bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
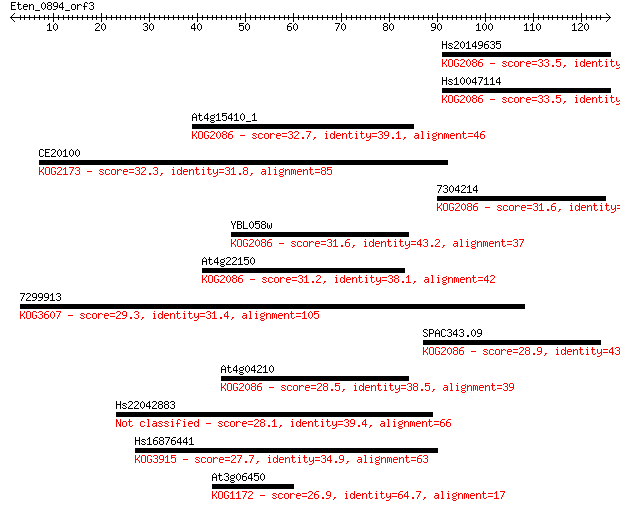
Query= Eten_0894_orf3
Length=125
Score E
Sequences producing significant alignments: (Bits) Value
Hs20149635 33.5 0.10
Hs10047114 33.5 0.11
At4g15410_1 32.7 0.17
CE20100 32.3 0.21
7304214 31.6 0.40
YBL058w 31.6 0.41
At4g22150 31.2 0.51
7299913 29.3 1.7
SPAC343.09 28.9 2.2
At4g04210 28.5 2.9
Hs22042883 28.1 4.2
Hs16876441 27.7 5.1
At3g06450 26.9 8.5
> Hs20149635
Length=370
Score = 33.5 bits (75), Expect = 0.10, Method: Compositional matrix adjust.
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query 91 FSLLEGFPPRPLSAARSATLQEAGLLNATLSQKLS 125
F L+ FP + L A S TL+EA LLNA + Q+L+
Sbjct 337 FILMTTFPNKEL-ADESQTLKEANLLNAVIVQRLT 370
> Hs10047114
Length=371
Score = 33.5 bits (75), Expect = 0.11, Method: Compositional matrix adjust.
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query 91 FSLLEGFPPRPLSAARSATLQEAGLLNATLSQKLS 125
F L+ FP + L A S TL+EA LLNA + Q+L+
Sbjct 338 FILMTTFPNKEL-ADESQTLKEANLLNAVIVQRLT 371
> At4g15410_1
Length=463
Score = 32.7 bits (73), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 18/51 (35%), Positives = 27/51 (52%), Gaps = 8/51 (15%)
Query 39 RVRAMAIRSLADLR-----GSGEDSPNENGATNSFSGGERSGLAIQNPSSA 84
R A IR+ ADL G G DS + A ++GG++SG+ +Q+P
Sbjct 129 RQHAGNIRTFADLNRSPADGEGSDS---DEANEYYTGGQKSGMMVQDPKKV 176
> CE20100
Length=880
Score = 32.3 bits (72), Expect = 0.21, Method: Composition-based stats.
Identities = 27/85 (31%), Positives = 39/85 (45%), Gaps = 8/85 (9%)
Query 7 LEREPPRSSATAGPSAAAAAAAAAGAAAASLLRVRAMAIRSLADLRGSGEDSPNENGATN 66
L +E PR++ + S A+ GA A +R+ A+ +R L D S N G T+
Sbjct 714 LYQEQPRAAESLSNSLRASGVDIDGAGAE--MRINALFLRGLHDESIIHSSSRNYGGTTS 771
Query 67 SFSGGERSGLAIQNPSSAFPAPGGF 91
SF+ A+Q S F P GF
Sbjct 772 SFN---MHPTAMQ---SVFAMPDGF 790
> 7304214
Length=407
Score = 31.6 bits (70), Expect = 0.40, Method: Compositional matrix adjust.
Identities = 17/35 (48%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Query 90 GFSLLEGFPPRPLSAARSATLQEAGLLNATLSQKL 124
F L+ FP R LS S T+++AGL NA L Q+L
Sbjct 373 NFILVSSFPTRELSDDNS-TIEKAGLKNAALMQRL 406
> YBL058w
Length=423
Score = 31.6 bits (70), Expect = 0.41, Method: Compositional matrix adjust.
