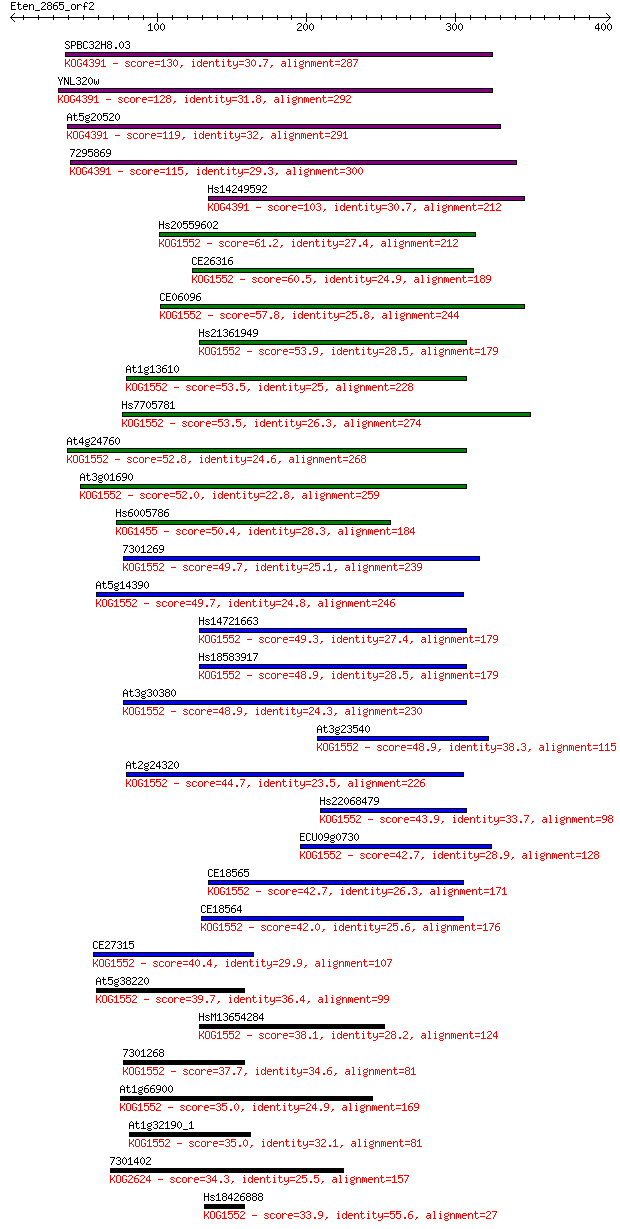
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_2865_orf2
Length=403
Score E
Sequences producing significant alignments: (Bits) Value
SPBC32H8.03 130 6e-30
YNL320w 128 3e-29
At5g20520 119 1e-26
7295869 115 1e-25
Hs14249592 103 9e-22
Hs20559602 61.2 4e-09
CE26316 60.5 7e-09
CE06096 57.8 4e-08
Hs21361949 53.9 6e-07
At1g13610 53.5 8e-07
Hs7705781 53.5 9e-07
At4g24760 52.8 1e-06
At3g01690 52.0 2e-06
Hs6005786 50.4 7e-06
7301269 49.7 1e-05
At5g14390 49.7 1e-05
Hs14721663 49.3 2e-05
Hs18583917 48.9 2e-05
At3g30380 48.9 2e-05
At3g23540 48.9 2e-05
At2g24320 44.7 4e-04
Hs22068479 43.9 6e-04
ECU09g0730 42.7 0.001
CE18565 42.7 0.002
CE18564 42.0 0.002
CE27315 40.4 0.008
At5g38220 39.7 0.013
HsM13654284 38.1 0.034
7301268 37.7 0.040
At1g66900 35.0 0.28
At1g32190_1 35.0 0.30
7301402 34.3 0.46
Hs18426888 33.9 0.62
> SPBC32H8.03
Length=299
Score = 130 bits (326), Expect = 6e-30, Method: Compositional matrix adjust.
Identities = 88/295 (29%), Positives = 145/295 (49%), Gaps = 59/295 (20%)
Query 38 FSFQEKLVFDPSYPIEPAEIKKFPHIIGSPASYGIPYDDLFLTTKDGVRIHGWFLKQTEN 97
+ +Q+ LV+ ++P E + +P + + Y+ + L T+D V + + + Q+E+
Sbjct 32 YKYQKTLVYPSAFPQGSREN------VPTPKEFNMEYERIELRTRDKVTLDSYLMLQSES 85
Query 98 -EKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDYRGYGRSEGTPSEAGVYMDAD 155
E PT + FH N G++G LP AR Y MNV +I YRGYG+S G+PSEAG+ +D+
Sbjct 86 PESRPTLLYFHANAGNMGHRLPIARVFYSALNMNVFIISYRGYGKSTGSPSEAGLKIDSQ 145
Query 156 AALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVMGHS 215
AL++++ + I + K + V G S
Sbjct 146 TALEYLM---------------------EHPICSKTK----------------IVVYGQS 168
Query 216 MGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAVFRLFRWLVKAIQRI------S 269
+GGAVAI L + + ++ L++ENTFTS+++ + +F + I R S
Sbjct 169 IGGAVAIALTAKNQDRISALILENTFTSIKD------MIPTVFPYGGSIISRFCTEIWSS 222
Query 270 MNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKFLVEVPEGDHEET 324
+ I K+ +LPVLFL G +DE + P L+ CGS++K P+ H +T
Sbjct 223 QDEIRKIK--KLPVLFLSGEKDEIVPPPQMVLLFGLCGSAKKKFHSFPKCTHNDT 275
> YNL320w
Length=284
Score = 128 bits (321), Expect = 3e-29, Method: Compositional matrix adjust.
