bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_2248_orf2
Length=175
Score E
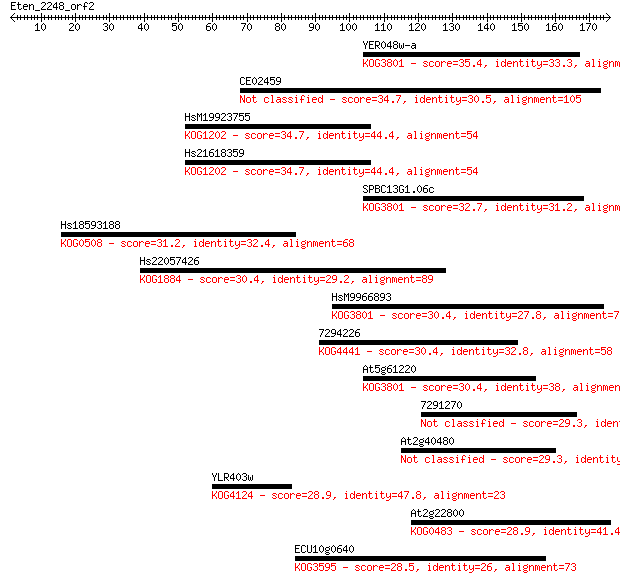
Sequences producing significant alignments: (Bits) Value
YER048w-a 35.4 0.065
CE02459 34.7 0.094
HsM19923755 34.7 0.11
Hs21618359 34.7 0.11
SPBC13G1.06c 32.7 0.36
Hs18593188 31.2 1.00
Hs22057426 30.4 1.7
HsM9966893 30.4 1.7
7294226 30.4 1.8
At5g61220 30.4 1.9
7291270 29.3 4.1
At2g40480 29.3 4.3
YLR403w 28.9 5.3
At2g22800 28.9 5.7
ECU10g0640 28.5 7.2
> YER048w-a
Length=94
Score = 35.4 bits (80), Expect = 0.065, Method: Compositional matrix adjust.
Identities = 21/67 (31%), Positives = 34/67 (50%), Gaps = 4/67 (5%)
Query 104 IHLCKEVLRAARSFKNLNFRDYFVRRAKEKLRELRCTDTADAEAQLKKEIED----LKRQ 159
+ L KE ++ A F N NFR+YF+ + + R+ L KE ++ LKRQ
Sbjct 13 LSLYKEFIKNANQFNNYNFREYFLSKTRTTFRKNMNQQDPKVLMNLFKEAKNDLGVLKRQ 72
Query 160 ATITNIY 166
+ I+ +Y
Sbjct 73 SVISQMY 79
> CE02459
Length=333
Score = 34.7 bits (78), Expect = 0.094, Method: Compositional matrix adjust.
Identities = 32/113 (28%), Positives = 50/113 (44%), Gaps = 10/113 (8%)
Query 68 TTSFHHFNHFNSFFHLNCAHLSIPTKGLACRAKPLDIHLCKEVLRAARSFKNLNFRDYFV 127
+ S H F+ + H + LS+ G A P D HL E + S+ L+
Sbjct 135 SISVHGFSDSDVNHHRQVSILSVIISG-AGEILPFDSHLKDEPKQTEMSYIELDKGILLP 193
Query 128 RRAKEKLRELRC------TDTADAEAQLKKEIED--LKRQATITNIYHLPTVT 172
++K K+RE + TD+ D E KE+ + KR+A + N LPT +
Sbjct 194 LKSKPKMRETKTIVGLYDTDSEDEEILFSKEMLNRLRKREAVMKN-PQLPTTS 245
> HsM19923755
Length=2509
Score = 34.7 bits (78), Expect = 0.11, Method: Composition-based stats.
Identities = 24/56 (42%), Positives = 27/56 (48%), Gaps = 3/56 (5%)
Query 52 PELRALNWSFSAVANPTTSFHHFNHFNSFFHLNCAHLSIPTKGLAC-RAKPLD-IH 105
P L LN S + P H + FH + LSIPT GL C RA PLD IH
Sbjct 2227 PTLMRLN-SVQSSERPLFLVHPIEGSTTVFHSLASRLSIPTYGLQCTRAAPLDSIH 2281
> Hs21618359
Length=2509
Score = 34.7 bits (78), Expect = 0.11, Method: Composition-based stats.
