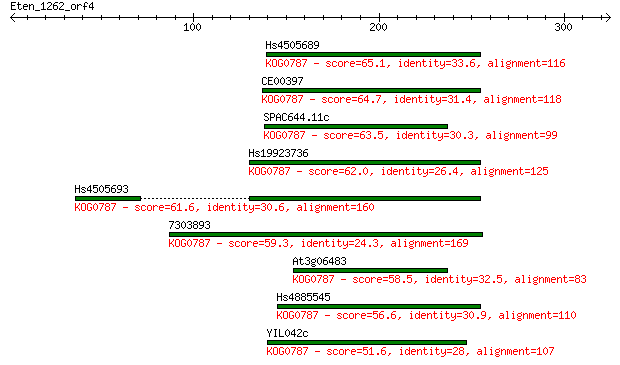
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_1262_orf4
Length=324
Score E
Sequences producing significant alignments: (Bits) Value
Hs4505689 65.1 2e-10
CE00397 64.7 3e-10
SPAC644.11c 63.5 6e-10
Hs19923736 62.0 1e-09
Hs4505693 61.6 2e-09
7303893 59.3 1e-08
At3g06483 58.5 2e-08
Hs4885545 56.6 7e-08
YIL042c 51.6 2e-06
> Hs4505689
Length=436
Score = 65.1 bits (157), Expect = 2e-10, Method: Compositional matrix adjust.
Identities = 39/116 (33%), Positives = 61/116 (52%), Gaps = 7/116 (6%)
Query 139 PPVDIHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNKTRAGSAAAL 198
PP+ +H+ L E + VK+ D G G+ L + + ++YST P R +RA A
Sbjct 299 PPIQVHVTLGNED-LTVKMSDRGGGVPLRKIDRLFNYMYSTAPRP--RVETSRAVPLA-- 353
Query 199 FSGWGVGLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTASPQTNSAA 254
G+G GLP++R+YA+ GD + S G GT + +Y+ +S + P N AA
Sbjct 354 --GFGYGLPISRLYAQYFQGDLKLYSLEGYGTDAVIYIKALSTDSIERLPVYNKAA 407
> CE00397
Length=401
Score = 64.7 bits (156), Expect = 3e-10, Method: Compositional matrix adjust.
Identities = 37/118 (31%), Positives = 64/118 (54%), Gaps = 8/118 (6%)
Query 137 KLPPVDIHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNKTRAGSAA 196
LP + ++++ E + +K+CD G G+ T + + ++YST P R G+ A
Sbjct 265 DLPDIKVYVVKGQED-LSIKICDRGGGVSRTILERLYNYMYSTAPPP------PRDGTQA 317
Query 197 ALFSGWGVGLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTASPQTNSAA 254
L +G+G GLPL+R+YAR GD F+ S G GT + +Y+ + A P ++++
Sbjct 318 PL-AGYGYGLPLSRLYARYFLGDLFLVSMEGHGTDACIYLKAVPVEASEVLPIYSTSS 374
> SPAC644.11c
Length=425
Score = 63.5 bits (153), Expect = 6e-10, Method: Compositional matrix adjust.
Identities = 30/101 (29%), Positives = 53/101 (52%), Gaps = 4/101 (3%)
Query 138 LPPVDIHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTR--TMPVDRQNKTRAGSA 195
PP+ + ++ + + +K+ D G G+ L W ++++T T+ D + A S
Sbjct 316 FPPIKV-IVAKGQEDITIKISDEGGGISRRNIPLVWSYMFTTASPTLTDDPHDIVSANST 374
Query 196 AALFSGWGVGLPLTRMYARALGGDAFMESRIGRGTRSYLYV 236
+ +G+G GLPL R+Y R GGD + S G GT Y+++
Sbjct 375 TPM-AGFGFGLPLARLYTRYFGGDLELISMEGYGTDVYIHL 414
> Hs19923736
Length=407
Score = 62.0 bits (149), Expect = 1e-09, Method: Compositional matrix adjust.
