bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_0160_orf4
Length=246
Score E
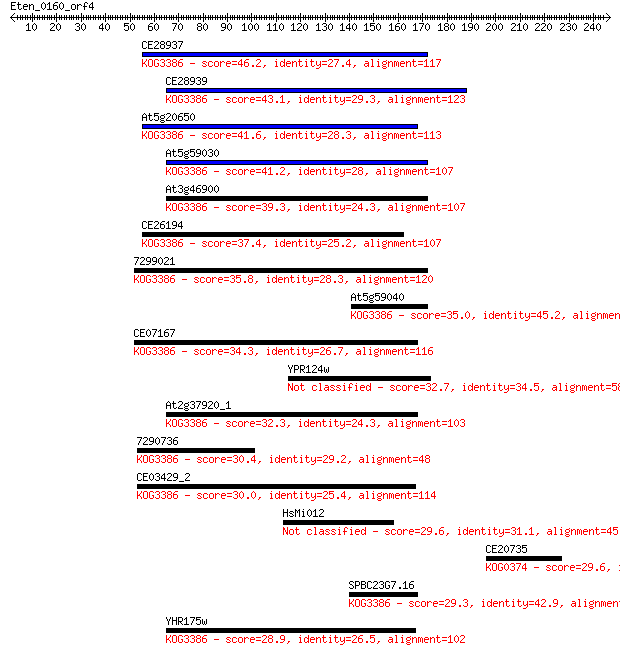
Sequences producing significant alignments: (Bits) Value
CE28937 46.2 6e-05
CE28939 43.1 5e-04
At5g20650 41.6 0.001
At5g59030 41.2 0.002
At3g46900 39.3 0.008
CE26194 37.4 0.026
7299021 35.8 0.073
At5g59040 35.0 0.13
CE07167 34.3 0.27
YPR124w 32.7 0.72
At2g37920_1 32.3 0.86
7290736 30.4 3.6
CE03429_2 30.0 4.5
HsMi012 29.6 5.5
CE20735 29.6 6.0
SPBC23G7.16 29.3 8.1
YHR175w 28.9 9.0
> CE28937
Length=134
Score = 46.2 bits (108), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 32/117 (27%), Positives = 57/117 (48%), Gaps = 4/117 (3%)
Query 55 MSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENRSK 114
M F P+ +LFS W+ + + C+ I G I A+K RR+ R + + S
Sbjct 18 MWFHTKPQDTVLFSTWNITSAGKMVWACILVAIAGIILEAIKYNRRLIQKRQSPSKKESY 77
Query 115 PTLLFGCFPVFHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFVSMLGGMALGFLTIG 171
+ L F + F+ + Y LML+ MTF++ + ++++ G+++GFL G
Sbjct 78 ISRLLSTMHFFQT----FLFFVQLGFSYCLMLIFMTFSIWLGLAVVIGLSIGFLIFG 130
> CE28939
Length=166
Score = 43.1 bits (100), Expect = 5e-04, Method: Compositional matrix adjust.
Identities = 36/150 (24%), Positives = 62/150 (41%), Gaps = 30/150 (20%)
Query 65 LLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENR------------ 112
+LF W Y +C+ + F ALK R ++ ++E +
Sbjct 18 ILFREWKPLNTTAYVFSCIEIFLIAFCLEALKFGRTKLSPKVKIVEKKVDCCCSTEKDGL 77
Query 113 ---------SKPTLLFGCFP------VFHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFV 157
++ T+ F FH A + F+ + DY LMLV+MT+N IF+
Sbjct 78 WNIPETIPLTQKTVTLAPFTRDSLISKFHMA-SSLLVFVQHFIDYSLMLVSMTYNWPIFL 136
Query 158 SMLGGMALGFLTIGRFLDLPFEPAKVSGGC 187
S+L G G+ +G + + E ++ +G C
Sbjct 137 SLLAGHTTGYFFLGPMMTV--EESEAAGSC 164
> At5g20650
Length=146
Score = 41.6 bits (96), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 32/122 (26%), Positives = 55/122 (45%), Gaps = 9/122 (7%)
Query 55 MSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLR----RISDVRLAMME 110
M+F + +LF +W + Y +T + C +F L+ R +S R A
Sbjct 4 MTFYWGIKATILFDFWKTDSWLSYILTLIACFVFSAFYQYLENRRIQFKSLSSSRRAPPP 63
Query 111 NRSKPTLLFGCFPV--FHNAIRGCVTFL---NYSWDYMLMLVAMTFNVGIFVSMLGGMAL 165
RS + P +A + L N + Y+LML AM+FN G+F++++ G+
Sbjct 64 PRSSSGVSAPLIPKSGTRSAAKAASVLLFGVNAAIGYLLMLAAMSFNGGVFIAIVVGLTA 123
Query 166 GF 167
G+
Sbjct 124 GY 125
> At5g59030
Length=170
Score = 41.2 bits (95), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 30/107 (28%), Positives = 52/107 (48%), Gaps = 13/107 (12%)
Query 65 LLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENRSKPTLLFGCFPV 124
+LFS W ++ YA+ C I F+++ + L S +R + ++ ++ L
Sbjct 54 VLFSGWPGTSSGMYAL---CLIFVFFLAVLTEWLAHSSLLRGSTGDSANRAAGL------ 104
Query 125 FHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFVSMLGGMALGFLTIG 171
I+ V L Y++ML M+FN G+F+ L G A+GF+ G
Sbjct 105 ----IQTAVYTLRIGLAYLVMLAVMSFNAGVFLVALAGHAVGFMLFG 147
> At3g46900
Length=158
Score = 39.3 bits (90), Expect = 0.008, Method: Compositional matrix adjust.
