bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3502_orf2
Length=77
Score E
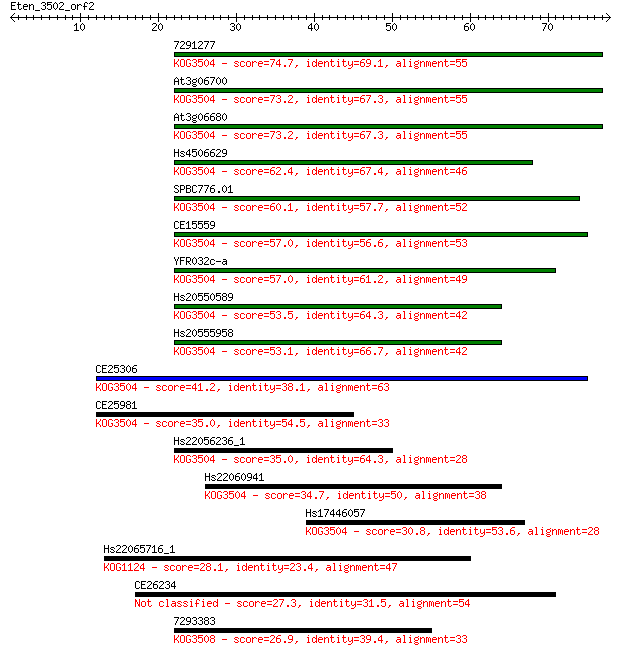
Sequences producing significant alignments: (Bits) Value
7291277 74.7 4e-14
At3g06700 73.2 1e-13
At3g06680 73.2 1e-13
Hs4506629 62.4 2e-10
SPBC776.01 60.1 1e-09
CE15559 57.0 7e-09
YFR032c-a 57.0 8e-09
Hs20550589 53.5 1e-07
Hs20555958 53.1 1e-07
CE25306 41.2 4e-04
CE25981 35.0 0.031
Hs22056236_1 35.0 0.032
Hs22060941 34.7 0.045
Hs17446057 30.8 0.68
Hs22065716_1 28.1 3.7
CE26234 27.3 7.5
7293383 26.9 9.1
> 7291277
Length=76
Score = 74.7 bits (182), Expect = 4e-14, Method: Compositional matrix adjust.
Identities = 38/55 (69%), Positives = 44/55 (80%), Gaps = 0/55 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKGMLKQKE 76
MAKSKNHTNHNQN+KAHRNGIK+ R+R ST GM KFL NQRY +KG L ++E
Sbjct 1 MAKSKNHTNHNQNKKAHRNGIKRPLRKRHESTLGMDVKFLINQRYARKGNLSREE 55
> At3g06700
Length=61
Score = 73.2 bits (178), Expect = 1e-13, Method: Compositional matrix adjust.
Identities = 37/55 (67%), Positives = 42/55 (76%), Gaps = 0/55 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKGMLKQKE 76
MAKSKNHT HNQ+ KAH+NGIKK R R T T+GM PKFLRNQRY +K +K E
Sbjct 1 MAKSKNHTAHNQSAKAHKNGIKKPRRHRHTPTRGMDPKFLRNQRYARKHNVKAGE 55
> At3g06680
Length=61
Score = 73.2 bits (178), Expect = 1e-13, Method: Compositional matrix adjust.
Identities = 37/55 (67%), Positives = 42/55 (76%), Gaps = 0/55 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKGMLKQKE 76
MAKSKNHT HNQ+ KAH+NGIKK R R T T+GM PKFLRNQRY +K +K E
Sbjct 1 MAKSKNHTAHNQSAKAHKNGIKKPRRHRHTPTRGMDPKFLRNQRYARKHNVKSGE 55
> Hs4506629
Length=159
Score = 62.4 bits (150), Expect = 2e-10, Method: Compositional matrix adjust.
Identities = 31/46 (67%), Positives = 35/46 (76%), Gaps = 0/46 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYC 67
MAKSKNHT HNQ+RK HRNGIKK +R S KG+ PKFLRN R+
Sbjct 1 MAKSKNHTTHNQSRKWHRNGIKKPRSQRYESLKGVDPKFLRNMRFA 46
> SPBC776.01
Length=61
Score = 60.1 bits (144), Expect = 1e-09, Method: Compositional matrix adjust.
Identities = 30/52 (57%), Positives = 38/52 (73%), Gaps = 0/52 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKGMLK 73
MAKSKNHTNHNQN+KAHRNGIK+ + R S K KF RNQ++ +G ++
Sbjct 1 MAKSKNHTNHNQNKKAHRNGIKRPQKHRYDSLKYRDAKFRRNQKFANRGTVE 52
> CE15559
Length=62
Score = 57.0 bits (136), Expect = 7e-09, Method: Compositional matrix adjust.
