bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_0756_orf1
Length=106
Score E
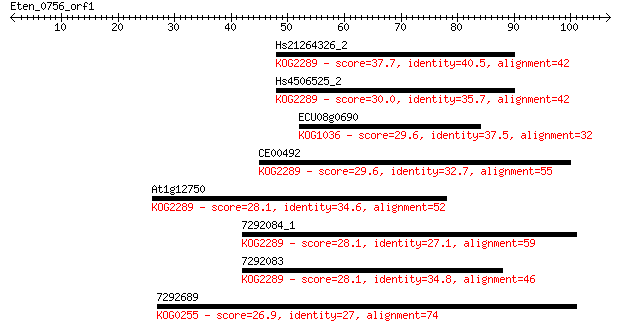
Sequences producing significant alignments: (Bits) Value
Hs21264326_2 37.7 0.006
Hs4506525_2 30.0 1.1
ECU08g0690 29.6 1.3
CE00492 29.6 1.4
At1g12750 28.1 3.8
7292084_1 28.1 3.8
7292083 28.1 4.4
7292689 26.9 8.1
> Hs21264326_2
Length=302
Score = 37.7 bits (86), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 17/46 (36%), Positives = 28/46 (60%), Gaps = 4/46 (8%)
Query 48 MGHLGGLICGLSLGFL----YNKDMQDKPRWWSFAMWCSVFLLVAL 89
+ HLGG+ G++LG + Y + +QD+ WW F +VF+L A+
Sbjct 239 VAHLGGVAVGITLGVVVLRNYEQRLQDQSLWWIFVAMYTVFVLFAV 284
> Hs4506525_2
Length=302
Score = 30.0 bits (66), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 15/46 (32%), Positives = 24/46 (52%), Gaps = 4/46 (8%)
Query 48 MGHLGGLICGLSLGFL----YNKDMQDKPRWWSFAMWCSVFLLVAL 89
M HL G + G+S+G Y + ++D+ WW + FLL A+
Sbjct 239 MAHLAGAVVGVSMGLTILRSYEERLRDQCGWWVVLLAYGTFLLFAV 284
> ECU08g0690
Length=287
Score = 29.6 bits (65), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 12/32 (37%), Positives = 18/32 (56%), Gaps = 0/32 (0%)
Query 52 GGLICGLSLGFLYNKDMQDKPRWWSFAMWCSV 83
GG G + G +Y +D +D+ R + F CSV
Sbjct 171 GGFFAGTTDGRIYYEDFEDRSRSYVFNAHCSV 202
> CE00492
Length=397
Score = 29.6 bits (65), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 18/56 (32%), Positives = 30/56 (53%), Gaps = 3/56 (5%)
Query 45 IDQMGHLGGLICGLSLGFLYNKDMQDKP-RWWSFAMWCSVFLLVALPASCVPVIFA 99
+ GHLGGL+ G+ F+ + KP R+++ + W S+ L A C+ +I A
Sbjct 252 VSMYGHLGGLVAGILFTFILFRG--SKPSRFYTVSFWVSLVLSGFFIAICITLIAA 305
> At1g12750
Length=307
Score = 28.1 bits (61), Expect = 3.8, Method: Compositional matrix adjust.
Identities = 18/52 (34%), Positives = 26/52 (50%), Gaps = 8/52 (15%)
Query 26 IISFFLLMGLFAFTLNGGYIDQMGHLGGLICGLSLGFLYNKDMQDKPRWWSF 77
II+ L +GL ++D H+GGL+ G LGF+ MQ + W F
Sbjct 198 IIAINLAIGLLP------WVDNFAHIGGLLTGFCLGFIL--LMQPQSGWEEF 241
> 7292084_1
Length=356
Score = 28.1 bits (61), Expect = 3.8, Method: Composition-based stats.
Identities = 16/63 (25%), Positives = 28/63 (44%), Gaps = 15/63 (23%)
Query 42 GGYIDQMGHLGGLICGLSLGFL----YNKDMQDKPRWWSFAMWCSVFLLVALPASCVPVI 97
G + + HL G I GL++G L + + + ++ WW +AL V+
Sbjct 249 AGAVSYVAHLAGAIAGLTIGLLVLKSFEQKLHEQLLWW-----------IALGTYLALVV 297
Query 98 FAV 100
FA+
Sbjct 298 FAI 300
> 7292083
Length=355
Score = 28.1 bits (61), Expect = 4.4, Method: Composition-based stats.
Identities = 16/53 (30%), Positives = 28/53 (52%), Gaps = 7/53 (13%)
Query 42 GGYIDQMGHLGGLICGLSLGFLYNKDMQDKPR----WW-SFAMWC--SVFLLV 87
G + + HL G + GL++GFL K+ + WW + ++C +VF +V
Sbjct 273 GPQVSYIAHLTGALAGLTIGFLVLKNFGHREYEQLIWWLALGVYCAFTVFAIV 325
> 7292689
Length=529
Score = 26.9 bits (58), Expect = 8.1, Method: Compositional matrix adjust.
Identities = 20/77 (25%), Positives = 38/77 (49%), Gaps = 5/77 (6%)
Query 27 ISFFLLMGLFAFTLNGGYIDQMGHLGG---LICGLSLGFLYNKDMQDKPRWWSFAMWCSV 83
I +FL G+F + +D++G G L+C + GF N +D+ S + S
Sbjct 305 IIYFLAQGIFMWFPT--ILDELGTRNGENTLLCTVLQGFNINSSSEDEASSCSVEVDTST 362
Query 84 FLLVALPASCVPVIFAV 100
+ ++ + A+C VI+ +
Sbjct 363 YQVMIIIAACFVVIYLI 379
Lambda K H
0.333 0.145 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1167556980
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40