bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3332_orf2
Length=197
Score E
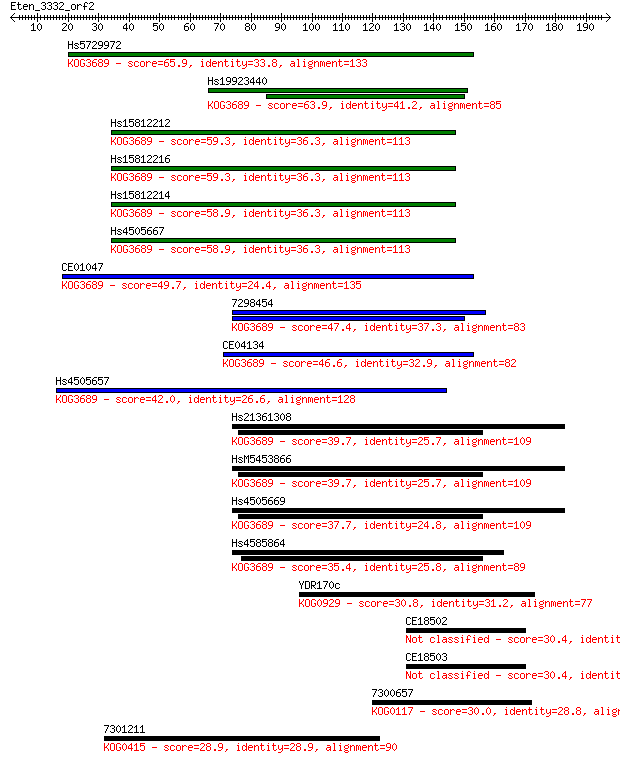
Sequences producing significant alignments: (Bits) Value
Hs5729972 65.9 6e-11
Hs19923440 63.9 2e-10
Hs15812212 59.3 5e-09
Hs15812216 59.3 5e-09
Hs15812214 58.9 6e-09
Hs4505667 58.9 6e-09
CE01047 49.7 4e-06
7298454 47.4 2e-05
CE04134 46.6 3e-05
Hs4505657 42.0 8e-04
Hs21361308 39.7 0.004
HsM5453866 39.7 0.004
Hs4505669 37.7 0.015
Hs4585864 35.4 0.077
YDR170c 30.8 1.8
CE18502 30.4 2.4
CE18503 30.4 2.6
7300657 30.0 3.2
7301211 28.9 6.8
> Hs5729972
Length=779
Score = 65.9 bits (159), Expect = 6e-11, Method: Compositional matrix adjust.
Identities = 45/144 (31%), Positives = 70/144 (48%), Gaps = 16/144 (11%)
Query 20 LDTIFETVLIDGPKLFHCERAALFVYDPSRDEVWS-----------KAVFGEGKALRMQL 68
+D++ E ++I L + +R ALF D E++S K VF + K +R +
Sbjct 267 IDSLLEHIMIYAKNLVNADRCALFQVDHKNKELYSDLFDIGEEKEGKPVFKKTKEIRFSI 326
Query 69 HQSHCIITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALL 128
+ I V E+LN +A DPRFN E D G TR++L P+V++ G I +
Sbjct 327 EKG--IAGQVARTGEVLNIPDAYADPRFNREVDLYTGYTTRNILCMPIVSR-GSVIGVVQ 383
Query 129 FINKHPQFAGHFSEDDERLVEMMS 152
+NK A FS+ DE +M +
Sbjct 384 MVNKISGSA--FSKTDENNFKMFA 405
> Hs19923440
Length=934
Score = 63.9 bits (154), Expect = 2e-10, Method: Compositional matrix adjust.
Identities = 35/85 (41%), Positives = 52/85 (61%), Gaps = 1/85 (1%)
Query 66 MQLHQSHCIITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIA 125
+Q+ II +V E+ E +N +A QD RFN E D+ G KT+S+L P+ + GE I
Sbjct 278 VQVPWGKGIIGYVGEHGETVNIPDAYQDRRFNDEIDKLTGYKTKSLLCMPIRSSDGEIIG 337
Query 126 ALLFINKHPQFAGHFSEDDERLVEM 150
INK P+ A F+EDDE++++M
Sbjct 338 VAQAINKIPEGAP-FTEDDEKVMQM 361
Score = 42.7 bits (99), Expect = 5e-04, Method: Compositional matrix adjust.