Identities = 26/107 (24%), Positives = 47/107 (43%), Gaps = 15/107 (14%)
Query 65 LLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENRSKPTLLFGCFPV 124
+LFS W ++ YA+ C I+ +++ + L +R++ NR+
Sbjct 42 VLFSGWPGTSSGMYAL---CLIVIFLLAVIAEWLAHSPILRVSGSTNRAA---------- 88
Query 125 FHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFVSMLGGMALGFLTIG 171
+ V L Y++ML M+FN G+F+ + G +GF G
Sbjct 89 --GLAQTAVYTLKTGLSYLVMLAVMSFNAGVFIVAIAGYGVGFFLFG 133
> CE26194
Length=130
Score = 37.4 bits (85), Expect = 0.026, Method: Compositional matrix adjust.
Identities = 27/108 (25%), Positives = 44/108 (40%), Gaps = 2/108 (1%)
Query 55 MSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENRSK 114
MSF E +LF +W T AV C ++ F+ L+ R + + +
Sbjct 1 MSFHFGTEETILFDFWKTETAVGIAVACFITVLLAFLMETLRFFRDYRKAQTQLHQPPIS 60
Query 115 PTLLFGCFPVFHNAIRGCVTFLNYSWDYMLMLVAMTFNVGI-FVSMLG 161
P P + I + + Y LML+ MTFN + F +++G
Sbjct 61 PEDRLKRSPQL-DLIDPLLQLFQLTIAYFLMLIFMTFNAYLCFFTVVG 107
> 7299021
Length=174
Score = 35.8 bits (81), Expect = 0.073, Method: Compositional matrix adjust.
Identities = 34/147 (23%), Positives = 60/147 (40%), Gaps = 28/147 (19%)
Query 52 PLPMSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLR---------RIS 102
P+ M F A +L+ W A T ++ ++ + + F+ ALK LR R S
Sbjct 16 PMIMVFHAGHCERILWRGWVASTVTEFVLSALAIFLVSFLYEALKFLRQQLARREARRAS 75
Query 103 DVRLAMMENRSKPTLLFGC------------------FPVFHNAIRGCVTFLNYSWDYML 144
+ A +++ GC F H ++ + L Y+L
Sbjct 76 EQLAAEQRRKNEAPAAGGCCSEAPLAEPREQTYWQRLFASSH-IVQSLLNLLQIVISYLL 134
Query 145 MLVAMTFNVGIFVSMLGGMALGFLTIG 171
ML+ MTFN + ++++ G+ LG+ G
Sbjct 135 MLIFMTFNYWLCLAVILGLGLGYFFFG 161
> At5g59040
Length=151
Score = 35.0 bits (79), Expect = 0.13, Method: Compositional matrix adjust.
Identities = 14/31 (45%), Positives = 21/31 (67%), Gaps = 0/31 (0%)
Query 141 DYMLMLVAMTFNVGIFVSMLGGMALGFLTIG 171
Y++ML M+FN G+FV+ + G LGF+ G
Sbjct 102 SYLVMLAVMSFNGGVFVAAMAGFGLGFMIFG 132
> CE07167
Length=162
Score = 34.3 bits (77), Expect = 0.27, Method: Compositional matrix adjust.
Identities = 31/148 (20%), Positives = 62/148 (41%), Gaps = 36/148 (24%)
Query 52 PLPMSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLR------------ 99
+ M+ +LFSWW + + AV+ + + + A+K R
Sbjct 2 DMDMTLHFGEREKILFSWWKTGSLSGMAVSMLITFLLCILYEAIKSFRYFLAVWNNQKRQ 61
Query 100 ------RISDVRLAMMENRSKPTL--------------LFGCFPVFHNAIRGCVTFLNYS 139
I++ + + +N S+ ++ LF + + A+ G L Y+
Sbjct 62 QRHAEASITNPQNSGGDNISEDSIHIAPLVQLSGFTKRLFTSYRLAQGALYGLQALLAYT 121
Query 140 WDYMLMLVAMTFNVGIFVSMLGGMALGF 167
LML+AMT+N+ + +S++ G A+G+
Sbjct 122 ----LMLIAMTYNMNLILSIVVGEAVGY 145
> YPR124w
Length=406
Score = 32.7 bits (73), Expect = 0.72, Method: Compositional matrix adjust.
