRPEEASKAKMVFHGRFFAGRKLNVSFIKDEEFPKA 560
> dre:406251 rbm39a, rnpc2, zgc:55780; RNA binding motif protein
39a; K13091 RNA-binding protein 39
Length=523
Score = 42.4 bits (98), Expect = 6e-04, Method: Compositional matrix adjust.
Identities = 26/66 (39%), Positives = 36/66 (54%), Gaps = 3/66 (4%)
Query 83 EVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
+V EE K+G V + V + GNVYVK AA A+SAL GR++ GK I A +
Sbjct 442 DVIEECNKHGGVIHIYVDKKSAE---GNVYVKCPTIPAAMAAVSALHGRWFGGKMITAAY 498
Query 143 TPVTDF 148
P+ +
Sbjct 499 VPLPTY 504
Score = 31.2 bits (69), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 20/67 (29%), Positives = 33/67 (49%), Gaps = 5/67 (7%)
Query 76 HF---EAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRY 132
HF E +FE + ++ M+ D+ G ++ +SD + AKKAL L G
Sbjct 254 HFNITEDMLRGIFEPFGRIDSIQLMM--DSETGRSKGYGFITFSDAECAKKALEQLNGFE 311
Query 133 YAGKPIQ 139
AG+P++
Sbjct 312 LAGRPMK 318
> dre:541556 rbm39b, rnpc2l, wu:fa97g07, wu:fb09c08, zgc:112139,
zgc:113117; RNA binding motif protein 39b
Length=539
Score = 41.6 bits (96), Expect = 0.001, Method: Composition-based stats.
Identities = 29/98 (29%), Positives = 45/98 (45%), Gaps = 3/98 (3%)
Query 51 PNPPVAIAIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGN 110
P P+A + N+ + ++ +V EE K+G V + V N GN
Sbjct 426 PTQPLATHCLQLSNMFNPQMENEPGWDIEIRDDVIEECRKHGGVIHIYVDKNSAQ---GN 482
Query 111 VYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDF 148
VYVK A +S+L GR++AGK I A + P+ +
Sbjct 483 VYVKCPTIPVAMAVVSSLHGRWFAGKMITAAYVPLPTY 520
Score = 29.6 bits (65), Expect = 4.8, Method: Composition-based stats.
Identities = 20/68 (29%), Positives = 36/68 (52%), Gaps = 7/68 (10%)
Query 76 HF---EAFFSEVFEELYKYGEVEDM-VVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGR 131
HF E +FE +G++E + ++ D+ G ++ ++D + AKKAL L G
Sbjct 272 HFNITEDMLRGIFE---PFGKIEGIQLMMDSETGRSKGYGFISFADAECAKKALEQLNGF 328
Query 132 YYAGKPIQ 139
AG+P++
Sbjct 329 ELAGRPMK 336
> pfa:MAL13P1.35 U1 small nuclear ribonucleoprotein A, putative
Length=449
Score = 41.2 bits (95), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 30/98 (30%), Positives = 49/98 (50%), Gaps = 11/98 (11%)
Query 63 QNVSDEL-LDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAA 121
+N++D + DE + + F+ YGE++D++V + G +V Y D + A
Sbjct 180 KNLNDRVKTDEMKKNLKDLFNT-------YGEIKDLIVMKSFWRK--GQAWVVYDDKECA 230
Query 122 KKALSALQGRYYAGKPIQAEFT-PVTDFREARCRQFVD 158
KAL+ALQG GK +Q F+ +D R FV+
Sbjct 231 TKALNALQGYVLFGKIMQINFSHNKSDIHAKRDGTFVE 268
> ath:AT1G30480 DRT111; DRT111; nucleic acid binding / nucleotide
binding; K12840 splicing factor 45
Length=387
Score = 40.8 bits (94), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 41/146 (28%), Positives = 63/146 (43%), Gaps = 22/146 (15%)