Identities = 93/295 (31%), Positives = 142/295 (48%), Gaps = 53/295 (17%)
Query 33 SIDSEFSFQEKLVFDPSYPIEPAEIKKFPHIIGSPASYGIPYDDLFLTTKDGVRIHGWFL 92
S+ + + +Q +LV+ P+ + + + +P S GIPY+ L L T+D +++ W +
Sbjct 20 SVATLYHYQNRLVY-------PSWAQGARNHVDTPDSRGIPYEKLTLITQDHIKLEAWDI 72
Query 93 KQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDYRGYGRSEGTPSEAGVY 151
K EN + IL N G++G + Y Q GM+V + YRGYG SEG+PSE G+
Sbjct 73 KN-ENSTSTVLILC-PNAGNIGYFILIIDIFYRQFGMSVFIYSYRGYGNSEGSPSEKGLK 130
Query 152 MDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFV 211
+DAD + S S+ S SK V +
Sbjct 131 LDADCVI------------------------------------SHLSTDSFHSKRKLV-L 153
Query 212 MGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAVFRLFRWLVKAIQRISMN 271
G S+GGA A+ +A + + G+++ENTF S+R+ + + + F L I N
Sbjct 154 YGRSLGGANALYIASKFRDLCDGVILENTFLSIRKVIPYIFPLLKRFTLLCHEIW----N 209
Query 272 NISKVGSL--ELPVLFLCGLRDENIKPRHSSRLYEACGSSQKFLVEVPEGDHEET 324
+ +GS E P LFL GL+DE + P H +LYE C SS K + E P G H +T
Sbjct 210 SEGLMGSCSSETPFLFLSGLKDEIVPPFHMRKLYETCPSSNKKIFEFPLGSHNDT 264
> At5g20520
Length=340
Score = 119 bits (298), Expect = 1e-26, Method: Compositional matrix adjust.
Identities = 93/333 (27%), Positives = 141/333 (42%), Gaps = 89/333 (26%)
Query 39 SFQEKLVFDPSYPIEPAEIKKFPHIIGSPASYGIPYDDLFLTTKDGVRIHGWFLKQTENE 98
+FQEKLV+ P+ P K +P +PA + Y+D++L + DGVR+H WF+K
Sbjct 25 AFQEKLVY---VPVLPGLSKSYPI---TPARLNLIYEDIWLQSSDGVRLHAWFIKMFPEC 78
Query 99 KAP----TFILFHGNYGHVGLTLPRARW----LYDQGM---------------------N 129
+ P +F + + ++ W YD + N
Sbjct 79 REPFLGFCLFIFVCSKEPILCSM-SVSWDTLSCYDLDLRIPDIAHRLEMVRIMIQKLKCN 137
Query 130 VLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISK 189
V ++ YRGYG SEG PS+ G+ DA AALD + G+ +S
Sbjct 138 VFMLSYRGYGASEGYPSQQGIIKDAQAALDHLSGRTDIDTSR------------------ 179
Query 190 QQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAE 249
+ V G S+GGAV L K ++++ L++ENTFTS+ + A
Sbjct 180 -------------------IVVFGRSLGGAVGAVLTKNNPDKVSALILENTFTSILDMAG 220
Query 250 DTYAVFRLFRW-----------LVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRH 298
+ +W L+ + R I + ++ PVLFL GL+DE + P H
Sbjct 221 ---VLLPFLKWFIGGSGTKSLKLLNFVVRSPWKTIDAIAEIKQPVLFLSGLQDEMVPPFH 277
Query 299 SSRLY--EACGSSQKFLVEVPEGDHEETFEIGG 329
LY A + Q VE P G H +T+ GG
Sbjct 278 MKMLYAKAAARNPQCTFVEFPSGMHMDTWLSGG 310
> 7295869
Length=338
Score = 115 bits (289), Expect = 1e-25, Method: Compositional matrix adjust.
Identities = 88/305 (28%), Positives = 148/305 (48%), Gaps = 52/305 (17%)
Query 41 QEKLVFDPSYPIEPAEIKKFPHIIGSPASYGIPYDDLFLTTKDGVRIHGWFLKQTE--NE 98
Q+ L++ P P A + + I P + +P+ + + T D V +H +++ Q E ++
Sbjct 53 QDLLLYHPDLP---ANSRIY---IPIPTMHNLPHITVSIKTPDDVTLHAFWVTQPEERSK 106
Query 99 KAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDYRGYGRSEGTPSEAGVYMDADAA 157
+PT + FHGN G++G + +Y NVL+++YRGYG S G P+E G+ DA AA
Sbjct 107 SSPTLLYFHGNAGNMGHRMQNVWGIYHHLHCNVLMVEYRGYGLSTGVPTERGLVTDARAA 166
Query 158 LDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVMGHSMG 217
+D++ + S + + G S+G
Sbjct 167 IDYLHTRHDLDHSQ-------------------------------------LILFGRSLG 189
Query 218 GAVAIDLAKRR--GNELAGLVVENTFTSLREAAEDTYAVFRLFRWLVKAIQRISMNNISK 275
GAV +D+A G +L +VENTF+S+ E A + V +++ + + +++SK
Sbjct 190 GAVVVDVAADTVYGQKLMCAIVENTFSSIPEMAVEL--VHPAVKYIPNLLFKNKYHSMSK 247
Query 276 VGSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKFLVEVPEGDHEETFEIGGLETKRA 335
+G +P LF+ GL D + PR LY CGS K L+E P G H +T+ + G +A
Sbjct 248 IGKCSVPFLFISGLADNLVPPRMMRALYTKCGSEIKRLLEFPGGSHNDTWIVDG--YYQA 305
Query 336 LGDFV 340
+G F+
Sbjct 306 IGGFL 310
> Hs14249592
Length=201
Score = 103 bits (256), Expect = 9e-22, Method: Compositional matrix adjust.