Identities = 24/56 (42%), Positives = 27/56 (48%), Gaps = 3/56 (5%)
Query 52 PELRALNWSFSAVANPTTSFHHFNHFNSFFHLNCAHLSIPTKGLAC-RAKPLD-IH 105
P L LN S + P H + FH + LSIPT GL C RA PLD IH
Sbjct 2227 PTLMRLN-SVQSSERPLFLVHPIEGSTTVFHSLASRLSIPTYGLQCTRAAPLDSIH 2281
> SPBC13G1.06c
Length=102
Score = 32.7 bits (73), Expect = 0.36, Method: Compositional matrix adjust.
Identities = 20/68 (29%), Positives = 41/68 (60%), Gaps = 5/68 (7%)
Query 104 IHLCKEVLRAARSFKNLNFRDYFVRRAKEKLRELRC-TDTADAEAQLK---KEIEDLKRQ 159
+ L + +L+ ++ F +R+Y +RR ++K +EL+ +D A E +K K +E ++RQ
Sbjct 9 VRLYRNILKTSKLFP-YTYREYTIRRTRDKFKELKVESDPAKFEQGIKDSEKLLEIIQRQ 67
Query 160 ATITNIYH 167
+ I +Y+
Sbjct 68 SIINGMYN 75
> Hs18593188
Length=1172
Score = 31.2 bits (69), Expect = 1.00, Method: Composition-based stats.
Identities = 22/71 (30%), Positives = 30/71 (42%), Gaps = 3/71 (4%)
Query 16 SFSSFQSKLFLFSFL---PMAAAAAAGRCGFAFSPGVCTPELRALNWSFSAVANPTTSFH 72
S SSF S LFS++ P A + + GFA GV T +R + W+ + P S
Sbjct 934 SASSFLSFAELFSYVLQDPAAKGSLGTQIGFADLMGVLTKGVREVEWALQLLREPRDSAQ 993
Query 73 HFNHFNSFFHL 83
HL
Sbjct 994 FNKALAIILHL 1004
> Hs22057426
Length=2407
Score = 30.4 bits (67), Expect = 1.7, Method: Composition-based stats.
Identities = 26/104 (25%), Positives = 43/104 (41%), Gaps = 15/104 (14%)
Query 39 GRCGF---AFSPGVCTPELRALNW---SFSAVANPTTSFH---HFNHFNSFFHLNCAHLS 89
GRC A G+ +L +N+ S + NP++ F+H +S +L
Sbjct 524 GRCSIPIHALEAGIENRQLDTVNFFLKSKENLFNPSSKSSVSDQFDHLSSHLYLRNVEEL 583
Query 90 IPTKGLACRA------KPLDIHLCKEVLRAARSFKNLNFRDYFV 127
IP L C A +P H +++L SF N ++ F+
Sbjct 584 IPALDLLCSAIRESYSEPQSKHFSEQLLNLTLSFLNNQIKELFI 627
> HsM9966893
Length=91
Score = 30.4 bits (67), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 22/83 (26%), Positives = 40/83 (48%), Gaps = 4/83 (4%)
Query 95 LACRAKPLDIHLCKEVLRAARSFKNLNFRDYFVRRAKEKLRELR-CTDTADAEAQLKKEI 153
+A ++ + L + +LR ++ F N+R Y VRR ++ RE + D + + + K
Sbjct 1 MAASSRAQVLSLYRAMLRESKRFSAYNYRTYAVRRIRDAFRENKNVKDPVEIQTLVNKAK 60
Query 154 EDL---KRQATITNIYHLPTVTI 173
DL +RQ I +Y + I
Sbjct 61 RDLGVIRRQVHIGQLYSTDKLII 83
> 7294226
Length=623
Score = 30.4 bits (67), Expect = 1.8, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 6/63 (9%)
Query 91 PTKGLACRAKPLDIHLCKEVLRAARSFKNLNFRDY-----FVRRAKEKLRELRCTDTADA 145
PT L RA D H C+E+LR A F NF++ F+ +L ++ C+D +
Sbjct 171 PTNCLGIRAFA-DTHSCRELLRIADKFTQHNFQEVMESEEFLLLPVGQLVDIICSDELNV 229
Query 146 EAQ 148
++
Sbjct 230 RSE 232
> At5g61220
Length=87
Score = 30.4 bits (67), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 19/54 (35%), Positives = 30/54 (55%), Gaps = 4/54 (7%)
Query 104 IHLCKEVLRAARSFKNLNFRDYFVRRAKEKLRELR-CTD---TADAEAQLKKEI 153
+ LC+ +LRA R F + N R+Y RR + R + TD +A A+ KK++
Sbjct 8 LSLCRALLRAGRQFPDYNIREYSKRRTLDGFRMNKNLTDPSKVTEAYAEAKKQL 61
> 7291270
Length=587
Score = 29.3 bits (64), Expect = 4.1, Method: Composition-based stats.