Query 7 IGTEEDRVNCPFYWKIGACRHGDQCSRSHYKPSSSPTIVLRHMYPN-PPVAI-----AIA 60
+G E + P K R G + S K SS+ V++ + N P + +
Sbjct 233 LGKSEQGITTPLMAKKTDRRAGVIVNASENKSSSAEKKVVKSVNINGEPTRVLLLRNMVG 292
Query 61 EGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCD----NIGDHIIGNVYVKYS 116
GQ V DEL DE V E KYG V +++ + N H ++V++S
Sbjct 293 PGQ-VDDELEDE-----------VGGECGKYGTVTRVLIFEITEPNFPVHEAVRIFVQFS 340
Query 117 DDDAAKKALSALQGRYYAGKPIQAEF 142
+ KAL L GRY+ G+ ++A F
Sbjct 341 RPEETTKALVDLDGRYFGGRTVRATF 366
> cel:Y55F3AM.3 hypothetical protein; K13091 RNA-binding protein
39
Length=580
Score = 40.8 bits (94), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 19/40 (47%), Positives = 26/40 (65%), Gaps = 0/40 (0%)
Query 109 GNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDF 148
GNVYVK A +A+SAL GR+++GK I A + PV +
Sbjct 470 GNVYVKCPSIVIAHQAVSALHGRWFSGKVITANYVPVNSY 509
> dre:767721 uhmk1, MGC153241, zgc:153241; U2AF homology motif
(UHM) kinase 1 (EC:2.7.11.1); K08877 U2AF homology motif (UHM)
kinase 1 [EC:2.7.11.1]
Length=410
Score = 40.8 bits (94), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 31/110 (28%), Positives = 53/110 (48%), Gaps = 9/110 (8%)
Query 49 MYPNPPVAIAIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVV-CDNIGDHI 107
+ P P + + NV D+ D +E ++ EE KYG V +++ +N G
Sbjct 308 LLPTPVLRLL-----NVIDDSHLYNEDEYEDIIEDMKEECQKYGTVVSLLIPKENPGK-- 360
Query 108 IGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCRQFV 157
G V+V+Y++ +K+A L GR + GK + A F P+ ++ Q V
Sbjct 361 -GQVFVEYANAGDSKEAQRLLTGRTFDGKFVVATFYPLGAYKRGYLYQTV 409
> mmu:170791 Rbm39, 1500012C14Rik, 2310040E03Rik, B330012G18Rik,
C79248, R75070, Rnpc2, caper; RNA binding motif protein 39;
K13091 RNA-binding protein 39
Length=530
Score = 40.0 bits (92), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 27/66 (40%), Positives = 37/66 (56%), Gaps = 3/66 (4%)
Query 83 EVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
+V EE K+G V + V N GNVYVK AA A++AL GR++AGK I A +
Sbjct 449 DVIEECNKHGGVIHIYVDKNSAQ---GNVYVKCPSIAAAIAAVNALHGRWFAGKMITAAY 505
Query 143 TPVTDF 148
P+ +
Sbjct 506 VPLPTY 511
Score = 32.7 bits (73), Expect = 0.55, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 35/68 (51%), Gaps = 7/68 (10%)
Query 76 HF---EAFFSEVFEELYKYGEVEDM-VVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGR 131
HF E +FE +G +E + ++ D+ G ++ +SD + AKKAL L G
Sbjct 258 HFNITEDMLRGIFE---PFGRIESIQLMMDSETGRSKGYGFITFSDSECAKKALEQLNGF 314
Query 132 YYAGKPIQ 139
AG+P++
Sbjct 315 ELAGRPMK 322
> hsa:9584 RBM39, CAPER, CAPERalpha, CC1.3, DKFZp781C0423, FLJ44170,
HCC1, RNPC2, fSAP59; RNA binding motif protein 39; K13091
RNA-binding protein 39
Length=524
