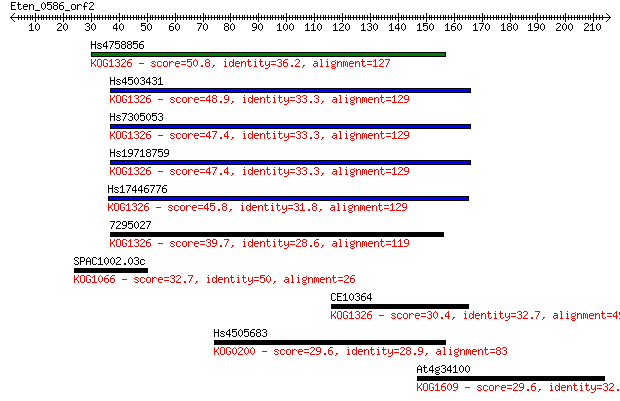
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_0586_orf2
Length=215
Score E
Sequences producing significant alignments: (Bits) Value
Hs4758856 50.8 2e-06
Hs4503431 48.9 8e-06
Hs7305053 47.4 2e-05
Hs19718759 47.4 2e-05
Hs17446776 45.8 7e-05
7295027 39.7 0.005
SPAC1002.03c 32.7 0.59
CE10364 30.4 2.5
Hs4505683 29.6 4.7
At4g34100 29.6 5.0
> Hs4758856
Length=1230
Score = 50.8 bits (120), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 46/167 (27%), Positives = 68/167 (40%), Gaps = 41/167 (24%)
Query 30 QGPQPLLQSTDVHYYSKTGTAIFNWRVV--YSRVAAPVNSCI------------------ 69
+G Q Q TDVHY+S TG FNWR + + +AA I
Sbjct 1000 KGQQEDKQDTDVHYHSLTGEGNFNWRYLFPFDYLAAEEKIVISKKESMFSWDETEYKIPA 1059
Query 70 -LQISAYDFRNIGESPFIGEVNLEVRKYLERVAQT-------MASIEVDAELR---LQNR 118
L + +D + F+G + L++ ++ R A+T MA+ EVD L Q R
Sbjct 1060 RLTLQIWDADHFSADDFLGAIELDLNRF-PRGAKTAKQCTMEMATGEVDVPLVSIFKQKR 1118
Query 119 AQ---------ETVDVQTFGYVQVTLQFMAQSEATSRPVGLGREPPN 156
+ E + + G V+ L + EA PVGL R P+
Sbjct 1119 VKGWWPLLARNENDEFELTGKVEAELHLLTAEEAEKNPVGLARNEPD 1165
> Hs4503431
Length=2080
Score = 48.9 bits (115), Expect = 8e-06, Method: Compositional matrix adjust.