Identities = 18/48 (37%), Positives = 27/48 (56%), Gaps = 3/48 (6%)
Query 121 NFRDYFVRRAKEKLRELRCTDTADAEAQL---KKEIEDLKRQATITNI 165
+F FV +AK+K RC A+AE+ L KK I L+R + ++ I
Sbjct 244 DFNATFVNQAKKKRCARRCEKMAEAESNLLDRKKSIVRLQRPSEMSGI 291
> At2g40480
Length=541
Score = 29.3 bits (64), Expect = 4.3, Method: Composition-based stats.
Identities = 16/54 (29%), Positives = 27/54 (50%), Gaps = 9/54 (16%)
Query 115 RSFKNLNFRDYFVRRAKEK---------LRELRCTDTADAEAQLKKEIEDLKRQ 159
R F N+ ++ ++R +E ++EL D +A K+ +EDLKRQ
Sbjct 107 RGFSNIYNGEFDIKRMEEHAAELEKDLIVKELETLDVLEALGSTKRIVEDLKRQ 160
> YLR403w
Length=683
Score = 28.9 bits (63), Expect = 5.3, Method: Composition-based stats.
Identities = 11/23 (47%), Positives = 16/23 (69%), Gaps = 0/23 (0%)
Query 60 SFSAVANPTTSFHHFNHFNSFFH 82
S ++V NPTT H++N F+S H
Sbjct 500 SMNSVVNPTTGSHNYNTFHSSVH 522
> At2g22800
Length=274
Score = 28.9 bits (63), Expect = 5.7, Method: Compositional matrix adjust.
Identities = 24/74 (32%), Positives = 36/74 (48%), Gaps = 16/74 (21%)
Query 118 KNLNFRDYFV------RRAKEKLREL--------RCTDT-ADAEAQLKKEIEDLKRQATI 162
+ LN R V RRA+ KL++ +C +T AD +L+KEI++LK
Sbjct 145 RQLNLRPRQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLADENIRLQKEIQELKTLKLT 204
Query 163 TNIY-HLPTVTIYK 175
Y H+P T+ K
Sbjct 205 QPFYMHMPASTLTK 218
> ECU10g0640
Length=3151
Score = 28.5 bits (62), Expect = 7.2, Method: Composition-based stats.
Identities = 19/77 (24%), Positives = 39/77 (50%), Gaps = 9/77 (11%)
Query 84 NCAHLSIPTKGLACRAKPLDIHLCKEVLRAARSFKNLNF-RDYFVRRAKEKLRELR---C 139
+C H SI T G C+A I +R ++ F++ R F+ +K+ + R C
Sbjct 2398 SCLHPSIVTTGFLCKA----IEFGMSFIRISKEFRDKEIRRRQFLTEGLQKIEDFREEAC 2453
Query 140 TDTADAEAQLKKEIEDL 156
+ +++ + +K+++DL
Sbjct 2454 SLGSESSRK-RKDLQDL 2469
Lambda K H
0.330 0.138 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2671071884
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40