Identities = 65/212 (30%), Positives = 100/212 (47%), Gaps = 40/212 (18%)
Query 134 DYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKG 193
DYRGYG+SEG SE G+Y+D++A LD+V+ + +
Sbjct 13 DYRGYGKSEGEASEEGLYLDSEAVLDYVMTRPDLDKTK---------------------- 50
Query 194 KSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYA 253
+F+ G S+GGAVAI LA + ++ ++VENTF S+ A ++
Sbjct 51 ---------------IFLFGRSLGGAVAIHLASENSHRISAIMVENTFLSIPHMASTLFS 95
Query 254 VFRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKFL 313
F + R+L + + K+ +P LF+ GL D+ I P +LYE S K L
Sbjct 96 FFPM-RYLPLWCYKNKFLSYRKISQCRMPSLFISGLSDQLIPPVMMKQLYELSPSRTKRL 154
Query 314 VEVPEGDHEETFEIGGLETKRALGDFVKLGIK 345
P+G H +T++ G T AL F+K +K
Sbjct 155 AIFPDGTHNDTWQCQGYFT--ALEQFIKEVVK 184
> Hs20559602
Length=404
Score = 61.2 bits (147), Expect = 4e-09, Method: Compositional matrix adjust.
Identities = 58/225 (25%), Positives = 87/225 (38%), Gaps = 52/225 (23%)
Query 101 PTFILFHGNYGHVG--LTLPRARWLYDQGMNVLVIDYRGYGRSEGTPSEAGVYMDADAAL 158
P + HGN G G + + L G +V+ DYRG+G S GTPSE G+ DA
Sbjct 169 PIILYLHGNAGTRGGDHRVELYKVLSSLGYHVVTFDYRGWGDSVGTPSERGMTYDALHVF 228
Query 159 DFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVMGHSMGG 218
D++ + S N V++ GHS+G
Sbjct 229 DWI---------------------------------------KARSGDNPVYIWGHSLGT 249
Query 219 AVAIDLAKR---RGNELAGLVVENTFTSLREAAED-----TYAVFRLFRW-LVKAIQR-- 267
VA +L +R R L++E+ FT++RE A+ Y F F W + I
Sbjct 250 GVATNLVRRLCERETPPDALILESPFTNIREEAKSHPFSVIYRYFPGFDWFFLDPITSSG 309
Query 268 ISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKF 312
I N V + P+L L D + + +LY ++ F
Sbjct 310 IKFANDENVKHISCPLLILHAEDDPVVPFQLGRKLYSIAAPARSF 354
> CE26316
Length=345
Score = 60.5 bits (145), Expect = 7e-09, Method: Compositional matrix adjust.
Identities = 47/203 (23%), Positives = 86/203 (42%), Gaps = 54/203 (26%)
Query 123 LYDQGMNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNN 182
L D +V+ DYRGYG SEGTP+E G+ D +++
Sbjct 138 LSDCNYHVVCFDYRGYGDSEGTPTEKGIVEDTKTVYEWL--------------------- 176
Query 183 NANAISKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAK---RRGNELAGLVVEN 239
K+ GK+ V V GHSMG V+ L + R GL++E+
Sbjct 177 ------KENCGKTP------------VIVWGHSMGTGVSCKLVQDLSREQQPPCGLILES 218
Query 240 TFTSLREAAEDTYAVFRLFRWL---------VKAIQRI--SMNNISKVGSLELPVLFLCG 288
F +L++A + + +F +F W+ ++ + + +M + ++ + P++ L
Sbjct 219 PFNNLKDAVTN-HPIFTVFSWMNDFMVDHIIIRPLNSVGLTMRSDKRIRLVSCPIIILHA 277
Query 289 LRDENIKPRHSSRLYEACGSSQK 311
D+ + + LYEA +++
Sbjct 278 EDDKILPVKLGRALYEAAKDAER 300
> CE06096
Length=405
Score = 57.8 bits (138), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 63/249 (25%), Positives = 93/249 (37%), Gaps = 57/249 (22%)
Query 102 TFILFHGNYGHVGLTLPRARWLYDQG----MNVLVIDYRGYGRSEGTPSEAGVYMDADAA 157
T + HGN +G +LY G NV DY GYG S G PSE +Y D AA
Sbjct 187 TLLFSHGNAVDLGQ---MTSFLYGLGFHLNCNVFSYDYSGYGCSTGKPSEKNLYADITAA 243
Query 158 LDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVMGHSMG 217
+ + + + K++ + + G S+G
Sbjct 244 FELL--------------------KSEFGVPKEK-----------------IILYGQSIG 266
Query 218 GAVAIDLAKRRGNELAGLVVENTFTS-LREAAEDTYAVFRLFRWLVKAIQRISMNNISKV 276
++DLA R +LA LV+ + S +R A T W A +I KV
Sbjct 267 TVPSVDLASRE--DLAALVLHSPLMSGMRVAFPGTTTT-----WCCDAFP-----SIEKV 314
Query 277 GSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKFLVEVPEGDHEETFEIGGLETKRAL 336
++ P L + G DE I H +YE C +S + L G ++ LE R+
Sbjct 315 PRVKCPTLVIHGTDDEVIDFSHGVSIYERCPTSVEPLWVPGAGHNDVELHAAYLERLRSF 374
Query 337 GDFVKLGIK 345
D I+
Sbjct 375 IDMEASAIR 383
> Hs21361949
Length=361
Score = 53.9 bits (128), Expect = 6e-07, Method: Compositional matrix adjust.