Identities = 43/169 (25%), Positives = 68/169 (40%), Gaps = 41/169 (24%)
Query 37 QSTDVHYYSKTGTAIFNWRVVYSRVAAPVNS-CIL--------------QISA------Y 75
Q TDVHY S G FNWR ++ P C + +I A +
Sbjct 1852 QKTDVHYRSLGGEGNFNWRFIFPFDYLPAEQVCTIAKKDAFWRLDKTESKIPARVVFQIW 1911
Query 76 DFRNIGESPFIGEVNLEVRKYLERVAQTMASIEVD-----------AELRLQNR------ 118
D F+G + L++ + + + A+T +D L Q
Sbjct 1912 DNDKFSFDDFLGSLQLDLNR-MPKPAKTAKKCSLDQLDDAFHPEWFVSLFEQKTVKGWWP 1970
Query 119 --AQETVDVQTFGYVQVTLQFMAQSEATSRPVGLGREPPNRDPRLTTPQ 165
A+E G +++TL+ +A+SE RP G GR+ PN +P+L P+
Sbjct 1971 CVAEEGEKKILAGKLEMTLEIVAESEHEERPAGQGRDEPNMNPKLEDPR 2019
> Hs7305053
Length=2061
Score = 47.4 bits (111), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 43/173 (24%), Positives = 66/173 (38%), Gaps = 45/173 (26%)
Query 37 QSTDVHYYSKTGTAIFNWRVVYSRVAAPVNS-CI--------------------LQISAY 75
Q TDVHY S G FNWR V+ P CI L I +
Sbjct 1829 QKTDVHYRSLDGEGNFNWRFVFPFDYLPAEQLCIVAKKEHFWSIDQTEFRIPPRLIIQIW 1888
Query 76 DFRNIGESPFIGEVNLEVR------KYLERVAQTMASIEVDAELRLQNRAQETVDVQTF- 128
D ++G + L++R K E+ M ++ A L+ + + ++
Sbjct 1889 DNDKFSLDDYLGFLELDLRHTIIPAKSPEKCRLDMIP-DLKAMNPLKAKTASLFEQKSMK 1947
Query 129 ----------------GYVQVTLQFMAQSEATSRPVGLGREPPNRDPRLTTPQ 165
G V++TL+ + + EA RP G GR+ PN +P+L P
Sbjct 1948 GWWPCYAEKDGARVMAGKVEMTLEILNEKEADERPAGKGRDEPNMNPKLDLPN 2000
> Hs19718759
Length=2048
Score = 47.4 bits (111), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 43/173 (24%), Positives = 66/173 (38%), Gaps = 45/173 (26%)
Query 37 QSTDVHYYSKTGTAIFNWRVVYSRVAAPVNS-CI--------------------LQISAY 75
Q TDVHY S G FNWR V+ P CI L I +
Sbjct 1816 QKTDVHYRSLDGEGNFNWRFVFPFDYLPAEQLCIVAKKEHFWSIDQTEFRIPPRLIIQIW 1875
Query 76 DFRNIGESPFIGEVNLEVR------KYLERVAQTMASIEVDAELRLQNRAQETVDVQTF- 128
D ++G + L++R K E+ M ++ A L+ + + ++
Sbjct 1876 DNDKFSLDDYLGFLELDLRHTIIPAKSPEKCRLDMIP-DLKAMNPLKAKTASLFEQKSMK 1934
Query 129 ----------------GYVQVTLQFMAQSEATSRPVGLGREPPNRDPRLTTPQ 165
G V++TL+ + + EA RP G GR+ PN +P+L P
Sbjct 1935 GWWPCYAEKDGARVMAGKVEMTLEILNEKEADERPAGKGRDEPNMNPKLDLPN 1987
> Hs17446776
Length=802
Score = 45.8 bits (107), Expect = 7e-05, Method: Compositional matrix adjust.
Identities = 41/173 (23%), Positives = 71/173 (41%), Gaps = 44/173 (25%)
Query 36 LQSTDVHYYSKTGTAIFNWRVVYS-RVAAPVNSCI--------------------LQISA 74
+Q TD+HY+S TG A FNWR +++ A +C+ L I
Sbjct 506 MQKTDIHYHSLTGEADFNWRFIFTMDYLAAERTCVQSQKDYIWSLDATSMKFPARLIIQV 565
Query 75 YDFRNIGESPFIG--EVNLEVRKYLERVAQTMASIEVDAELR----LQNRAQETVDVQTF 128
+D F+G E++L R A+ + +DA+ + +Q + +T
