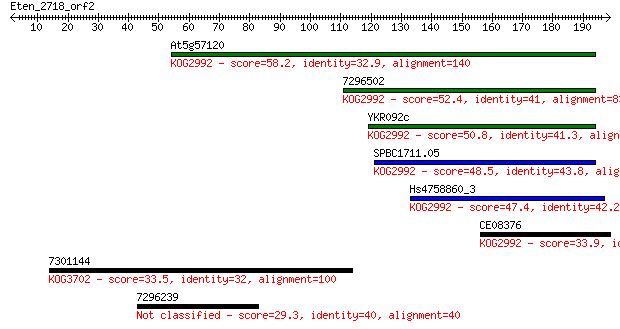
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_2718_orf2
Length=198
Score E
Sequences producing significant alignments: (Bits) Value
At5g57120 58.2 1e-08
7296502 52.4 6e-07
YKR092c 50.8 2e-06
SPBC1711.05 48.5 8e-06
Hs4758860_3 47.4 2e-05
CE08376 33.9 0.23
7301144 33.5 0.28
7296239 29.3 6.1
> At5g57120
Length=330
Score = 58.2 bits (139), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 46/146 (31%), Positives = 73/146 (50%), Gaps = 27/146 (18%)
Query 54 KKKARAIPENENSEAEATQAAPETDSSCSKQNGKTSEKKAKSEEETAAQEEPQDSQRGGK 113
+KK + EN+ E Q P ++ +K+NG + + KS +Q+ GK
Sbjct 199 RKKEENVVEND----EGVQETPVKETE-TKENGNAEKSETKS-----------TNQKSGK 242
Query 114 GAAGFKR----FQRIDDTRWISALPAELKDNSFWKKKN--DSFAAKAAEQLGRVRGKDFR 167
G + K FQR++ + + NS++ K + KA E LG+VRG+DFR
Sbjct 243 GLSNSKEPKKPFQRVN----VDEIVYTENSNSYYSKGGAEIGYGLKAQEVLGQVRGRDFR 298
Query 168 HEKSKKKRATWKGCGEIPMTVNSIQF 193
HEK+KKKR +++G G I +S +F
Sbjct 299 HEKTKKKRGSYRG-GLIDQESHSTKF 323
> 7296502
Length=654
Score = 52.4 bits (124), Expect = 6e-07, Method: Compositional matrix adjust.
Identities = 34/85 (40%), Positives = 50/85 (58%), Gaps = 7/85 (8%)
Query 111 GGKGAAGFKRFQRIDDTRWISALPAELKDNSFWKKKN--DSFAAKAAEQLGRVRGKDFRH 168
GG G R R +D + + ++D SF KKN S+ +A + L RGK F+H
Sbjct 574 GGSGRRSPFRRVRTEDV----VVDSRVQDMSFEAKKNAAGSWGERANKDLKHTRGKSFKH 629
Query 169 EKSKKKRATWKGCGEIPMTVNSIQF 193
EK+KKKR +++G G+I + VNSI+F
Sbjct 630 EKTKKKRGSYRG-GQIDVGVNSIKF 653
> YKR092c
Length=406
Score = 50.8 bits (120), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 31/75 (41%), Positives = 46/75 (61%), Gaps = 3/75 (4%)
Query 119 KRFQRIDDTRWISALPAELKDNSFWKKKNDSFAAKAAEQLGRVRGKDFRHEKSKKKRATW 178
K F R+D ++ I+ EL DN++ K ++ KA E+LGRVRGKDF K+K KR ++
Sbjct 333 KHFSRVDRSK-INFEAWELTDNTY-KGAAGTWGEKANEKLGRVRGKDFTKNKNKMKRGSY 390
Query 179 KGCGEIPMTVNSIQF 193
+G G I + S +F
Sbjct 391 RG-GSITLESGSYKF 404
> SPBC1711.05
Length=451
Score = 48.5 bits (114), Expect = 8e-06, Method: Compositional matrix adjust.
Identities = 32/74 (43%), Positives = 42/74 (56%), Gaps = 5/74 (6%)
Query 121 FQRIDD-TRWISALPAELKDNSFWKKKNDSFAAKAAEQLGRVRGKDFRHEKSKKKRATWK 179
F R+ D ++W A PA L+DNSF D + A L RGK FR EK+KKKR +++
Sbjct 382 FTRVGDPSQWDFASPA-LRDNSF--NFEDDYGTLANRDLIVTRGKGFRQEKNKKKRGSYR 438
Query 180 GCGEIPMTVNSIQF 193
G G I V S +F
Sbjct 439 G-GRINTEVRSFKF 451
> Hs4758860_3
Length=142
Score = 47.4 bits (111), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 27/66 (40%), Positives = 43/66 (65%), Gaps = 3/66 (4%)
Query 133 LPAELKDNSFWKKKNDS--FAAKAAEQLGRVRGKDFRHEKSKKKRATWKGCGEIPMTVNS 190
+ + + DNSF K+ + + +A + L +GK FRHEK+KKKR +++G G I + VNS
Sbjct 78 VDSRVADNSFDAKRGAAGDWGERANQVLKFTKGKSFRHEKTKKKRGSYRG-GSISVQVNS 136
Query 191 IQFASD 196
I+F S+
Sbjct 137 IKFDSE 142
> CE08376
Length=971
Score = 33.9 bits (76), Expect = 0.23, Method: Compositional matrix adjust.
Identities = 22/43 (51%), Positives = 28/43 (65%), Gaps = 2/43 (4%)
Query 156 EQLGRVRGKDFRHEKSKKKRATWKGCGEIPMTVNSIQFASDSE 198
E LG+V GK FRHE KK+ G G I ++NSI+F SDS+
Sbjct 930 ESLGKVVGKAFRHE-KTKKKKGSYGGGPINQSINSIKF-SDSD 970
> 7301144
Length=1004
Score = 33.5 bits (75), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 32/104 (30%), Positives = 52/104 (50%), Gaps = 4/104 (3%)
Query 14 DLLQLCSAAARASEEAEE---AVEKPSKKSKKKKINDEESTKPKKKARAIPENENSEAEA 70
DLL L + +E EE +E P +K+K I + P+K+ A PE ++ + EA
Sbjct 239 DLLNLSAREESVDQEPEEDPPKLEAPPRKAKSPIIFNRREKTPEKRPEAEPEIDSRQEEA 298
Query 71 TQAAPETDSSCSKQNGKTSEKKAKSEEETAAQEEPQD-SQRGGK 113
+ +P+ DSS K SE+++ + + + P D SQR K
Sbjct 299 EEKSPKKDSSADKAKEDRSERRSVHDRLGSKAQTPADRSQRNQK 342
> 7296239
Length=2151
Score = 29.3 bits (64), Expect = 6.1, Method: Compositional matrix adjust.
Identities = 16/40 (40%), Positives = 26/40 (65%), Gaps = 5/40 (12%)
Query 43 KKINDEESTKPKKKARAIPENENSEAEATQAAPETDSSCS 82
KK ++E+ + P A+P+ + SE EA +AAP T+ +CS
Sbjct 426 KKFSEEKPSSP-----AVPDGKVSEMEADEAAPNTEENCS 460
Lambda K H
0.303 0.118 0.326
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3407623970
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40