bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
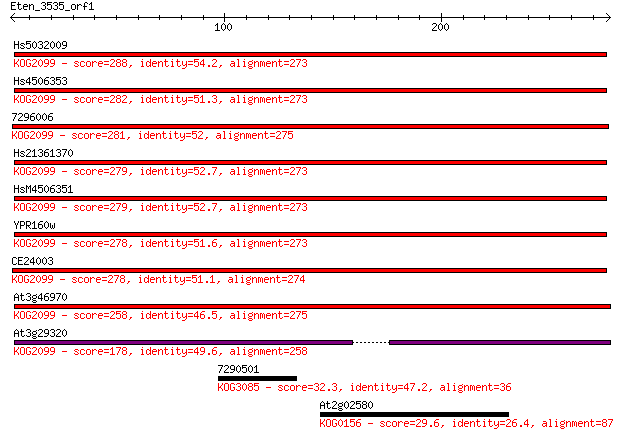
Query= Eten_3535_orf1
Length=277
Score E
Sequences producing significant alignments: (Bits) Value
Hs5032009 288 7e-78
Hs4506353 282 6e-76
7296006 281 7e-76
Hs21361370 279 4e-75
HsM4506351 279 4e-75
YPR160w 278 8e-75
CE24003 278 1e-74
At3g46970 258 8e-69
At3g29320 178 1e-44
7290501 32.3 1.1
At2g02580 29.6 7.9
> Hs5032009
Length=842
Score = 288 bits (738), Expect = 7e-78, Method: Compositional matrix adjust.
Identities = 148/280 (52%), Positives = 194/280 (69%), Gaps = 13/280 (4%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFK--KV 60
F+VG Y+++V +R AE+IS VLYPNDN EGKELRLKQ+YF AT+QD++RRFK K
Sbjct 258 FNVGGYIQAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFVVAATLQDIIRRFKSSKF 317
Query 61 PGRD-----WKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTN 115
RD + PDK+ QLNDTHP++AIPELMRIL+D+E +DWD AWD+T R YTN
Sbjct 318 GCRDPVRTNFDAFPDKVAIQLNDTHPSLAIPELMRILVDLERMDWDKAWDVTVRTCAYTN 377
Query 116 HTVLPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGNE 175
HTVLPEALE+W L+ LLPRHL II EIN RFLN + F D +++ RMS+ EEG
Sbjct 378 HTVLPEALERWPVHLLETLLPRHLQIIYEINQRFLNRVAAAFPGDVDRLRRMSLVEEGAV 437
Query 176 KRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRW 235
KRI MA+L + G VNGVA IHSE++KK +F +F E KF N TNG+TPRRW
Sbjct 438 KRINMAHLCIAGSHAVNGVARIHSEILKKTIFKDFYELEPH-----KFQNKTNGITPRRW 492
Query 236 IYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSF 275
+ N GLA++ + +G D ++ +LD + L++ +D+ +F
Sbjct 493 LVLCNPGLAEVIAERIGED-FISDLDQLRKLLSFVDDEAF 531
> Hs4506353
Length=847
Score = 282 bits (721), Expect = 6e-76, Method: Compositional matrix adjust.
Identities = 140/280 (50%), Positives = 190/280 (67%), Gaps = 13/280 (4%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFK---- 58
F+VG Y+++V +R AE+IS VLYPNDN EGKELRLKQ+YF AT+QD++RRFK
Sbjct 258 FNVGDYIQAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFVVAATLQDIIRRFKASKF 317
Query 59 ---KVPGRDWKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTN 115
+ G + PD++ QLNDTHP +AIPELMRI +D+E L W AW+LT++ F YTN
Sbjct 318 GSTRGAGTVFDAFPDQVAIQLNDTHPALAIPELMRIFVDIEKLPWSKAWELTQKTFAYTN 377
Query 116 HTVLPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGNE 175
HTVLPEALE+W DL+ +LLPRHL II EIN + L+ ++F D +++ RMS+ EE
Sbjct 378 HTVLPEALERWPVDLVEKLLPRHLEIIYEINQKHLDRIVALFPKDVDRLRRMSLIEEEGS 437
Query 176 KRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRW 235
KRI MA+L ++G VNGVA IHS++VK +F +F E DKF N TNG+TPRRW
Sbjct 438 KRINMAHLCIVGSHAVNGVAKIHSDIVKTKVFKDFSELEP-----DKFQNKTNGITPRRW 492
Query 236 IYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSF 275
+ N GLA+L + +G D ++K+L + L + + + F
Sbjct 493 LLLCNPGLAELIAEKIGED-YVKDLSQLTKLHSFLGDDVF 531
> 7296006
Length=844
Score = 281 bits (720), Expect = 7e-76, Method: Compositional matrix adjust.