Identities = 20/63 (31%), Positives = 31/63 (49%), Gaps = 5/63 (7%)
Query 115 PTLLFGCF-----PVFHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFVSMLGGMALGFLT 169
P LL F +FH+ IR + F + YMLML M+F + +++ G+AL +
Sbjct 221 PNLLSDIFVPSLMDLFHDIIRAFLVFTSTMIIYMLMLATMSFVLTYVFAVITGLALSEVF 280
Query 170 IGR 172
R
Sbjct 281 FNR 283
> At2g37920_1
Length=185
Score = 32.3 bits (72), Expect = 0.86, Method: Compositional matrix adjust.
Identities = 25/103 (24%), Positives = 46/103 (44%), Gaps = 13/103 (12%)
Query 65 LLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENRSKPTLLFGCFPV 124
+LFS W YA+ + F++ + L R SD ++ G +
Sbjct 42 VLFSGWPGSDRGMYALALIFVFFLAFLA---EWLARCSDAS----------SIKQGADKL 88
Query 125 FHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFVSMLGGMALGF 167
A R + + + Y+++L ++FN G+F++ + G ALGF
Sbjct 89 AKVAFRTAMYTVKSGFSYLVILAVVSFNGGVFLAAIFGHALGF 131
> 7290736
Length=220
Score = 30.4 bits (67), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 14/48 (29%), Positives = 21/48 (43%), Gaps = 0/48 (0%)
Query 53 LPMSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRR 100
+PM+F +LFSWWH T A + + + + LK R
Sbjct 58 MPMAFHFGYNETILFSWWHIETVAGLIGSMIAIFLLALMYEGLKYYRE 105
> CE03429_2
Length=156
Score = 30.0 bits (66), Expect = 4.5, Method: Compositional matrix adjust.
Identities = 29/124 (23%), Positives = 50/124 (40%), Gaps = 10/124 (8%)
Query 53 LPMSFEASPEVLLLFSWWHARTNAQYAVTCVCCIIFGFISIALKGLRRISDVRLAMMENR 112
+ M + E +LF W TC G + ALK R ++ R+ + +
Sbjct 21 MWMWYHVDVEDTVLFKSWTVFDAGTMVWTCFVVAAAGILLEALKYARWATEERMKIDQEN 80
Query 113 SKPTLLFGCFPV------FHNAIRGCVTFLNYSWD----YMLMLVAMTFNVGIFVSMLGG 162
+G + ++ R + L + W Y+LM V M F+V I +S+ G
Sbjct 81 VDSKTKYGGIKIPGKSEKYNFWKRHIIDSLYHFWQLLLAYILMNVYMVFSVYICLSLCFG 140
Query 163 MALG 166
+A+G
Sbjct 141 LAIG 144
> HsMi012
Length=174
Score = 29.6 bits (65), Expect = 5.5, Method: Compositional matrix adjust.
Identities = 14/45 (31%), Positives = 26/45 (57%), Gaps = 0/45 (0%)
Query 113 SKPTLLFGCFPVFHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFV 157
SKP+ ++G + + + GCV LN+ YM ++V + + G+ V
Sbjct 21 SKPSPIYGGLVLIVSGVVGCVIILNFGGGYMGLMVFLIYLGGMMV 65
> CE20735
Length=329
Score = 29.6 bits (65), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 11/31 (35%), Positives = 19/31 (61%), Gaps = 0/31 (0%)
Query 196 GCHKGRPCTCAASQLVASCRRSKMSFGEQPV 226
GC G+P T + +++ A C +S+ F QP+
Sbjct 21 GCRPGKPVTMSEAEIRALCHKSREIFLSQPI 51
> SPBC23G7.16
Length=148
Score = 29.3 bits (64), Expect = 8.1, Method: Compositional matrix adjust.
Identities = 12/28 (42%), Positives = 19/28 (67%), Gaps = 0/28 (0%)
Query 140 WDYMLMLVAMTFNVGIFVSMLGGMALGF 167
+ Y LMLVAMT+N + +++ G A G+
Sbjct 106 FSYFLMLVAMTYNAYVILAIAIGAAFGY 133
> YHR175w
Length=189
Score = 28.9 bits (63), Expect = 9.0, Method: Compositional matrix adjust.
Identities = 27/105 (25%), Positives = 52/105 (49%), Gaps = 10/105 (9%)
Query 65 LLFSWWHARTNAQYAVTCVCCIIFGFISIALK---GLRRISDVRLAMMENRSKPTLLFGC 121
++F WWH +T ++C+ ++ LK R++S + ++ NRS T +
Sbjct 72 VVFEWWHIKTLPGLILSCLAIFGLAYLYEYLKYCVHKRQLS--QRVLLPNRS-LTKINQA 128
Query 122 FPVFHNAIRGCVTFLNYSWDYMLMLVAMTFNVGIFVSMLGGMALG 166
V ++ + G L + +MLMLV MT+N + ++++ G G
Sbjct 129 DKVSNSILYG----LQVGFSFMLMLVFMTYNGWLMLAVVCGAIWG 169
Lambda K H
0.328 0.140 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4974335240
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40