Identities = 51/180 (28%), Positives = 68/180 (37%), Gaps = 50/180 (27%)
Query 128 MNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAI 187
N+ DY GYG S G PSE +Y D DAA +
Sbjct 191 CNIFSYDYSGYGASSGRPSERNLYADIDAAWQAL-------------------------- 224
Query 188 SKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LRE 246
+ + G S S + + G S+G +DLA R E A +V+ + TS +R
Sbjct 225 -RTRYGISPDS----------IILYGQSIGTVPTVDLASR--YECAAVVLHSPLTSGMRV 271
Query 247 AAEDTYAVFRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
A DT + + NI KV + PVL + G DE I H LYE C
Sbjct 272 AFPDTKKTYCF----------DAFPNIEKVSKITSPVLIIHGTEDEVIDFSHGLALYERC 321
> At1g13610
Length=351
Score = 53.5 bits (127), Expect = 8e-07, Method: Compositional matrix adjust.
Identities = 57/229 (24%), Positives = 88/229 (38%), Gaps = 52/229 (22%)
Query 79 LTTKDGVRIHGWFLKQTENEKAPTFILF-HGNYGHVGLTLPRARWLYDQGMNVLVIDYRG 137
+ TK G I ++K N A +LF HGN + L +N++ DY G
Sbjct 46 IRTKRGNEIVAMYVK---NPTAKLTVLFSHGNASDLAQIFYILAELIQLNVNLMGYDYSG 102
Query 138 YGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSS 197
YG+S G PSE Y D +AA +++ +Q G
Sbjct 103 YGQSSGKPSEQDTYADIEAAYNWL---------------------------RQTYG---- 131
Query 198 SSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAVFRL 257
+K + + G S+G +++LA R L LV+ + F S Y V
Sbjct 132 ------TKDERIILYGQSVGSGPSLELASRLP-RLRALVLHSPFLS---GLRVMYPVKHS 181
Query 258 FRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
F + + NI K+ +E PVL + G D+ + H L+ C
Sbjct 182 FPFDI-------YKNIDKIHLVECPVLVIHGTDDDVVNISHGKHLWGLC 223
> Hs7705781
Length=293
Score = 53.5 bits (127), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 72/277 (25%), Positives = 105/277 (37%), Gaps = 56/277 (20%)
Query 76 DLFLT-TKDGVRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVI 133
+ F+T T G RI F++ + N K T + HGN +G L + N+
Sbjct 67 ECFMTRTSKGNRIACMFVRCSPNAKY-TLLFSHGNAVDLGQMSSFYIGLGSRINCNIFSY 125
Query 134 DYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKG 193
DY GYG S G P+E +Y D +AA L + N
Sbjct 126 DYSGYGASSGKPTEKNLYADIEAAW---LALRTRYGIRPEN------------------- 163
Query 194 KSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LREAAEDTY 252
V + G S+G ++DLA R E A +++ + TS +R A DT
Sbjct 164 ---------------VIIYGQSIGTVPSVDLAARY--ESAAVILHSPLTSGMRVAFPDTK 206
Query 253 AVFRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKF 312
+ + NI K+ + PVL + G DE I H L+E C +
Sbjct 207 ETYCF----------DAFPNIDKISKITSPVLIIHGTEDEVIDFSHGLALFERCQRPVEP 256
Query 313 LVEVPEGDHEETFEIGGLETKRALGDFVKLGIKEKKK 349
L EG E+ G +R L FV + +K K
Sbjct 257 LWV--EGAGHNDVELYGQYLER-LKQFVSQELVQKHK 290
> At4g24760
Length=365
Score = 52.8 bits (125), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 66/274 (24%), Positives = 101/274 (36%), Gaps = 59/274 (21%)
Query 39 SFQEKLVF----DPSYPIEPAEIKKFPHIIGSPASYGIPYDDLFLTTKDGVRIHGWFLKQ 94
S KL F PSY + E + + SP + D L L T+ G I +++
Sbjct 7 SMAAKLAFFPPNPPSYKLVRDETTEL--FLMSPFPHRENVDILRLPTRRGTEIVAMYIRY 64
Query 95 TENEKAPTFILF-HGNYGHVGLTLPRARWL-YDQGMNVLVIDYRGYGRSEGTPSEAGVYM 152
A T +L+ HGN +G L +N++ DY GYG+S G P+E Y
Sbjct 65 P---MAVTTLLYSHGNAADIGQMYELFIELSIHLRVNLMGYDYSGYGQSSGKPTEQNTYA 121
Query 153 DADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVM 212
D +AA + N A N + +
Sbjct 122 DIEAAYKCL------------EENYGAKQEN-------------------------IILY 144
Query 213 GHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAVFRLFRWLVKAIQRISMNN 272
G S+G +DLA R A ++ + LR Y V R + + + N
Sbjct 145 GQSVGSGPTVDLAARLPRLRASILHSPILSGLRV----MYPVKRTYWFDI-------YKN 193
Query 273 ISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
I K+ + PVL + G D+ + H +L+E C
Sbjct 194 IDKITLVRCPVLVIHGTADDVVDFSHGKQLWELC 227
> At3g01690
Length=361
Score = 52.0 bits (123), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 59/260 (22%), Positives = 97/260 (37%), Gaps = 53/260 (20%)
