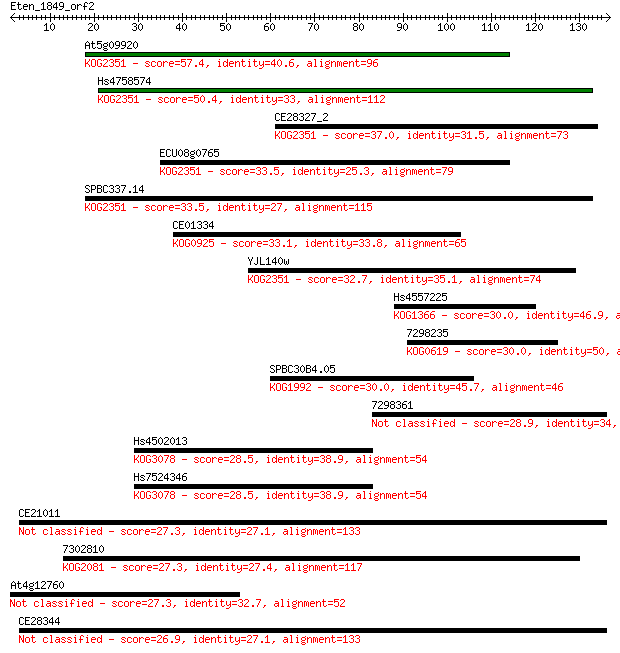
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_1849_orf2
Length=136
Score E
Sequences producing significant alignments: (Bits) Value
At5g09920 57.4 8e-09
Hs4758574 50.4 1e-06
CE28327_2 37.0 0.010
ECU08g0765 33.5 0.11
SPBC337.14 33.5 0.13
CE01334 33.1 0.14
YJL140w 32.7 0.21
Hs4557225 30.0 1.2
7298235 30.0 1.2
SPBC30B4.05 30.0 1.4
7298361 28.9 2.9
Hs4502013 28.5 4.0
Hs7524346 28.5 4.2
CE21011 27.3 7.9
7302810 27.3 8.2
At4g12760 27.3 9.0
CE28344 26.9 9.7
> At5g09920
Length=138
Score = 57.4 bits (137), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 39/100 (39%), Positives = 55/100 (55%), Gaps = 5/100 (5%)
Query 18 LGPEFKNAKCLNLCELQLILG---DQLRLSVARPA-EAHNLIKASYEYASRFGKMTVRTA 73
+G EF AKCL CE+ LIL +QL+ P + + + S +Y RF + A
Sbjct 14 IGDEFLKAKCLMNCEVSLILEHKFEQLQQISEDPMNQVSQVFEKSLQYVKRFSRYKNPDA 73
Query 74 VVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSL 113
V ++R+ L R L +FEL +L NL P T +EA A +PSL
Sbjct 74 VRQVREILSRH-QLTEFELCVLGNLCPETVEEAVAMVPSL 112
> Hs4758574
Length=142
Score = 50.4 bits (119), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 37/115 (32%), Positives = 56/115 (48%), Gaps = 4/115 (3%)
Query 21 EFKNAKCLNLCELQLILG--DQLRLSVARPAEAHNLIKASYEYASRFGKMTVRTAVVEIR 78
EF+ A+ L E+ ++L Q S E + + Y +RF + R + +R
Sbjct 25 EFETAETLLNSEVHMLLEHRKQQNESAEDEQELSEVFMKTLNYTARFSRFKNRETIASVR 84
Query 79 KHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSLI-RLSAARLSRIIDTLEVFR 132
L + L +FELA L NL P T +E+KA IPSL R L +I+D ++ R
Sbjct 85 SLL-LQKKLHKFELACLANLCPETAEESKALIPSLEGRFEDEELQQILDDIQTKR 138
> CE28327_2
Length=133
Score = 37.0 bits (84), Expect = 0.010, Method: Compositional matrix adjust.
Identities = 23/74 (31%), Positives = 37/74 (50%), Gaps = 2/74 (2%)
Query 61 YASRFGKMTVRTAVVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSL-IRLSAA 119
YA R + R + +R E L +FE+A + NL P +EAKA +PSL ++ +
Sbjct 59 YARRMSRFKNRETIRAVRAIF-SEKHLHKFEVAQIANLCPENAEEAKALVPSLENKIEES 117
Query 120 RLSRIIDTLEVFRV 133
L ++ L+ R
Sbjct 118 ELEEVLKDLQSKRT 131
> ECU08g0765
Length=104
Score = 33.5 bits (75), Expect = 0.11, Method: Compositional matrix adjust.
