bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
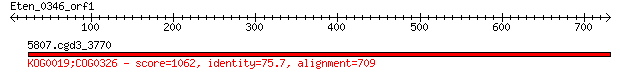
Query= Eten_0346_orf1
Length=731
Score E
Sequences producing significant alignments: (Bits) Value
5807.cgd3_3770 1062 0.0
> 5807.cgd3_3770
Length=711
Score = 1062 bits (2747), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 537/709 (75%), Positives = 616/709 (86%), Gaps = 12/709 (1%)
Query 23 ETFAFNADIQQLMSLIINTFYSNKEIFLRELISNASDALDKIRYEAITEPEKLKTKPELF 82
ETFAFNADIQQLMSLIINTFYSNKEIFLRELISNASDALDKIRYE++T+PE+LK+ E+
Sbjct 15 ETFAFNADIQQLMSLIINTFYSNKEIFLRELISNASDALDKIRYESLTDPEQLKSNEEMH 74
Query 83 IRLIPDKANNTLTIEDSGIGMTKAELVNNLGTIARSGTKAFMEALQAGGDISMIGQFGVG 142
IR+IPDK NNTLTIEDSGIGMTK EL+NNLGTIARSGTKAFMEA+QAGGD+SMIGQFGVG
Sbjct 75 IRIIPDKVNNTLTIEDSGIGMTKNELINNLGTIARSGTKAFMEAIQAGGDVSMIGQFGVG 134
Query 143 FYSAYLVADSVTVVSKHNDDEQYVWESAAGGSFTVQKDDKYEPLGRGTRIILHLKEDQGE 202
FYSAYLVAD VTV++KHN DEQY+WES+AGGSFT+ D L RGTRIILHLKEDQ +
Sbjct 135 FYSAYLVADKVTVITKHNGDEQYIWESSAGGSFTITNDTSDNKLQRGTRIILHLKEDQLD 194
Query 203 YLEERRLKDLVKKHSEFISFPIELAVEKTHEREVTESEDEEEKKADEKAEEKEGEEKKEG 262
YLEER L+DLVKKHSEFISFPIEL+VEKT E+E+T+S+ +E+E +++ E
Sbjct 195 YLEERTLRDLVKKHSEFISFPIELSVEKTTEKEITDSD----------VDEEEEKKEGED 244
Query 263 EEKKEGEEEKKEKTKKTKKVKEVTREWEQLNKQKPLWMRKPEEVTEEEYASFYKSLSNDW 322
E EE KEK K KK+ EVT+ W+ LNK KP+WMRKPEEVT EEY+SFYKS+SNDW
Sbjct 245 GEDAPKIEEVKEKEPKKKKITEVTQSWDLLNKNKPIWMRKPEEVTFEEYSSFYKSISNDW 304
Query 323 EEHLAVKHFSVEGQLEFKALLFVPKRAPFDLFETRKKRNNIKLYVRRVFIMDDCEDIIPE 382
E+ LAVKHFSVEGQLEFKA+LF+P+RAPFDLFETRKKRNNIKLYVRRVFIMDDCE++IPE
Sbjct 305 EDPLAVKHFSVEGQLEFKAILFIPRRAPFDLFETRKKRNNIKLYVRRVFIMDDCEELIPE 364
Query 383 WLNFVKGVVDSEDLPLNISRESLQQNKILKVIRKNLVKKCLEMFAEIEEKKENYAKFYEQ 442
+L FV+GVVDSEDLPLNISRESLQQNKILKVI+KN+VKKCLE+ EI EK ++Y KFYEQ
Sbjct 365 FLGFVRGVVDSEDLPLNISRESLQQNKILKVIKKNIVKKCLELITEITEKPDDYKKFYEQ 424
Query 443 FSKNLKLGIHEDSANRAKIAELLRFHSSKSGEDMVSFKEYVDRMKEGQKDIYYITGESRQ 502
FSKNLKLGIHED+ NR KI+ELLR+ +SKSGE+++S +EYVDRMKE QK+IYYITGES Q
Sbjct 425 FSKNLKLGIHEDTTNRNKISELLRYQTSKSGEELISLREYVDRMKENQKEIYYITGESIQ 484
Query 503 TVANSPFLEKLTKKGYEVLYMTDPIDEYAVQQLKEFDNHKLRCCTKEGLEIDESEEEKKK 562
V NSPFLEKL K YEV+YM DPIDEY VQQ+KEFD KLRCCTKEGL ++E+ EEK+
Sbjct 485 AVQNSPFLEKLRKLDYEVIYMVDPIDEYCVQQMKEFDGKKLRCCTKEGLTLEETAEEKEA 544
Query 563 FEELKAEFEPLLKLIKEVLHDKVDKVVLSNRITDSPCVLVTTEFGWSANMERIMKAQALR 622
FE L+ E+EPL +LIKEVLHDKVDKV+ S RI+DSPCVLVT+EFGWSANMERIMKAQALR
Sbjct 545 FEALQKEYEPLCQLIKEVLHDKVDKVITSQRISDSPCVLVTSEFGWSANMERIMKAQALR 604
Query 623 DNSMTSYMVSKKTMEVNGHHSIMIEIKNKAAVDKSDKTVKDLIWLLYDTALLTSGFSLEE 682
D SMTSYM+SKKTME+N ++SI+ E+K K A DKSDKTVKDLIWLLYDT+LLTSGFSLE+
Sbjct 605 DTSMTSYMMSKKTMEINPYNSIITELKTKIANDKSDKTVKDLIWLLYDTSLLTSGFSLED 664
Query 683 PTQFAARIHRMIKLGLSIDDDEEAKDDDLPPLEEVEGAADEASKMEEVD 731
PTQF++RI+RMIKLGLSI DEE DDLPPLE V A +ASKMEEVD
Sbjct 665 PTQFSSRINRMIKLGLSI--DEEDIVDDLPPLEPVNDAELQASKMEEVD 711
Lambda K H
0.313 0.131 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 343748210575
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40