Score = 40.0 bits (92), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 27/66 (40%), Positives = 37/66 (56%), Gaps = 3/66 (4%)
Query 83 EVFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
+V EE K+G V + V N GNVYVK AA A++AL GR++AGK I A +
Sbjct 443 DVIEECNKHGGVIHIYVDKNSAQ---GNVYVKCPSIAAAIAAVNALHGRWFAGKMITAAY 499
Query 143 TPVTDF 148
P+ +
Sbjct 500 VPLPTY 505
Score = 32.7 bits (73), Expect = 0.56, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 35/68 (51%), Gaps = 7/68 (10%)
Query 76 HF---EAFFSEVFEELYKYGEVEDM-VVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGR 131
HF E +FE +G +E + ++ D+ G ++ +SD + AKKAL L G
Sbjct 258 HFNITEDMLRGIFE---PFGRIESIQLMMDSETGRSKGYGFITFSDSECAKKALEQLNGF 314
Query 132 YYAGKPIQ 139
AG+P++
Sbjct 315 ELAGRPMK 322
> ath:AT5G16260 RNA recognition motif (RRM)-containing protein;
K13093 HIV Tat-specific factor 1
Length=519
Score = 40.0 bits (92), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 29/100 (29%), Positives = 49/100 (49%), Gaps = 18/100 (18%)
Query 43 TIVLRHMYPNPPVAIAIAEGQNVSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDN 102
T+VLR+M+ + + ++DE D +V EE K+G + + VC++
Sbjct 407 TVVLRYMF---------SPAELMADE------DLVAELEEDVKEESLKHGPFDSVKVCEH 451
Query 103 IGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEF 142
H G V V++ D A+K + A+ GR+YA + I A
Sbjct 452 ---HPQGVVLVRFKDRRDAQKCIEAMNGRWYAKRQIHASL 488
> dre:415211 puf60b, SIAHBP1, fe37c05, repressor, si:ch211-12p8.2,
si:zc12p8.2, wu:fb33e11, wu:fe37c05, zgc:86806; poly-U
binding splicing factor b
Length=502
Score = 39.7 bits (91), Expect = 0.004, Method: Composition-based stats.
Identities = 23/65 (35%), Positives = 37/65 (56%), Gaps = 5/65 (7%)
Query 83 EVFEELYKYGEVEDMVVC-DNIGDH----IIGNVYVKYSDDDAAKKALSALQGRYYAGKP 137
EV EE KYG V +++ + G+ II ++V++SD KA+ AL R++AG+
Sbjct 425 EVMEECGKYGAVNRVIIYQERQGEEDDAEIIVKIFVEFSDAGEMNKAIQALNNRWFAGRK 484
Query 138 IQAEF 142
+ AE
Sbjct 485 VVAEL 489
> pfa:PF14_0513 RNA binding protein, putative; K12840 splicing
factor 45
Length=511
Score = 39.7 bits (91), Expect = 0.005, Method: Composition-based stats.
Identities = 17/53 (32%), Positives = 31/53 (58%), Gaps = 0/53 (0%)
Query 96 DMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDF 148
++VV N+ D + +Y +Y D A+ AL+ +GR +AG+ +QA F ++
Sbjct 452 NIVVDKNLLDALAVKIYCEYESKDQAQNALNTFKGRTFAGRKVQASFATEEEY 504
> bbo:BBOV_II004960 18.m06413; RNA recognition motif (RRM)-containing
protein; K12837 splicing factor U2AF 65 kDa subunit
Length=383
Score = 39.3 bits (90), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 31/136 (22%), Positives = 63/136 (46%), Gaps = 22/136 (16%)
Query 37 KPSSSPTIVLRHMYPNPPVAIAIAEGQNVS---------------DELLDEAADHFEAFF 81
K ++ P V + + NP + + + G+ + ++L+ + H