Identities = 20/79 (25%), Positives = 41/79 (51%), Gaps = 1/79 (1%)
Query 35 LILGDQLRLSVARPAEAHNLIKASYEYASRFGKMTVRTAVVEIRKHLEREGDLQQFELAL 94
L+ G + R + A + +++ Y F ++ ++ ++R L G + E+AL
Sbjct 8 LLEGQKERFRADFRSNASKVFRSTLGYLDDFCRIKDKSVAEDLRTTLSGLG-FGEVEIAL 66
Query 95 LVNLLPRTPDEAKAHIPSL 113
+L P++ +EAK+ +PSL
Sbjct 67 FGSLFPQSVEEAKSLVPSL 85
> SPBC337.14
Length=135
Score = 33.5 bits (75), Expect = 0.13, Method: Compositional matrix adjust.
Identities = 31/120 (25%), Positives = 52/120 (43%), Gaps = 8/120 (6%)
Query 18 LGPEFKNAKCLNLCELQLILGDQLRLSVARPAEAH----NLIKASYEYASRFGKMTVRTA 73
LGPEF+N L + E ++++ + + AR +++K + Y + F + A
Sbjct 15 LGPEFENEDMLTVSEAKILI-ETVLAQRARETNGEIPMTDVMKKTVAYFNVFARFKTAEA 73
Query 74 VVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSLI-RLSAARLSRIIDTLEVFR 132
+ L +FE A L L +EA+ IPSL ++ L I+D L R
Sbjct 74 TYACERILGNR--FHKFERAQLGTLCCEDAEEARTLIPSLANKIDDQNLQGILDELSTLR 131
> CE01334
Length=739
Score = 33.1 bits (74), Expect = 0.14, Method: Compositional matrix adjust.
Identities = 22/69 (31%), Positives = 34/69 (49%), Gaps = 4/69 (5%)
Query 38 GDQLRLSVARPAEAHNLIKASYEYASRFGKMTVRTAVVE----IRKHLEREGDLQQFELA 93
D +R ++R + +NL + S ++ SR + +R A+V HLER G +
Sbjct 601 ADTVRTQLSRVMDKYNLRRVSTDFKSRDYYLNIRKALVAGFFMQVAHLERSGHYVTVKDN 660
Query 94 LLVNLLPRT 102
LVNL P T
Sbjct 661 QLVNLHPST 669
> YJL140w
Length=221
Score = 32.7 bits (73), Expect = 0.21, Method: Compositional matrix adjust.
Identities = 26/75 (34%), Positives = 40/75 (53%), Gaps = 2/75 (2%)
Query 55 IKASYEYASRFGKMTVRTAVVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSL- 113
+K + +Y + F + + V + + L+ G L FE+A L +L T DEAK IPSL
Sbjct 141 LKNTMQYLTNFSRFRDQETVGAVIQLLKSTG-LHPFEVAQLGSLACDTADEAKTLIPSLN 199
Query 114 IRLSAARLSRIIDTL 128
++S L RI+ L
Sbjct 200 NKISDDELERILKEL 214
> Hs4557225
Length=1474
Score = 30.0 bits (66), Expect = 1.2, Method: Composition-based stats.
Identities = 15/32 (46%), Positives = 19/32 (59%), Gaps = 0/32 (0%)
Query 88 QQFELALLVNLLPRTPDEAKAHIPSLIRLSAA 119
++F AL V LP+T DE KAH I LS +
Sbjct 1337 EEFPFALGVQTLPQTCDEPKAHTSFQISLSVS 1368
> 7298235
Length=1021
Score = 30.0 bits (66), Expect = 1.2, Method: Compositional matrix adjust.
Identities = 17/36 (47%), Positives = 22/36 (61%), Gaps = 2/36 (5%)
Query 91 ELALLVNLLPRTPDEAKAHIPSLIR--LSAARLSRI 124
EL L N + R PD+A H+P L++ LS RLS I
Sbjct 188 ELRLSGNPILRVPDDAFGHVPQLVKLELSDCRLSHI 223
> SPBC30B4.05
Length=967
Score = 30.0 bits (66), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 21/51 (41%), Positives = 29/51 (56%), Gaps = 5/51 (9%)
Query 60 EYASRFGK-MTVRTAVVEIRKH---LEREGDLQQF-ELALLVNLLPRTPDE 105
+Y GK M T+V+ IRKH +++ LQQF EL +L N+ R DE
Sbjct 300 KYDGLVGKAMAFLTSVIRIRKHAEFFQQDQVLQQFIELVVLPNICLRESDE 350
> 7298361
Length=1441
Score = 28.9 bits (63), Expect = 2.9, Method: Composition-based stats.
Identities = 18/54 (33%), Positives = 30/54 (55%), Gaps = 2/54 (3%)
Query 83 REGDLQQFELALLVNLLPRT-PDEAKAHIPSLIRLSAARLSRIIDTLEVFRVHA 135
++G + + ALL +L +T PD + P LI S +L ++ TL++ R HA
Sbjct 157 KKGGIMKGTRALLAHLSSQTRPDNVQGE-PILIETSEKQLEEVVITLDIGRAHA 209
> Hs4502013
Length=239
Score = 28.5 bits (62), Expect = 4.0, Method: Compositional matrix adjust.