Sbjct 566 WDNDIFSPDDFLGVLELDLSDMPLPARHAKQCSIRMMDADPKWPYFIQYKHFSLFKKKTV 625
Query 129 -----------------GYVQVTLQFMAQSEATSRPVGLGREPPNRDPRLTTP 164
G V+++L+ +++ EA +P G G+ PN+ P L P
Sbjct 626 TGWWPCQVLDGGKWRLSGKVKMSLEILSEKEALIKPAGRGQSEPNQYPTLHPP 678
> 7295027
Length=1782
Score = 39.7 bits (91), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 34/157 (21%), Positives = 62/157 (39%), Gaps = 39/157 (24%)
Query 37 QSTDVHYYSKTGTAIFNWRVVYSRVAAPVNSC-------------------ILQISAYDF 77
QSTD+HY S G A FNWR+V+ +P I+ + +D
Sbjct 1403 QSTDIHYRSFAGDAAFNWRMVFGLKYSPNEDLMVIRRKAGLLEEIEFKKPPIIYLQVWDK 1462
Query 78 RNIGESPFIGEVNLEVR----------------KYLERVAQTMASIEVDAELRLQNRAQE 121
+ + ++ + L + K +R+ + ++ LQ+ A
Sbjct 1463 DVLTKDEYLAALELNLSNMYAPYPAAQLCKPYPKKRKRINLFKRKV-INGWFPLQSNAHP 1521
Query 122 TVDVQTF---GYVQVTLQFMAQSEATSRPVGLGREPP 155
+ Q G +Q++++ EA +RP G G++PP
Sbjct 1522 SQRSQAAVQNGKMQLSIEVCTDVEAVNRPAGRGQDPP 1558
> SPAC1002.03c
Length=923
Score = 32.7 bits (73), Expect = 0.59, Method: Compositional matrix adjust.
Identities = 13/26 (50%), Positives = 17/26 (65%), Gaps = 0/26 (0%)
Query 24 MDCTKYQGPQPLLQSTDVHYYSKTGT 49
+D K GP P QST H+YS++GT
Sbjct 318 IDVEKESGPSPHSQSTSTHWYSESGT 343
> CE10364
Length=2034
Score = 30.4 bits (67), Expect = 2.5, Method: Compositional matrix adjust.
Identities = 16/54 (29%), Positives = 23/54 (42%), Gaps = 5/54 (9%)
Query 116 QNRAQETVDVQT-----FGYVQVTLQFMAQSEATSRPVGLGREPPNRDPRLTTP 164
QN ++ DV G V++ + + + EA P G R PN P L P
Sbjct 1919 QNAKKKNDDVGKDPKWIMGLVEMDMLLLTKQEADQEPAGKKRSEPNHSPFLEKP 1972
> Hs4505683
Length=1106
Score = 29.6 bits (65), Expect = 4.7, Method: Compositional matrix adjust.
Identities = 24/83 (28%), Positives = 37/83 (44%), Gaps = 3/83 (3%)
Query 74 AYDFRNIGESPFIGEVNLEVRKYLERVAQTMASIEVDAELRLQNRAQETVDVQTFGYVQV 133
A RN+ E+ ++ E+ L K E TM + DAE++L + Q V V+ ++
Sbjct 366 ALSTRNVSETRYVSELTLVRVKVAEAGHYTMRAFHEDAEVQLSFQLQINVPVRVL---EL 422
Query 134 TLQFMAQSEATSRPVGLGREPPN 156
+ E T R G G PN
Sbjct 423 SESHPDSGEQTVRCRGRGMPQPN 445
> At4g34100
Length=1051
Score = 29.6 bits (65), Expect = 5.0, Method: Compositional matrix adjust.
Identities = 22/69 (31%), Positives = 31/69 (44%), Gaps = 2/69 (2%)
Query 147 PVGLGREPPNRDPRLTTPQEGRKWEDVLGSAGLRVDYRPIWYWVRVVF--IVFLSLWLFV 204
PV LGR + P L + + + S V IW W +VF V L++W+F+
Sbjct 783 PVSLGRALFSAIPILPITHGIKYAIEHVKSKRTSVLLNQIWKWCGIVFKSSVLLAIWVFI 842
Query 205 IAFLYPALF 213
I L LF
Sbjct 843 IPVLIGLLF 851
Lambda K H
0.324 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3969005466
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40