Query 48 PSYPIEPAEIKKFPHIIGSPASYGIPYDDLFLTTKDGVRIHGWFLKQTENEKAPTFILFH 107
PSY + E+ ++ SP + + + L T+ G I G +++ T + H
Sbjct 20 PSYKVVTDELTGL--LLLSPFPHRENVEIVKLRTRRGTEIVGMYVRHPM--ATSTLLYSH 75
Query 108 GNYGHVGLTLPRARWL-YDQGMNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQ 166
GN +G L +N++ DY GYG+S G PSE Y D +A +
Sbjct 76 GNAADLGQMYELFIELSIHLKVNLMGYDYSGYGQSTGKPSEHNTYADIEAVYKCL----- 130
Query 167 SSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAK 226
+ SK V + G S+G +DLA
Sbjct 131 --------------------------------EETFGSKQEGVILYGQSVGSGPTLDLAS 158
Query 227 RRGNELAGLVVENTFTSLREAAEDTYAVFRLFRWLVKAIQRISMNNISKVGSLELPVLFL 286
R A ++ + LR Y+V + + + + NI K+ ++ PVL +
Sbjct 159 RLPQLRAVVLHSPILSGLRV----MYSVKKTYWFDI-------YKNIDKIPYVDCPVLII 207
Query 287 CGLRDENIKPRHSSRLYEAC 306
G DE + H +L+E C
Sbjct 208 HGTSDEVVDCSHGKQLWELC 227
> Hs6005786
Length=313
Score = 50.4 bits (119), Expect = 7e-06, Method: Compositional matrix adjust.
Identities = 52/185 (28%), Positives = 72/185 (38%), Gaps = 37/185 (20%)
Query 72 IPYDDL-FLTTKDGVRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQGMNV 130
IPY DL L DG + + K T KA F+ HG H G AR L + V
Sbjct 24 IPYQDLPHLVNADGQYLFCRYWKPTGTPKALIFV-SHGAGEHSGRYEELARMLMGLDLLV 82
Query 131 LVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQ 190
D+ G+G+SEG E V D + VL + +++ K
Sbjct 83 FAHDHVGHGQSEG---ERMVVSDFHVFVRDVL-------------------QHVDSMQKD 120
Query 191 QKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAED 250
G VF++GHSMGGA+AI A R AG+V+ + +
Sbjct 121 YPGLP-------------VFLLGHSMGGAIAILTAAERPGHFAGMVLISPLVLANPESAT 167
Query 251 TYAVF 255
T+ V
Sbjct 168 TFKVL 172
> 7301269
Length=286
Score = 49.7 bits (117), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 60/242 (24%), Positives = 91/242 (37%), Gaps = 53/242 (21%)
Query 77 LFLTTKDGVRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDY 135
F T G I +++ ++N K T + HGN +G L Q N+ DY
Sbjct 68 FFTRTSRGNLITCIYVRCSKNAKY-TLLFSHGNAVDLGQMSSFYLTLGSQINCNIFGYDY 126
Query 136 RGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKS 195
GYG S G PSE +Y D +AA + + S +
Sbjct 127 SGYGMSGGKPSEKNLYADIEAAWQAMRTRFNISPET------------------------ 162
Query 196 SSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LREAAEDTYAV 254
+ + G S+G +DLA R +E+ +++ + S LR +T
Sbjct 163 -------------IILYGQSIGTVPTVDLASR--HEVGAVILHSPLMSGLRVVFRNTKRT 207
Query 255 FRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC-GSSQKFL 313
W A +I KV ++ PVL + G DE I H +YE C + + F
Sbjct 208 -----WFFDAFP-----SIDKVAKVKAPVLVIHGTDDEVIDFSHGIGIYERCPKTVEPFW 257
Query 314 VE 315
VE
Sbjct 258 VE 259
> At5g14390
Length=369
Score = 49.7 bits (117), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 61/267 (22%), Positives = 94/267 (35%), Gaps = 73/267 (27%)
Query 59 KFPHIIGSPASYGIPYDDLF------------------LTTKDGVRIHGWFLKQTENEKA 100
KF SP+SY + YD+L L T+ G I +++
Sbjct 11 KFAFFPPSPSSYKLVYDELTGLLLMNPFPHRENVEILKLPTRRGTEIVAMYVRHPM--AT 68
Query 101 PTFILFHGNYGHVGLTLPRARWL-YDQGMNVLVIDYRGYGRSEGTPSEAGVYMDADAALD 159
T + HGN +G L +N++ DY GYG+S G PSE Y D +AA
Sbjct 69 STLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDYSGYGQSTGKPSEHHTYADIEAAYK 128
Query 160 FVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGA 219
+ + +K + + G S+G
Sbjct 129 CL-------------------------------------EETYGAKQEDIILYGQSVGSG 151
Query 220 VAIDLAKRRGNELAGLVVENTFTSLREA--AEDTYAVFRLFRWLVKAIQRISMNNISKVG 277
+DLA R A ++ + LR + TY F +F+ NI K+
Sbjct 152 PTLDLAARLPQLRAAVLHSPILSGLRVMYPVKKTYW-FDIFK------------NIDKIP 198
Query 278 SLELPVLFLCGLRDENIKPRHSSRLYE 304
+ PVL + G DE + H +L+E
Sbjct 199 LVNCPVLVIHGTCDEVVDCSHGKQLWE 225
> Hs14721663
Length=310
Score = 49.3 bits (116), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 49/180 (27%), Positives = 66/180 (36%), Gaps = 50/180 (27%)
Query 128 MNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAI 187