Sbjct 246 KATNLPNSVTQSILSNPLLGLQMQSGRRIGSKPSRIVQLINIVFHEDLIQDKRYH--EVK 303
Query 82 SEVFEELYKYGEVEDMVV---CDNIG-DHIIGNVYVKYSDDDAAKKALSALQGRYYAGKP 137
+ EE KYG +ED+V+ D++ +G V++K+ D+ ++++A L GR + G
Sbjct 304 DAIMEEAKKYGHLEDIVIPRPNDDLSYKEGVGKVFLKFGDEISSRRAQYMLNGRVFDGNR 363
Query 138 IQ-AEFTPVTDFREAR 152
I A F P+ F + +
Sbjct 364 IVCAAFFPLDRFLKGK 379
> tgo:TGME49_034520 U2 snRNP auxiliary factor or splicing factor,
putative ; K12837 splicing factor U2AF 65 kDa subunit
Length=553
Score = 39.3 bits (90), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 32/134 (23%), Positives = 62/134 (46%), Gaps = 22/134 (16%)
Query 39 SSSPTIVLRHMYPNPPVAIAIAEGQNVSD---------------ELLDEAADHFEAFFSE 83
+S P + + + +P VA+ + + + + +L+D +EA +
Sbjct 418 TSLPNSMTQKLLSDPLVAVQVQAARKIGERPSKVVQLLNCVYQEDLID--PKEYEAICDD 475
Query 84 VFEELYKYGEVEDMVVCDNIGD----HIIGNVYVKYSDDDAAKKALSALQGRYY-AGKPI 138
+ +E K+G +E+++V D +G V+++YSD AA+KA L GR + + + +
Sbjct 476 IKQEAEKHGALEEVLVPKPNEDLSYREGVGKVFLRYSDVTAARKAQLMLNGRRFDSNRVV 535
Query 139 QAEFTPVTDFREAR 152
A F P F R
Sbjct 536 CAAFFPEEKFAAGR 549
> ath:AT1G60830 U2 snRNP auxiliary factor large subunit, putative
Length=111
Score = 38.9 bits (89), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 23/84 (27%), Positives = 45/84 (53%), Gaps = 6/84 (7%)
Query 65 VSDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDH----IIGNVYVKYSDDDA 120
+D+L D+A + ++ +E K+G + ++V+ DH +G V+++Y+D D
Sbjct 19 TADDLRDDA--EYADIMEDMSQEGGKFGNLVNVVIPRPNPDHDPTPGVGKVFLEYADVDG 76
Query 121 AKKALSALQGRYYAGKPIQAEFTP 144
+ KA S + GR + G + A + P
Sbjct 77 SSKARSGMNGRKFGGNQVVAVYYP 100
> sce:YGR159C NSR1, SHE5; Nsr1p; K11294 nucleolin
Length=414
Score = 38.9 bits (89), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 24/68 (35%), Positives = 39/68 (57%), Gaps = 1/68 (1%)
Query 77 FEAFFSEVFEELYKYGEVEDMVVCDNI-GDHIIGNVYVKYSDDDAAKKALSALQGRYYAG 135
F A +FE K+GEV + + + + G YV++S+ + AKKAL ALQG Y
Sbjct 276 FNADRDAIFELFAKHGEVVSVRIPTHPETEQPKGFGYVQFSNMEDAKKALDALQGEYIDN 335
Query 136 KPIQAEFT 143
+P++ +F+
Sbjct 336 RPVRLDFS 343
> ath:AT2G16940 RNA recognition motif (RRM)-containing protein
Length=561
Score = 38.5 bits (88), Expect = 0.010, Method: Compositional matrix adjust.
Identities = 26/72 (36%), Positives = 39/72 (54%), Gaps = 5/72 (6%)
Query 72 EAADHFEAFFSE-VFEELYKYGEVEDMVVCDNIGDHIIGNVYVKYSDDDAAKKALSALQG 130
E D F+ E V EE K+G++ + V N +G VY+++ + AA A AL G
Sbjct 478 ETEDDFDEDIKEDVKEECSKFGKLNHIFVDKNS----VGFVYLRFENAQAAIGAQRALHG 533
Query 131 RYYAGKPIQAEF 142
R++AGK I A +
Sbjct 534 RWFAGKMITATY 545
> ath:AT1G60900 U2 snRNP auxiliary factor large subunit, putative;
K12837 splicing factor U2AF 65 kDa subunit
Length=589
Score = 38.5 bits (88), Expect = 0.010, Method: Composition-based stats.