Identities = 24/65 (36%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Query 85 LNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHPQFAGHFSEDD 144
+N S+A QDPRF+ E D+ G RSVL P+ N + + I +N+ F + D
Sbjct 486 VNISDAYQDPRFDAEADQISGFHIRSVLCVPIWNSNHQIIGVAQVLNRLD--GKPFDDAD 543
Query 145 ERLVE 149
+RL E
Sbjct 544 QRLFE 548
> Hs15812212
Length=833
Score = 59.3 bits (142), Expect = 5e-09, Method: Compositional matrix adjust.
Identities = 41/123 (33%), Positives = 59/123 (47%), Gaps = 10/123 (8%)
Query 34 LFHCERAALFVY--DPSRDEVWSKAVF--GEGKALR------MQLHQSHCIITWVMENKE 83
L +R +LF+ D S D+ +F EG L ++L + I+ V E
Sbjct 137 LISADRYSLFLVCEDSSNDKFLISRLFDVAEGSTLEEVSNNCIRLEWNKGIVGHVAALGE 196
Query 84 ILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHPQFAGHFSED 143
LN +A +DPRFN E D+ G KT+S+L P+ N E + INK G F+E
Sbjct 197 PLNIKDAYEDPRFNAEVDQITGYKTQSILCMPIKNHREEVVGVAQAINKKSGNGGTFTEK 256
Query 144 DER 146
DE+
Sbjct 257 DEK 259
> Hs15812216
Length=823
Score = 59.3 bits (142), Expect = 5e-09, Method: Compositional matrix adjust.
Identities = 41/123 (33%), Positives = 59/123 (47%), Gaps = 10/123 (8%)
Query 34 LFHCERAALFVY--DPSRDEVWSKAVF--GEGKALR------MQLHQSHCIITWVMENKE 83
L +R +LF+ D S D+ +F EG L ++L + I+ V E
Sbjct 127 LISADRYSLFLVCEDSSNDKFLISRLFDVAEGSTLEEVSNNCIRLEWNKGIVGHVAALGE 186
Query 84 ILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHPQFAGHFSED 143
LN +A +DPRFN E D+ G KT+S+L P+ N E + INK G F+E
Sbjct 187 PLNIKDAYEDPRFNAEVDQITGYKTQSILCMPIKNHREEVVGVAQAINKKSGNGGTFTEK 246
Query 144 DER 146
DE+
Sbjct 247 DEK 249
> Hs15812214
Length=865
Score = 58.9 bits (141), Expect = 6e-09, Method: Compositional matrix adjust.
Identities = 41/123 (33%), Positives = 59/123 (47%), Gaps = 10/123 (8%)
Query 34 LFHCERAALFVY--DPSRDEVWSKAVF--GEGKALR------MQLHQSHCIITWVMENKE 83
L +R +LF+ D S D+ +F EG L ++L + I+ V E
Sbjct 179 LISADRYSLFLVCEDSSNDKFLISRLFDVAEGSTLEEVSNNCIRLEWNKGIVGHVAALGE 238
Query 84 ILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHPQFAGHFSED 143
LN +A +DPRFN E D+ G KT+S+L P+ N E + INK G F+E
Sbjct 239 PLNIKDAYEDPRFNAEVDQITGYKTQSILCMPIKNHREEVVGVAQAINKKSGNGGTFTEK 298
Query 144 DER 146
DE+
Sbjct 299 DEK 301
> Hs4505667
Length=875
Score = 58.9 bits (141), Expect = 6e-09, Method: Compositional matrix adjust.
Identities = 41/123 (33%), Positives = 59/123 (47%), Gaps = 10/123 (8%)
Query 34 LFHCERAALFVY--DPSRDEVWSKAVF--GEGKALR------MQLHQSHCIITWVMENKE 83
L +R +LF+ D S D+ +F EG L ++L + I+ V E
Sbjct 179 LISADRYSLFLVCEDSSNDKFLISRLFDVAEGSTLEEVSNNCIRLEWNKGIVGHVAALGE 238
Query 84 ILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHPQFAGHFSED 143
LN +A +DPRFN E D+ G KT+S+L P+ N E + INK G F+E
Sbjct 239 PLNIKDAYEDPRFNAEVDQITGYKTQSILCMPIKNHREEVVGVAQAINKKSGNGGTFTEK 298
Query 144 DER 146
DE+
Sbjct 299 DEK 301
> CE01047
Length=918
Score = 49.7 bits (117), Expect = 4e-06, Method: Compositional matrix adjust.