N+ D GYG S G PSE +Y D DA +
Sbjct 140 CNIFTYDSSGYGASSGRPSERNLYADIDATWQAL-------------------------- 173
Query 188 SKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LRE 246
+ + G S S + + G S+G +DLA R E A +V+ + TS +R
Sbjct 174 -RTRYGISPDS----------IILYGQSIGTVPTMDLASR--YECAAVVLHSPLTSGMRV 220
Query 247 AAEDTYAVFRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
A DT + + NI KV + PVL + G DE I H LYE C
Sbjct 221 AFRDTKKTYCF----------DAFPNIEKVSKITSPVLIIHGREDEVIDFSHGLALYERC 270
> Hs18583917
Length=329
Score = 48.9 bits (115), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 51/180 (28%), Positives = 70/180 (38%), Gaps = 50/180 (27%)
Query 128 MNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAI 187
N+ DY GYG S G PSE +Y D DAA +
Sbjct 161 CNIFSYDYSGYGVSSGKPSEKNLYADIDAAWQAL-------------------------- 194
Query 188 SKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LRE 246
+ + G S + + + G S+G +DLA R E A +++ + S LR
Sbjct 195 -RTRYGVSPEN----------IILYGQSIGTVPTVDLASRY--ECAAVILHSPLMSGLRV 241
Query 247 AAEDTYAVFRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
A DT + A S++ ISKV S PVL + G DE I H +YE C
Sbjct 242 AFPDTRKTY-----CFDAFP--SIDKISKVTS---PVLVIHGTEDEVIDFSHGLAMYERC 291
> At3g30380
Length=399
Score = 48.9 bits (115), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 56/232 (24%), Positives = 87/232 (37%), Gaps = 53/232 (22%)
Query 77 LFLTTKDGVRIHGWFLKQTENEKAPTFILF-HGNYGHVGLTLPRARWL-YDQGMNVLVID 134
L L TK G ++ ++K N A +L+ HGN +G L +N++ D
Sbjct 46 LKLKTKRGNQVVAAYIK---NPTASLTLLYSHGNAADLGQMFELFSELSLHLRVNLIGYD 102
Query 135 YRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGK 194
Y GYGRS G PSE Y D +A + K
Sbjct 103 YSGYGRSSGKPSEQNTYSDIEAVYRCLEEKY----------------------------- 133
Query 195 SSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAV 254
K V + G S+G ++LA R N L +V+ + S Y V
Sbjct 134 --------GVKEQDVILYGQSVGSGPTLELASRLPN-LRAVVLHSAIAS---GLRVMYPV 181
Query 255 FRLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
R + + + N+ K+ ++ PVL + G D+ + H +L+E C
Sbjct 182 KRTYWFDI-------YKNVEKISFVKCPVLVIHGTSDDVVNWSHGKQLFELC 226
> At3g23540
Length=568
Score = 48.9 bits (115), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 44/138 (31%), Positives = 70/138 (50%), Gaps = 28/138 (20%)
Query 207 NFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAE---DTYAVFRLFRWLVK 263
+ + + G SMG ++ + +AG+++++ F+ L + DTY FRL ++ VK
Sbjct 132 SLIGLWGRSMGAVTSL-MYGVEDPSIAGMILDSPFSDLVDLMMELVDTYK-FRLPKFTVK 189
Query 264 --------AIQR------ISMNNISKVGSLEL------PVLFLCGLRDENIKPRHSSRLY 303
AIQ+ + +N I KVG L + PVLF L D+ I+P HS R+Y
Sbjct 190 FAIQFMRRAIQKKAKFDIMELNTI-KVGPLPVAKASFVPVLFGHALDDDFIRPHHSDRIY 248
Query 304 EACGSSQKFLVEVPEGDH 321
EA K +++ P GDH
Sbjct 249 EAY-VGDKNIIKFP-GDH 264
> At2g24320
Length=316
Score = 44.7 bits (104), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 53/240 (22%), Positives = 86/240 (35%), Gaps = 54/240 (22%)
Query 79 LTTKDGVRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDYRG 137
LTTK G ++ F K + T + HGN +G + L +N++ DY G
Sbjct 48 LTTKSGNKVIATFWKHPFSRF--TLLYSHGNAADLGQMVDLFIELRAHLRVNIMSYDYSG 105
Query 138 YGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKGKSSS 197
YG S G P+E Y D +A + + I +++
Sbjct 106 YGASTGKPTELNTYYDIEAVYNCL--------------------RTEYGIMQEE------ 139
Query 198 SSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAVFRL 257
+ + G S+G + LA R L G+V+ + S F
Sbjct 140 -----------MILYGQSVGSGPTLHLASRV-KRLRGIVLHSAILSGLRVLYPVKMTFWF 187
Query 258 FRWLVKAIQRIS-------------MNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYE 304
+ V I +S + NI K+ + PVL + G +D+ + H RL+E
Sbjct 188 DMYKVSLISLVSGYYYRVSLSNSGILQNIDKIRHVTCPVLVIHGTKDDIVNMSHGKRLWE 247
> Hs22068479
Length=161
Score = 43.9 bits (102), Expect = 6e-04, Method: Compositional matrix adjust.