Identities = 21/55 (38%), Positives = 27/55 (49%), Gaps = 5/55 (9%)
Query 29 NLCELQLILGDQLRLSVARPAEAHNLIKASYEYASRFGKMTVRTAVVE-IRKHLE 82
N C L GD LR VA +E +KA+ + GK+ VVE I K+LE
Sbjct 38 NFCVCHLATGDMLRAMVASGSELGKKLKATMDA----GKLVSDEMVVELIEKNLE 88
> Hs7524346
Length=232
Score = 28.5 bits (62), Expect = 4.2, Method: Compositional matrix adjust.
Identities = 21/55 (38%), Positives = 27/55 (49%), Gaps = 5/55 (9%)
Query 29 NLCELQLILGDQLRLSVARPAEAHNLIKASYEYASRFGKMTVRTAVVE-IRKHLE 82
N C L GD LR VA +E +KA+ + GK+ VVE I K+LE
Sbjct 38 NFCVCHLATGDMLRAMVASGSELGKKLKATMDA----GKLVSDEMVVELIEKNLE 88
> CE21011
Length=650
Score = 27.3 bits (59), Expect = 7.9, Method: Compositional matrix adjust.
Identities = 36/142 (25%), Positives = 60/142 (42%), Gaps = 16/142 (11%)
Query 3 WPKMEASVPESLAGDLGPEFKNAKCLNLCELQLILGDQLRLSVARPAE---------AHN 53
+P + A + G +G F A C + EL+LI+ D + PAE
Sbjct 162 FPSIIADMLSDAIGCVG--FSWAACPAMTELELIMLDWFGKMIGLPAEFLPLTENGKGGG 219
Query 54 LIKASYEYASRFGKMTVRTAVVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSL 113
+I++S AS +T+ A E+ K L + + E LL L+ EA + +
Sbjct 220 VIQSS---ASECNFVTLLAARFEVMKELRQRFPFVE-EGLLLSKLIAYCSKEAHSSVEKA 275
Query 114 IRLSAARLSRIIDTLEVFRVHA 135
+ +L RI++T FR+
Sbjct 276 CMIGMVKL-RILETDSKFRLRG 296
> 7302810
Length=571
Score = 27.3 bits (59), Expect = 8.2, Method: Compositional matrix adjust.
Identities = 32/117 (27%), Positives = 58/117 (49%), Gaps = 6/117 (5%)
Query 13 SLAGDLGPEFKNAKCLNLCELQLILGDQLRLSVARPAEAHNLIKASYEYASRFGKMTVRT 72
SLA + F + + L L+ L D L+ + A ++++LI S G M +
Sbjct 48 SLAKKIKTSFYSLQKFQLMSLKNSLIDHLKYA-AMMRDSNSLIVQLAVGISALGLMFSQW 106
Query 73 AVVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSLIRLSAARLSRIIDTLE 129
E++ + + + Q+ +ALL +L P+E + PS + + A +L+ +IDTLE
Sbjct 107 DY-ELQDFVRKLSENPQYVMALL-EVLKVIPEETR---PSNLPVEAKKLNSVIDTLE 158
> At4g12760
Length=227
Score = 27.3 bits (59), Expect = 9.0, Method: Compositional matrix adjust.
Identities = 17/54 (31%), Positives = 28/54 (51%), Gaps = 11/54 (20%)
Query 1 WCWPKMEASVPESLAGDLGPEFKNAKCLNLCELQLILGDQLRLSV--ARPAEAH 52
WC P G+L ++ C N+C++QL G+++ L++ R AEAH
Sbjct 144 WC--------PICKKGELMENHRHIDC-NMCDMQLNKGEEVNLNILQERLAEAH 188
> CE28344
Length=705
Score = 26.9 bits (58), Expect = 9.7, Method: Compositional matrix adjust.
Identities = 36/142 (25%), Positives = 60/142 (42%), Gaps = 16/142 (11%)
Query 3 WPKMEASVPESLAGDLGPEFKNAKCLNLCELQLILGDQLRLSVARPAE---------AHN 53
+P + A + G +G F A C + EL+LI+ D + PAE
Sbjct 162 FPSIIADMLSDAIGCVG--FSWAACPAMTELELIMLDWFGKMIGLPAEFLPLTENGKGGG 219
Query 54 LIKASYEYASRFGKMTVRTAVVEIRKHLEREGDLQQFELALLVNLLPRTPDEAKAHIPSL 113
+I++S AS +T+ A E+ K L + + E LL L+ EA + +
Sbjct 220 VIQSS---ASECNFVTLLAARFEVMKELRQRFPFVE-EGLLLSKLIAYCSKEAHSSVEKA 275
Query 114 IRLSAARLSRIIDTLEVFRVHA 135
+ +L RI++T FR+
Sbjct 276 CMIGMVKL-RILETDSKFRLRG 296
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1425342594
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40