Identities = 33/138 (23%), Positives = 63/138 (45%), Gaps = 11/138 (7%)
Query 18 QQLDTIFETVLIDGPKLFHCERAALFVYDPSRDEVW---SKAVFGEGKALRMQLHQSHCI 74
L + ++ + K E A+F++D ++ ++ + GK M + I
Sbjct 235 NNLSALISCIIAEAKKNTEAEDYAVFLHDEDNKQMVLFNNETMLMTGKKFDM----GYGI 290
Query 75 ITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHP 134
+ V +N + S+ P FN E DE+F K R+++A P+++ S I ++ NK
Sbjct 291 VGKVASTMRTMNIRDVSRCPFFNEEIDEQFSIKARNLIAFPLIDSSCSLIGVIVLYNKEN 350
Query 135 QFAGHFSEDDERLVEMMS 152
F+ H DE+ ++ S
Sbjct 351 GFSRH----DEKYIKRFS 364
> 7298454
Length=1284
Score = 47.4 bits (111), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 31/83 (37%), Positives = 44/83 (53%), Gaps = 5/83 (6%)
Query 74 IITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKH 133
I V E+ E +N +A QD RFN E D G +T+++L P+ + SG+ I INK
Sbjct 296 IAGHVAESGEPVNIPDAYQDERFNCEIDSLTGYRTKALLCMPIKDSSGDVIGVAQVINK- 354
Query 134 PQFAGHFSEDDERLVEMMSRHLQ 156
FSE DE++ S +LQ
Sbjct 355 -MNGECFSEIDEKV---FSSYLQ 373
Score = 41.6 bits (96), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 25/76 (32%), Positives = 36/76 (47%), Gaps = 2/76 (2%)
Query 74 IITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKH 133
I V E +N A +D RF+ DE K RS+L + N G+ I + INK
Sbjct 478 ITGHVATTGETVNVPNAYEDDRFDASVDENSCFKHRSILCMAIKNSLGQIIGVIQLINKF 537
Query 134 PQFAGHFSEDDERLVE 149
+ F+++DE VE
Sbjct 538 NEL--DFTKNDENFVE 551
> CE04134
Length=393
Score = 46.6 bits (109), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 27/82 (32%), Positives = 41/82 (50%), Gaps = 4/82 (4%)
Query 71 SHCIITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFI 130
S I +V E LN A +D RFN + D K G T+++L P++ + G I + +
Sbjct 2 SKGIAGYVASTGEGLNIENAYEDERFNADVDSKTGYTTKTILCMPILIR-GIVIGVVQMV 60
Query 131 NKHPQFAGHFSEDDERLVEMMS 152
NKH G F+ DE E+ +
Sbjct 61 NKHD---GVFTRQDEDAFEIFA 79
> Hs4505657
Length=941
Score = 42.0 bits (97), Expect = 8e-04, Method: Compositional matrix adjust.
Identities = 34/134 (25%), Positives = 64/134 (47%), Gaps = 10/134 (7%)
Query 16 HQQQLDTIFETVLIDGPKLFHCERAALFVYDPSRDEVWSKAVFGEG----KALRMQLHQS 71
H + + + ++ + L + E ++F+ D ++E+ +K VF G ++ +++
Sbjct 406 HLDDVSVLLQEIITEARNLSNAEICSVFLLD--QNELVAK-VFDGGVVDDESYEIRIPAD 462
Query 72 HCIITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFIN 131
I V +ILN +A P F D+ G +TR++L P+ N++ E I +N
Sbjct 463 QGIAGHVATTGQILNIPDAYAHPLFYRGVDDSTGFRTRNILCFPIKNENQEVIGVAELVN 522
Query 132 K--HPQFAGHFSED 143
K P F+ F ED
Sbjct 523 KINGPWFS-KFDED 535
> Hs21361308
Length=858
Score = 39.7 bits (91), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 28/113 (24%), Positives = 51/113 (45%), Gaps = 7/113 (6%)
Query 74 IITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKH 133
I+ W K+ N + ++ F+ D++ G T+++LA P+V E +A ++ +NK
Sbjct 142 IVGWAAHTKKTHNVPDVKKNSHFSDFMDKQTGYVTKNLLATPIV-VGKEVLAVIMAVNKV 200
Query 134 PQFAGHFSEDDERLVEMMSRHLQIIME----KFEYGAAEDRKMIKLPETDHVF 182
A FS+ DE + + II+ + Y R I + + VF
Sbjct 201 N--ASEFSKQDEEVFSKYLNFVSIILRLHHTSYMYNIESRRSQILMWSANKVF 251
Score = 35.4 bits (80), Expect = 0.071, Method: Compositional matrix adjust.