Identities = 33/99 (33%), Positives = 45/99 (45%), Gaps = 14/99 (14%)
Query 209 VFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LREAAEDTYAVFRLFRWLVKAIQR 267
+ + G S+G +DLA R E A +V+ + TS +R A DT K
Sbjct 36 IILYGQSIGTVPTVDLASR--YECAAVVLHSPLTSGMRVAFPDT-----------KTYCF 82
Query 268 ISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEAC 306
+ NI KV + PVL + G+ DE I H LYE C
Sbjct 83 DAFPNIEKVSKITSPVLIIHGMEDEVIDFSHGLALYERC 121
> ECU09g0730
Length=326
Score = 42.7 bits (99), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 37/128 (28%), Positives = 61/128 (47%), Gaps = 9/128 (7%)
Query 196 SSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYAVF 255
S++S S +S V+G S+G AV I LA + LV+ N F SLRE + +
Sbjct 181 STASEWLSRRSTRKVVVGFSIGAAVGIRLAGM--CRVDALVLVNPFISLREVVS-SIPLG 237
Query 256 RLFRWLVKAIQRISMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYEACGSSQKFLVE 315
R+ + LV +N++ V +++PV F+ DE + H+ L + +K ++
Sbjct 238 RVLKHLVVD----EWSNMNGVKDIDVPVYFVVSSDDEIVPESHADALIKRTRHPRKIVIR 293
Query 316 VPEGDHEE 323
DH E
Sbjct 294 --NADHNE 299
> CE18565
Length=305
Score = 42.7 bits (99), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 45/174 (25%), Positives = 60/174 (34%), Gaps = 52/174 (29%)
Query 134 DYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKG 193
DY GYG S GT E VY D A + +L +
Sbjct 116 DYSGYGFSSGTQGEKNVYADVRAVYEKILEMRPDKK------------------------ 151
Query 194 KSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAAEDTYA 253
+ VMG+S+G A+DLA + LAG+V+ FTS
Sbjct 152 ---------------IVVMGYSIGTTAAVDLAATNPDRLAGVVLIAPFTS---------- 186
Query 254 VFRLFRWLVKAIQRI---SMNNISKVGSLELPVLFLCGLRDENIKPRHSSRLYE 304
RLF S + K+ +++ VL G DE I H LYE
Sbjct 187 GLRLFSSKPDKPDTCWADSFKSFDKINNIDTRVLICHGDVDEVIPLSHGLALYE 240
> CE18564
Length=335
Score = 42.0 bits (97), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 45/183 (24%), Positives = 62/183 (33%), Gaps = 67/183 (36%)
Query 129 NVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAIS 188
+V DY GYG S GT SE +Y D A + +L
Sbjct 154 DVYAFDYSGYGFSSGTQSEKNMYADVRAVYEHIL-------------------------- 187
Query 189 KQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTSLREAA 248
K + K + V+G+S+G A+DLA + L G+V+ TS
Sbjct 188 KTRPDKK-------------IVVIGYSIGTTAAVDLAASNPDRLVGVVLIAPLTS----- 229
Query 249 EDTYAVFRLFRWLVKAIQRISMNN-------ISKVGSLELPVLFLCGLRDENIKPRHSSR 301
R+F NN I K+ + VL G D+ I H
Sbjct 230 -----ALRMF-----------CNNPDKETTCIDKICHINTRVLICHGDHDQRIPMTHGMA 273
Query 302 LYE 304
LYE
Sbjct 274 LYE 276
> CE27315
Length=490
Score = 40.4 bits (93), Expect = 0.008, Method: Compositional matrix adjust.
Identities = 32/120 (26%), Positives = 53/120 (44%), Gaps = 17/120 (14%)
Query 57 IKKFPHIIGSPASYGIPYDD------LFLTTKDGVRIHGWFL--KQTENEKAPTFILF-H 107
I F H++ + +PYD+ + L + +R W K + ++P I+F
Sbjct 191 IDVFYHLLRCKV-FTLPYDEKHKICAMELYCEQSIR---WLHRDKNRQRLRSPNLIIFSQ 246
Query 108 GNYGHVGLTLPRARWLYDQG----MNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLG 163
N +G L D ++L+ DY GYG SEGT +E VY +A + + +G
Sbjct 247 PNSSDLGCCLMMDPNFADIADFLQCDLLIYDYPGYGVSEGTTNEKNVYAAVEAVMKYAMG 306
> At5g38220
Length=320
Score = 39.7 bits (91), Expect = 0.013, Method: Compositional matrix adjust.
Identities = 36/116 (31%), Positives = 49/116 (42%), Gaps = 19/116 (16%)
Query 59 KFPHIIGSPASYG------------IPYDD----LFLTTKDGVRIHGWFLKQTENEKAPT 102
KF SP SYG +P D L L T+ G I ++K + T
Sbjct 11 KFAFFPPSPPSYGFVSDVDRLYITEVPRRDDVDVLKLKTRRGNEIVAIYIKHPKANG--T 68
Query 103 FILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDYRGYGRSEGTPSEAGVYMDADAA 157
+ HGN +G L ++ +N++ DY GYG+S G SE Y D DAA
Sbjct 69 LLYSHGNAADLGQMFELFIELSNRLRLNLMGYDYSGYGQSTGKASECNTYADIDAA 124
> HsM13654284
Length=236
Score = 38.1 bits (87), Expect = 0.034, Method: Compositional matrix adjust.