Identities = 16/38 (42%), Positives = 25/38 (65%), Gaps = 2/38 (5%)
Query 47 SLADL-RGSGEDSPNENGATNSFSGGERSGLAIQNPSS 83
S +D+ RG +D +E+ N+F+GGE SGL + +PS
Sbjct 128 SFSDMVRGQADDD-DEDQPRNTFAGGETSGLEVTDPSD 164
> At4g22150
Length=302
Score = 31.2 bits (69), Expect = 0.51, Method: Compositional matrix adjust.
Identities = 16/44 (36%), Positives = 27/44 (61%), Gaps = 2/44 (4%)
Query 41 RAMAIRSLADL--RGSGEDSPNENGATNSFSGGERSGLAIQNPS 82
R IR+L+DL R + + +G F+GGE+SG+ +Q+P+
Sbjct 15 RTGGIRTLSDLNRRSEPDSDSDSDGPQEYFTGGEKSGMLVQDPT 58
> 7299913
Length=1569
Score = 29.3 bits (64), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 33/108 (30%), Positives = 49/108 (45%), Gaps = 5/108 (4%)
Query 3 LRGRLEREPPRSSATAGPSAAAAAAAAAGAAAASLLRVR-AMAIRSLADLRGSGEDSPNE 61
LR +L+ PR +AT P A A + ASL + I + + G+G P
Sbjct 931 LRRKLDITAPRLNATTNPLALTEGAQFIQSDPASLKSSGGSKPITTRTNQNGTG--PPPS 988
Query 62 NGATNSF--SGGERSGLAIQNPSSAFPAPGGFSLLEGFPPRPLSAARS 107
NG S S + +++PS + P+ G F +GF RPL+ A S
Sbjct 989 NGLLKSTPSSTDDMDSALLKSPSDSTPSAGLFGKFDGFTLRPLNDASS 1036
> SPAC343.09
Length=410
Score = 28.9 bits (63), Expect = 2.2, Method: Compositional matrix adjust.
Identities = 16/37 (43%), Positives = 21/37 (56%), Gaps = 0/37 (0%)
Query 87 APGGFSLLEGFPPRPLSAARSATLQEAGLLNATLSQK 123
+PG F L FP + L S T++ A L NA+L QK
Sbjct 372 SPGNFILSVPFPAKTLEDDPSVTVEAASLKNASLVQK 408
> At4g04210
Length=308
Score = 28.5 bits (62), Expect = 2.9, Method: Compositional matrix adjust.
Identities = 15/41 (36%), Positives = 26/41 (63%), Gaps = 2/41 (4%)
Query 45 IRSLADL--RGSGEDSPNENGATNSFSGGERSGLAIQNPSS 83
IR+L+DL R + + +G ++GGE+SG+ +Q+PS
Sbjct 18 IRTLSDLNRRSGPDSDSDSDGPQEYYTGGEKSGMLVQDPSK 58
> Hs22042883
Length=268
Score = 28.1 bits (61), Expect = 4.2, Method: Compositional matrix adjust.
Identities = 26/79 (32%), Positives = 34/79 (43%), Gaps = 13/79 (16%)
Query 23 AAAAAAAAGAAAASLLRVRAMAIRSLADL------------RGSGEDSPNE-NGATNSFS 69
A A + AA +S RV+AMA+R ++ R G P E +GA N S
Sbjct 184 TAMALTQSRAACSSEYRVKAMAVRGRPEVQAISLPPPQGTWRLHGRRLPTEADGAQNELS 243
Query 70 GGERSGLAIQNPSSAFPAP 88
G R LA PS +P
Sbjct 244 GKPRLHLAKNRPSLKTASP 262
> Hs16876441
Length=599
Score = 27.7 bits (60), Expect = 5.1, Method: Compositional matrix adjust.
Identities = 22/63 (34%), Positives = 30/63 (47%), Gaps = 1/63 (1%)
Query 27 AAAAGAAAASLLRVRAMAIRSLADLRGSGEDSPNENGATNSFSGGERSGLAIQNPSSAFP 86
AAA A A L +++ MA+ +L GS + +E NS +GG S S F
Sbjct 216 TAAAMAEAMKLQKMKLMAMNTLQG-NGSQNGTESEPDDLNSNTGGSESSWDKDKMQSPFA 274
Query 87 APG 89
APG
Sbjct 275 APG 277
> At3g06450
Length=736
Score = 26.9 bits (58), Expect = 8.5, Method: Compositional matrix adjust.
Identities = 11/17 (64%), Positives = 13/17 (76%), Gaps = 0/17 (0%)
Query 43 MAIRSLADLRGSGEDSP 59
+ IRSL+D RG GE SP
Sbjct 689 VYIRSLSDFRGGGEISP 705
Lambda K H
0.309 0.125 0.339
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1183965632
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40