Identities = 27/83 (32%), Positives = 40/83 (48%), Gaps = 6/83 (7%)
Query 76 TWVMENKEILNCSEASQDPRFN-PE--FDEKFGSKTRSVLAAPVVNKSGEAIAALLFINK 132
T+V EN I N A D F P+ DE G ++VL+ P+VNK + + F N+
Sbjct 350 TYVAENGFICNMMNAPADEYFTFPKGPVDET-GWVIKNVLSLPIVNKKEDIVGVATFYNR 408
Query 133 HPQFAGHFSEDDERLVEMMSRHL 155
F E DE + E +++ L
Sbjct 409 KD--GKPFDEHDEYITETLTQFL 429
> HsM5453866
Length=858
Score = 39.7 bits (91), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 28/113 (24%), Positives = 51/113 (45%), Gaps = 7/113 (6%)
Query 74 IITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKH 133
I+ W K+ N + ++ F+ D++ G T+++LA P+V E +A ++ +NK
Sbjct 142 IVGWAAHTKKTHNVPDVKKNSHFSDFMDKQTGYVTKNLLATPIV-VGKEVLAVIMAVNKV 200
Query 134 PQFAGHFSEDDERLVEMMSRHLQIIME----KFEYGAAEDRKMIKLPETDHVF 182
A FS+ DE + + II+ + Y R I + + VF
Sbjct 201 N--ASEFSKQDEEVFSKYLNFVSIILRLHHTSYMYNIESRRSQILMWSANKVF 251
Score = 35.4 bits (80), Expect = 0.073, Method: Compositional matrix adjust.
Identities = 26/83 (31%), Positives = 39/83 (46%), Gaps = 6/83 (7%)
Query 76 TWVMENKEILNCSEASQDPRFNPE---FDEKFGSKTRSVLAAPVVNKSGEAIAALLFINK 132
T+V EN I N A D F + DE G ++VL+ P+VNK + + F N+
Sbjct 350 TYVAENGFICNMMNAPADEYFTFQKGPVDET-GWVIKNVLSLPIVNKKEDIVGVATFYNR 408
Query 133 HPQFAGHFSEDDERLVEMMSRHL 155
F E DE + E +++ L
Sbjct 409 KD--GKPFDEHDEYITETLTQFL 429
> Hs4505669
Length=854
Score = 37.7 bits (86), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 27/114 (23%), Positives = 59/114 (51%), Gaps = 9/114 (7%)
Query 74 IITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKH 133
++ V + K+++N + ++ P F+ DE KT+++LA P++N + +A ++ +NK
Sbjct 138 VVGHVAQTKKMVNVEDVAECPHFSSFADELTDYKTKNMLATPIMN-GKDVVAVIMAVNK- 195
Query 134 PQFAGHF--SEDDERLVEMM---SRHLQIIMEKFEYGAAEDRKMIKLPETDHVF 182
G F SED++ ++ + + +L+I + + R + L + VF
Sbjct 196 --LNGPFFTSEDEDVFLKYLNFATLYLKIYHLSYLHNCETRRGQVLLWSANKVF 247
Score = 35.0 bits (79), Expect = 0.087, Method: Compositional matrix adjust.
Identities = 27/84 (32%), Positives = 41/84 (48%), Gaps = 8/84 (9%)
Query 76 TWVMENKEILNCSEASQDPRFNPEFDEKF----GSKTRSVLAAPVVNKSGEAIAALLFIN 131
++V E+ I N AS D F +F E G ++VL+ P+VNK E + F N
Sbjct 346 SYVAESGFICNIMNASADEMF--KFQEGALDDSGWLIKNVLSMPIVNKKEEIVGVATFYN 403
Query 132 KHPQFAGHFSEDDERLVEMMSRHL 155
+ F E DE L+E +++ L
Sbjct 404 RKD--GKPFDEQDEVLMESLTQFL 425
> Hs4585864
Length=860
Score = 35.4 bits (80), Expect = 0.077, Method: Compositional matrix adjust.