Identities = 35/125 (28%), Positives = 48/125 (38%), Gaps = 40/125 (32%)
Query 128 MNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAI 187
N+ DY GYG S G PSE +Y D DAA +
Sbjct 140 CNIFSYDYSGYGASSGRPSERNLYADIDAAWQAL-------------------------- 173
Query 188 SKQQKGKSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS-LRE 246
+ + G S S + + G S+G +DLA R E A +V+ + TS +R
Sbjct 174 -RTRYGISPDS----------IILYGQSIGTVPTVDLASR--YECAAVVLHSPLTSGMRV 220
Query 247 AAEDT 251
A DT
Sbjct 221 AFPDT 225
> 7301268
Length=244
Score = 37.7 bits (86), Expect = 0.040, Method: Compositional matrix adjust.
Identities = 28/82 (34%), Positives = 38/82 (46%), Gaps = 2/82 (2%)
Query 77 LFLTTKDGVRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVIDY 135
F T G I +++ ++N K T + HGN +G L Q N+ DY
Sbjct 68 FFTRTSRGNLITCIYVRCSKNAKY-TLLFSHGNAVDLGQMSSFYLTLGSQINCNIFGYDY 126
Query 136 RGYGRSEGTPSEAGVYMDADAA 157
GYG S G PSE +Y D +AA
Sbjct 127 SGYGMSGGKPSEKNLYADIEAA 148
> At1g66900
Length=256
Score = 35.0 bits (79), Expect = 0.28, Method: Compositional matrix adjust.
Identities = 42/170 (24%), Positives = 67/170 (39%), Gaps = 41/170 (24%)
Query 75 DDLFLTTKDGVRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARWLYDQ-GMNVLVI 133
D L L T+ G I ++K ++ T + HGN +G L ++ +N++
Sbjct 46 DILKLRTRCGNEIVAVYVKHSKANG--TLLYSHGNAADLGQMFELFVELSNRLRVNLMGY 103
Query 134 DYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNNNANAISKQQKG 193
DY GYG+S G SE Y D +A+ + K++ G
Sbjct 104 DYSGYGQSTGQASECNTYADIEASYKCL---------------------------KEKYG 136
Query 194 KSSSSSSSSSSKSNFVFVMGHSMGGAVAIDLAKRRGNELAGLVVENTFTS 243
K + + V G S+G +DLA R N L G+V++ S
Sbjct 137 ----------VKDDQLIVYGQSVGSGPTVDLASRTPN-LRGVVLQCPILS 175
> At1g32190_1
Length=289
Score = 35.0 bits (79), Expect = 0.30, Method: Compositional matrix adjust.
Identities = 26/83 (31%), Positives = 42/83 (50%), Gaps = 5/83 (6%)
Query 81 TKDGVRIHGWFLKQTENEKAPTFILF-HGNYGHVGLTLPRARWL-YDQGMNVLVIDYRGY 138
T+ G ++ ++L+ N A +L+ HGN +G L + +N++ DY GY
Sbjct 60 TRRGNKVTAFYLR---NPNARLTLLYSHGNAADLGQLFDLFVQLKVNLRVNLMGYDYSGY 116
Query 139 GRSEGTPSEAGVYMDADAALDFV 161
G S G PSE Y D +AA + +
Sbjct 117 GASTGKPSEYDTYADIEAAYECL 139
> 7301402
Length=838
Score = 34.3 bits (77), Expect = 0.46, Method: Compositional matrix adjust.
Identities = 40/169 (23%), Positives = 67/169 (39%), Gaps = 36/169 (21%)
Query 68 ASYGIPYDDLFLTTKDG-----VRIHGWFLKQTENEKAPTFILFHGNYGHVGLTLPRARW 122
A++G P + + T+DG RI Q +NEK P +L H GLT W
Sbjct 470 AAHGYPSEHHHIVTEDGYILGVFRIPYSHKLQNQNEKRPIVLLQH------GLTSCSDAW 523
Query 123 LYDQGMNVLVIDYRGYGRSEGTPSEAGVYMDADAALDFVLGKQQSSSSSSSNSNSNANNN 182
+ G ++G P Y+ ADA D +G + +S S +++ + ++
Sbjct 524 ILQ-------------GPNDGLP-----YLLADAGFDVWMGNARGTSYSRNHTTLSPDHP 565
Query 183 NANAISKQQKGKSSSS-------SSSSSSKSNFVFVMGHSMGGAVAIDL 224
N S + G + S+ + + + +GHS G V L
Sbjct 566 NFWKFSWHEIGIYDITAIIDYALSTENGQGQDAIHYVGHSQGTTVFFAL 614
> Hs18426888
Length=166
Score = 33.9 bits (76), Expect = 0.62, Method: Compositional matrix adjust.
Identities = 15/27 (55%), Positives = 17/27 (62%), Gaps = 0/27 (0%)
Query 131 LVIDYRGYGRSEGTPSEAGVYMDADAA 157
+ DY GYG S G PSE +Y D DAA
Sbjct 133 IFYDYSGYGASAGRPSERNLYADIDAA 159
Lambda K H
0.314 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 10210178872
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40