Identities = 30/53 (56%), Positives = 38/53 (71%), Gaps = 0/53 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKGMLKQ 74
MAKSKNHTNHNQN+KAHRNGI K + S KG+ KF++N R+ +K +Q
Sbjct 1 MAKSKNHTNHNQNKKAHRNGITKPKKHIFLSMKGVDAKFIKNLRFSRKNNKRQ 53
> YFR032c-a
Length=59
Score = 57.0 bits (136), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 30/49 (61%), Positives = 34/49 (69%), Gaps = 0/49 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKG 70
MAKSKNHT HNQ RKAHRNGIKK + S KG+ PKF RN ++ G
Sbjct 1 MAKSKNHTAHNQTRKAHRNGIKKPKTYKYPSLKGVDPKFRRNHKHALHG 49
> Hs20550589
Length=209
Score = 53.5 bits (127), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 27/42 (64%), Positives = 31/42 (73%), Gaps = 0/42 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRN 63
MAKSKNH H+Q +K HRNGIKK +R S KG+ PKFLRN
Sbjct 21 MAKSKNHNTHDQFQKRHRNGIKKPQSQRSVSLKGVDPKFLRN 62
> Hs20555958
Length=220
Score = 53.1 bits (126), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 28/42 (66%), Positives = 31/42 (73%), Gaps = 0/42 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRN 63
MAKSKNHT HNQ+RK HRN IKK +R S KG+ PKFL N
Sbjct 1 MAKSKNHTTHNQSRKWHRNVIKKPLSQRYKSLKGVDPKFLGN 42
> CE25306
Length=87
Score = 41.2 bits (95), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 24/63 (38%), Positives = 36/63 (57%), Gaps = 0/63 (0%)
Query 12 FVLSTTSSKKMAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKGM 71
F + T K +NHTNHN+N KAHRNGI K + S +G +F+++ R+ +K
Sbjct 2 FSMKTIMPHNAWKPENHTNHNRNNKAHRNGITKPKKHIFLSIEGSRRQFIKSLRFFRKNN 61
Query 72 LKQ 74
+Q
Sbjct 62 KRQ 64
> CE25981
Length=223
Score = 35.0 bits (79), Expect = 0.031, Method: Compositional matrix adjust.
Identities = 18/33 (54%), Positives = 19/33 (57%), Gaps = 0/33 (0%)
Query 12 FVLSTTSSKKMAKSKNHTNHNQNRKAHRNGIKK 44
F + KSKN TNHNQN KAHR GI K
Sbjct 124 FEIVEREQHDAWKSKNRTNHNQNNKAHRIGITK 156
> Hs22056236_1
Length=29
Score = 35.0 bits (79), Expect = 0.032, Method: Compositional matrix adjust.
Identities = 18/28 (64%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 22 MAKSKNHTNHNQNRKAHRNGIKKVPRRR 49
MAKSKNH H+Q +K HRNGIKK +R
Sbjct 1 MAKSKNHNTHDQFQKRHRNGIKKPQSQR 28
> Hs22060941
Length=162
Score = 34.7 bits (78), Expect = 0.045, Method: Compositional matrix adjust.
Identities = 19/38 (50%), Positives = 24/38 (63%), Gaps = 0/38 (0%)
Query 26 KNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRN 63
KNH HNQ+ K HRNGI+K +R G+ KFLR+
Sbjct 42 KNHNTHNQSPKWHRNGIRKPRSQRYKCLNGVDLKFLRS 79
> Hs17446057
Length=258
Score = 30.8 bits (68), Expect = 0.68, Method: Composition-based stats.
Identities = 15/28 (53%), Positives = 19/28 (67%), Gaps = 0/28 (0%)
Query 39 RNGIKKVPRRRKTSTKGMCPKFLRNQRY 66
+NGIKK +R S KG+ PKFLRN +
Sbjct 3 QNGIKKPRLQRYESLKGVDPKFLRNMGF 30
> Hs22065716_1
Length=1131
Score = 28.1 bits (61), Expect = 3.7, Method: Composition-based stats.
Identities = 11/47 (23%), Positives = 24/47 (51%), Gaps = 0/47 (0%)
Query 13 VLSTTSSKKMAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPK 59
V + + S K + ++ H ++R+A ++ + + PR + K PK
Sbjct 967 VAAVSGSCKRRQKEHQKEHRRHRRACKDSVGRRPREGRAKAKAKVPK 1013
> CE26234
Length=323
Score = 27.3 bits (59), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 17/54 (31%), Positives = 30/54 (55%), Gaps = 6/54 (11%)
Query 17 TSSKKMAKSKNHTNHNQNRKAHRNGIKKVPRRRKTSTKGMCPKFLRNQRYCKKG 70
T+++K + K H ++ R A N I+ PR+ + + KG+C K + C+KG
Sbjct 2 TNAEKWGRLKAHADYGHTRFACVNKIE-APRKGEQNEKGICKKGI-----CEKG 49
> 7293383
Length=1436
Score = 26.9 bits (58), Expect = 9.1, Method: Composition-based stats.
Identities = 13/35 (37%), Positives = 21/35 (60%), Gaps = 2/35 (5%)
Query 22 MAKSKNHTNHNQNRKAHRNGIK--KVPRRRKTSTK 54
+A+ NH NHNQ ++ H+ +K VP + + S K
Sbjct 964 VAEPANHHNHNQGQQNHQGHLKPAAVPGKEQLSAK 998
Lambda K H
0.322 0.131 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1175087368
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40