Identities = 33/125 (26%), Positives = 65/125 (52%), Gaps = 8/125 (6%)
Query 130 QQQQQAIKLPPVDIHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNK 189
+ + ++ LPP+ + ++ E + +K+ D G G+ L + + + ++YST P
Sbjct 262 ESHESSLILPPIKV-MVALGEEDLSIKMSDRGGGVPLRKIERLFSYMYSTAPTP------ 314
Query 190 TRAGSAAALFSGWGVGLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTASPQ 249
+ G+ +G+G GLP++R+YA+ GD + S G GT + +Y+ +S + P
Sbjct 315 -QPGTGGTPLAGFGYGLPISRLYAKYFQGDLQLFSMEGFGTDAVIYLKALSTDSVERLPV 373
Query 250 TNSAA 254
N +A
Sbjct 374 YNKSA 378
> Hs4505693
Length=411
Score = 61.6 bits (148), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 36/125 (28%), Positives = 62/125 (49%), Gaps = 7/125 (5%)
Query 130 QQQQQAIKLPPVDIHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNK 189
+ Q+ L P+++ ++L E + +K+ D G G+ L + + YST PV
Sbjct 265 EHQENQPSLTPIEVIVVLGKED-LTIKISDRGGGVPLRIIDRLFSYTYSTAPTPV----- 318
Query 190 TRAGSAAALFSGWGVGLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTASPQ 249
S A +G+G GLP++R+YA+ GD + S G GT + +Y+ +S + P
Sbjct 319 -MDNSRNAPLAGFGYGLPISRLYAKYFQGDLNLYSLSGYGTDAIIYLKALSSESIEKLPV 377
Query 250 TNSAA 254
N +A
Sbjct 378 FNKSA 382
Score = 32.0 bits (71), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 13/35 (37%), Positives = 22/35 (62%), Gaps = 0/35 (0%)
Query 36 DNRVVVRCAPAYVYSGCFELIKNAIEATVQRQQKQ 70
D + + P++++ FEL KNA+ ATV+ Q+ Q
Sbjct 236 DQPIHIVYVPSHLHHMLFELFKNAMRATVEHQENQ 270
> 7303893
Length=413
Score = 59.3 bits (142), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 41/173 (23%), Positives = 78/173 (45%), Gaps = 17/173 (9%)
Query 87 VEEEQQQEWDALPVRQ----SRCIYSLPALYGAPSQEQQQLQLPQQQQQQQQAIKLPPVD 142
+++ + D LP+R S Y L L+ + ++ + LPP+
Sbjct 225 IQQHSSEPGDNLPIRTVYVPSHLYYMLFELF------KNSMRAVVEHHGHDNNDTLPPLK 278
Query 143 IHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNKTRAGSAAALFSGW 202
+ I + + VK+ D G G+ ++ ++++YST P +++ +G+
Sbjct 279 V-AICKGKEDICVKISDQGGGIPRSQTDQLFKYMYSTAPQP------SKSDLHTVPLAGY 331
Query 203 GVGLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTASPQTNSAAA 255
G GLP++R+YAR GD + S G GT + +Y+ +S A P N ++
Sbjct 332 GYGLPISRLYARYFHGDIVLLSCEGFGTDAIIYLKALSDEANELLPIFNKTSS 384
> At3g06483
Length=380
Score = 58.5 bits (140), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 27/83 (32%), Positives = 44/83 (53%), Gaps = 0/83 (0%)
Query 154 VVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNKTRAGSAAALFSGWGVGLPLTRMYA 213
++ V D G G+ + + +LYST P++ +G+G GLP++R+YA
Sbjct 287 IIVVSDEGGGIARSGLPRIFTYLYSTARNPLEEDVDLGIADVPVTMAGYGYGLPISRLYA 346
Query 214 RALGGDAFMESRIGRGTRSYLYV 236
R GGD + S G GT +YL++
Sbjct 347 RYFGGDLQIISMEGYGTDAYLHL 369
> Hs4885545
Length=406
Score = 56.6 bits (135), Expect = 7e-08, Method: Compositional matrix adjust.
Identities = 34/110 (30%), Positives = 55/110 (50%), Gaps = 6/110 (5%)
Query 145 LILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTMPVDRQNKTRAGSAAALFSGWGV 204
L+ + + +K+ D G G+ L + + ++YST P TRA A G+G
Sbjct 273 LVTLGKEDLSIKISDLGGGVPLRKIDRLFNYMYSTAPRP--SLEPTRAAPLA----GFGY 326
Query 205 GLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTASPQTNSAA 254
GLP++R+YAR GD + S G GT + +Y+ +S + P N +A
Sbjct 327 GLPISRLYARYFQGDLKLYSMEGVGTDAVIYLKALSSESFERLPVFNKSA 376
> YIL042c
Length=394
Score = 51.6 bits (122), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 30/108 (27%), Positives = 53/108 (49%), Gaps = 1/108 (0%)
Query 140 PVDIHLILAAESGVVVKVCDSGNGLCLTEQQLAWRFLYSTRTM-PVDRQNKTRAGSAAAL 198
P++I+L+ + + +++ D G G+ + L + + YST T D ++ G
Sbjct 285 PIEINLLKPDDDELYLRIRDHGGGITPEVEALMFNYSYSTHTQQSADSESTDLPGEQINN 344
Query 199 FSGWGVGLPLTRMYARALGGDAFMESRIGRGTRSYLYVPDISPTACTA 246
SG G GLP+ + Y GG ++S +G GT Y+ + S TA +
Sbjct 345 VSGMGFGLPMCKTYLELFGGKIDVQSLLGWGTDVYIKLKGPSKTALLS 392
Lambda K H
0.316 0.125 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 7566324290
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40