Identities = 23/89 (25%), Positives = 45/89 (50%), Gaps = 3/89 (3%)
Query 74 IITWVMENKEILNCSEASQDPRFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKH 133
I+ V +K+I N +D F D KT+++LA+P++N + +A ++ +NK
Sbjct 140 IVGHVAHSKKIANVPNTEEDEHFCDFVDILTEYKTKNILASPIMN-GKDVVAIIMAVNKV 198
Query 134 PQFAGHFSEDDERLVEMMSRHLQIIMEKF 162
HF++ DE ++ +IM+ +
Sbjct 199 D--GSHFTKRDEEILLKYLNFANLIMKWY 225
Score = 32.7 bits (73), Expect = 0.48, Method: Compositional matrix adjust.
Identities = 24/81 (29%), Positives = 39/81 (48%), Gaps = 4/81 (4%)
Query 77 WVMENKEILNCSEASQDP--RFNPEFDEKFGSKTRSVLAAPVVNKSGEAIAALLFINKHP 134
+V +N I N A + F E ++ G ++VL+ P+VNK E + F N+
Sbjct 349 YVAQNGLICNIMNAPAEDFFAFQKEPLDESGWMIKNVLSMPIVNKKEEIVGVATFYNRKD 408
Query 135 QFAGHFSEDDERLVEMMSRHL 155
F E DE L+E +++ L
Sbjct 409 --GKPFDEMDETLMESLTQFL 427
> YDR170c
Length=2009
Score = 30.8 bits (68), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 24/83 (28%), Positives = 46/83 (55%), Gaps = 10/83 (12%)
Query 96 FNPEFDEKFGSKTRSVLAAPVVNK-----SGEAIAALLFINKHPQFAGHFSEDDERLVEM 150
F+ +F+E +G +TR ++ A VV+K E AA + ++ Q + DDE+ ++
Sbjct 1863 FSRDFNEDYGLRTR-LVEARVVDKIPNLLKQETSAAAVLLDIMFQL---YLNDDEKKADL 1918
Query 151 MSRHLQIIMEKFE-YGAAEDRKM 172
++R + I ++ E Y + +DR M
Sbjct 1919 ITRLITICIQVVEGYVSLDDRTM 1941
> CE18502
Length=213
Score = 30.4 bits (67), Expect = 2.4, Method: Compositional matrix adjust.
Identities = 16/39 (41%), Positives = 21/39 (53%), Gaps = 0/39 (0%)
Query 131 NKHPQFAGHFSEDDERLVEMMSRHLQIIMEKFEYGAAED 169
+K P H SE ERL+ + SR L II E ++G D
Sbjct 50 SKAPFLLEHISEAKERLITVTSRGLMIIYENDDHGLVID 88
> CE18503
Length=289
Score = 30.4 bits (67), Expect = 2.6, Method: Compositional matrix adjust.
Identities = 16/39 (41%), Positives = 21/39 (53%), Gaps = 0/39 (0%)
Query 131 NKHPQFAGHFSEDDERLVEMMSRHLQIIMEKFEYGAAED 169
+K P H SE ERL+ + SR L II E ++G D
Sbjct 50 SKAPFLLEHISEAKERLITVTSRGLMIIYENDDHGLVID 88
> 7300657
Length=673
Score = 30.0 bits (66), Expect = 3.2, Method: Compositional matrix adjust.
Identities = 15/54 (27%), Positives = 28/54 (51%), Gaps = 2/54 (3%)
Query 120 SGEAIAALLFINKHPQFAGHFSEDDER--LVEMMSRHLQIIMEKFEYGAAEDRK 171
+G+ I + N H F G+ ++ +R L+E S+H + E Y + +D+K
Sbjct 269 TGKKIGVTISFNNHRLFVGNIPKNRDRDELIEEFSKHAPGLYEVIIYSSPDDKK 322
> 7301211
Length=653
Score = 28.9 bits (63), Expect = 6.8, Method: Compositional matrix adjust.
Identities = 26/91 (28%), Positives = 39/91 (42%), Gaps = 4/91 (4%)
Query 32 PKLFHCERAALFVYDPSRDEVWSKAVFGEGKALRMQLHQSHCIITWVMENKEILN-CSEA 90
PK+ H L + ++ V S+ G+ L L +HC+I V+E E+L ++A
Sbjct 84 PKINHSSAGMLSLVSAGKNLVGSQFFLTLGENL-TSLDGNHCVIGEVVEGHEVLRKLNDA 142
Query 91 SQDPRFNPEFDEKFGSKTRSVLAAPVVNKSG 121
D F P D + VL P N G
Sbjct 143 IVDDSFRPYQDIRITHTV--VLEDPFPNPRG 171
Lambda K H
0.319 0.133 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3371754244
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40