Identities = 22/83 (26%), Positives = 45/83 (54%), Gaps = 6/83 (7%)
Query 66 SDELLDEAADHFEAFFSEVFEELYKYGEVEDMVVCDNIGDHI----IGNVYVKYSDDDAA 121
+D+L D+ + + ++ +E K+G + ++V+ DH +G V+++Y+D D +
Sbjct 498 ADDLRDD--EEYAEIMEDMRQEGGKFGNLVNVVIPRPNPDHDPTPGVGKVFLEYADVDGS 555
Query 122 KKALSALQGRYYAGKPIQAEFTP 144
KA S + GR + G + A + P
Sbjct 556 SKARSGMNGRKFGGNQVVAVYYP 578
> cel:Y92C3B.2 uaf-1; U2AF splicing factor family member (uaf-1);
K12837 splicing factor U2AF 65 kDa subunit
Length=474
Score = 38.1 bits (87), Expect = 0.011, Method: Compositional matrix adjust.
Identities = 34/122 (27%), Positives = 57/122 (46%), Gaps = 19/122 (15%)
Query 51 PNPPVAIA---IAEGQNVSDELL--------DE--AADHFEAFFSEVFEELYKYGEVEDM 97
PN AIA +++G + E+L DE A D +E +V +E KYG V +
Sbjct 356 PNSASAIAGIDLSQGAGRATEILCLMNMVTEDELKADDEYEEILEDVRDECSKYGIVRSL 415
Query 98 VVCDNIGDHI---IGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTPVTDFREARCR 154
+ DH +G V+V+++ ++A +AL GR +A + + + V + R
Sbjct 416 EIPRPYEDHPVPGVGKVFVEFASTSDCQRAQAALTGRKFANRTVVTSYYDVDKYHN---R 472
Query 155 QF 156
QF
Sbjct 473 QF 474
> hsa:127933 UHMK1, DKFZp434C1613, FLJ23015, KIS, KIST; U2AF homology
motif (UHM) kinase 1 (EC:2.7.11.1); K08877 U2AF homology
motif (UHM) kinase 1 [EC:2.7.11.1]
Length=345
Score = 37.7 bits (86), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 24/71 (33%), Positives = 38/71 (53%), Gaps = 4/71 (5%)
Query 86 EELYKYGEVEDMVVC-DNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKPIQAEFTP 144
EE KYG V ++V +N G G V+V+Y++ +K A L GR + GK + A F P
Sbjct 275 EECQKYGPVVSLLVPKENPGR---GQVFVEYANAGDSKAAQKLLTGRMFDGKFVVATFYP 331
Query 145 VTDFREARCRQ 155
++ ++ Q
Sbjct 332 LSAYKRGYLYQ 342
> cel:F58B3.7 hypothetical protein; K12840 splicing factor 45
Length=371
Score = 37.4 bits (85), Expect = 0.022, Method: Compositional matrix adjust.
Identities = 20/71 (28%), Positives = 42/71 (59%), Gaps = 2/71 (2%)
Query 80 FFSEVFEELYKYGEVEDMVVC--DNIGDHIIGNVYVKYSDDDAAKKALSALQGRYYAGKP 137
F E+ EE+ K G+V +++V ++ + V+V+++++ A KA + GR++ G+
Sbjct 296 FADEIKEEMEKCGQVVNVIVHVDESQEEDRQVRVFVEFTNNAQAIKAFVMMNGRFFGGRS 355
Query 138 IQAEFTPVTDF 148
+ A F V+D+
Sbjct 356 VSAGFQNVSDY 366
> hsa:22827 PUF60, FIR, FLJ31379, RoBPI, SIAHBP1; poly-U binding
splicing factor 60KDa; K12838 poly(U)-binding-splicing factor
PUF60
Length=516
Score = 37.0 bits (84), Expect = 0.027, Method: Composition-based stats.
Identities = 22/64 (34%), Positives = 37/64 (57%), Gaps = 5/64 (7%)
Query 83 EVFEELYKYGEVEDMVVC-DNIGDH----IIGNVYVKYSDDDAAKKALSALQGRYYAGKP 137
EV EE K+G V +++ + G+ II ++V++S KA+ AL GR++AG+
Sbjct 439 EVTEECGKFGAVNRVIIYQEKQGEEEDAEIIVKIFVEFSIASETHKAIQALNGRWFAGRK 498
Query 138 IQAE 141
+ AE
Sbjct 499 VVAE 502
Lambda K H
0.320 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3712313100
Database: egene_temp_file_orthology_annotation_similarity_blast_database_865
Posted date: Sep 17, 2011 11:19 AM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40