bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eten_0412_orf6
Length=518
Score E
Sequences producing significant alignments: (Bits) Value
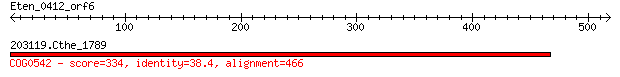
203119.Cthe_1789 334 4e-90
> 203119.Cthe_1789
Length=818
Score = 334 bits (857), Expect = 4e-90, Method: Compositional matrix adjust.
Identities = 179/483 (37%), Positives = 290/483 (60%), Gaps = 46/483 (9%)
Query 1 TREKALSILQRVRPHYEQFHQLEIPEEVLEAVTNMSDQYVKQRAFPDKALDLLDESCSWK 60
T+E+A+ IL+ +R YE H ++I +E L A NMSD+Y+ R PDKA+DL+DE+ S
Sbjct 344 TKEEAIEILRGLRDKYEAHHSVKITDEALVAAVNMSDRYITDRFLPDKAIDLIDEAASRV 403
Query 61 RV---SHNKKINVLNKQIAQLKKNK----LPEEEEELKKLQEELA----ALEALTAGGRR 109
R+ + + L +++ +L+K K + +E E+ ++++E LE R+
Sbjct 404 RLKSFTAPPDLKHLEEKVERLRKEKEDAIVCQEFEKAARIRDEEQRLKNELEKAKDSWRQ 463
Query 110 L------VLETNDVAHILSQWTGIPMGKMTEDEVSRVLRLADILSLRVVGQDRAVKAVAD 163
V+ +D+A I+S WTGIP+ ++ E+E R++++ DIL RV+GQD AVKA++
Sbjct 464 KNQTTTNVVSEDDIAVIVSDWTGIPVKRLAEEESERLMKMEDILHKRVIGQDEAVKAISK 523
Query 164 ALAIQRAGLSPKNKPLGTFLFLGSSGVGKTELAKAVAEEMFDSEKNLIRLDMVEYQEAHS 223
A+ R GL +P+G+F+FLG +GVGKTEL+KA+AE +F E +IR+DM EY E HS
Sbjct 524 AIRRGRVGLKDPKRPVGSFIFLGPTGVGKTELSKALAEALFGEENAMIRIDMSEYMEKHS 583
Query 224 ISRLIGPPPGYMGNDEGGQLTEAVRQKPHSVVLFDEVENAHKNLWSLLLPMLDEGHLSDS 283
+SRL+G PPGY+G +EGGQLTE VR+KP+SVVLFDE+E AH +++++LL +L++G L+DS
Sbjct 584 VSRLVGSPPGYVGYEEGGQLTEKVRRKPYSVVLFDEIEKAHPDIFNILLQILEDGRLTDS 643
Query 284 KGNRVDFTNCLIILTSNIGQQYILDSYKEVRALTAGNEKDKASSNGSSSSNGEEPGKSSF 343
+G VDF N +II+TSNIG + I EP + F
Sbjct 644 QGRVVDFRNTVIIMTSNIGARLIT-----------------------------EPKQLGF 674
Query 344 KGTGKSPKDILRRMRQKVLKEVFGYFKPQVIGRMSEIIIFEPLGETAMKGVLNLKLSALR 403
+ K M+ V+ E+ F+P+ + R+ EII+F PL E +K ++ L + L
Sbjct 675 APVAQDKKKSYEDMKNNVMNELKKNFRPEFLNRIDEIIVFHPLEEEHLKQIVGLMIDNLA 734
Query 404 NSLAAKGIDFKVADSALGYILERAWSHKFGGRRMAKYLEKYIKPRVAPLLISGKLKAGDR 463
L I+ +V+D A + ++ + +G R + + ++ ++ R+A ++ G++K+GD+
Sbjct 735 ERLKQNSIEIEVSDEAKALLAKKGFDPVYGARPLRRAVQSMVEDRLAEEMLEGRVKSGDK 794
Query 464 AVL 466
+
Sbjct 795 VFV 797
Lambda K H
0.314 0.132 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 225293496505
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40