Identities = 143/282 (50%), Positives = 190/282 (67%), Gaps = 13/282 (4%)
Query 2 LFDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFK--K 59
F+ G Y+++V +R AE+IS VLYPNDN EGKELRLKQ+YF C AT+QD++RR+K K
Sbjct 257 FFNDGDYIQAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFMCAATLQDIIRRYKASK 316
Query 60 VPGRD-----WKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYT 114
R+ + PDK+ QLNDTHP++AIPELMRIL+D E L W+ AWD+T R YT
Sbjct 317 FGSREAVRNTFDHFPDKVAIQLNDTHPSLAIPELMRILVDEEHLTWEKAWDITVRSCAYT 376
Query 115 NHTVLPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGN 174
NHTVLPEALE+W L+ +LPRHL II INF + + F DD +++ RMS+ EE
Sbjct 377 NHTVLPEALERWPVSLLESILPRHLQIIYHINFLHMENVKKKFPDDLDRMRRMSMVEEDG 436
Query 175 EKRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRR 234
EKRI MA+L+++G VNGVAAIHS+++K +LF +F E + KF N TNG+TPRR
Sbjct 437 EKRINMAHLSIVGSHAVNGVAAIHSQILKDSLFHDFYEMEPQ-----KFQNKTNGITPRR 491
Query 235 WIYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSFR 276
W+ N GL+DL + +G D W LD + L +P+F+
Sbjct 492 WLLLCNPGLSDLIAEKIG-DEWPVHLDQLVALKKWAKDPNFQ 532
> Hs21361370
Length=843
Score = 279 bits (714), Expect = 4e-75, Method: Compositional matrix adjust.
Identities = 144/280 (51%), Positives = 192/280 (68%), Gaps = 13/280 (4%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFK--KV 60
F+VG Y+E+V +R AE+IS VLYPNDN EGKELRLKQ+YF AT+QD++RRFK K
Sbjct 258 FNVGDYIEAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFVVAATLQDIIRRFKSSKF 317
Query 61 PGRD-----WKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTN 115
RD ++ PDK+ QLNDTHP ++IPELMRIL+DVE +DWD AW++T++ YTN
Sbjct 318 GCRDPVRTCFETFPDKVAIQLNDTHPALSIPELMRILVDVEKVDWDKAWEITKKTCAYTN 377
Query 116 HTVLPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGNE 175
HTVLPEALE+W + +LLPRHL II IN R L+ ++F D +++ RMS+ EEG+
Sbjct 378 HTVLPEALERWPVSMFEKLLPRHLEIIYAINQRHLDHVAALFPGDVDRLRRMSVIEEGDC 437
Query 176 KRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRW 235
KRI MA+L VIG VNGVA IHSE+VK+++F +F E +KF N TNG+TPRRW
Sbjct 438 KRINMAHLCVIGSHAVNGVARIHSEIVKQSVFKDFYELEP-----EKFQNKTNGITPRRW 492
Query 236 IYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSF 275
+ N GLAD +G + +L +L + L+ + + F
Sbjct 493 LLLCNPGLADTIVEKIG-EEFLTDLSQLKKLLPLVSDEVF 531
> HsM4506351
Length=843
Score = 279 bits (714), Expect = 4e-75, Method: Compositional matrix adjust.
Identities = 144/280 (51%), Positives = 192/280 (68%), Gaps = 13/280 (4%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFK--KV 60
F+VG Y+E+V +R AE+IS VLYPNDN EGKELRLKQ+YF AT+QD++RRFK K
Sbjct 258 FNVGDYIEAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFVVAATLQDIIRRFKSSKF 317
Query 61 PGRD-----WKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTN 115
RD ++ PDK+ QLNDTHP ++IPELMRIL+DVE +DWD AW++T++ YTN
Sbjct 318 GCRDPVRTCFETFPDKVAIQLNDTHPALSIPELMRILVDVEKVDWDKAWEITKKTCAYTN 377
Query 116 HTVLPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGNE 175
HTVLPEALE+W + +LLPRHL II IN R L+ ++F D +++ RMS+ EEG+
Sbjct 378 HTVLPEALERWPVSMFEKLLPRHLEIIYAINQRHLDHVAALFPGDVDRLRRMSVIEEGDC 437
Query 176 KRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRW 235
KRI MA+L VIG VNGVA IHSE+VK+++F +F E +KF N TNG+TPRRW
Sbjct 438 KRINMAHLCVIGSHAVNGVARIHSEIVKQSVFKDFYELEP-----EKFQNKTNGITPRRW 492
Query 236 IYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSF 275
+ N GLAD +G + +L +L + L+ + + F
Sbjct 493 LLLCNPGLADTIVEKIG-EEFLTDLSQLKKLLPLVSDEVF 531
> YPR160w
Length=902
Score = 278 bits (711), Expect = 8e-75, Method: Compositional matrix adjust.
Identities = 141/276 (51%), Positives = 189/276 (68%), Gaps = 8/276 (2%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFKKVPG 62
F+ G Y SV ++Q AESI+AVLYPNDN +GKELRLKQQYF+C A++ D+LRRFKK
Sbjct 316 FNNGDYKNSVAQQQRAESITAVLYPNDNFAQGKELRLKQQYFWCAASLHDILRRFKK-SK 374
Query 63 RDWKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTNHTVLPEA 122
R W E PD++ QLNDTHPT+AI EL R+L+D+E LDW AWD+ + F YTNHTV+ EA
Sbjct 375 RPWTEFPDQVAIQLNDTHPTLAIVELQRVLVDLEKLDWHEAWDIVTKTFAYTNHTVMQEA 434
Query 123 LEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGN-EKRIRMA 181
LEKW L LLPRHL II +IN+ FL + F D + + R+SI EE + E++IRMA
Sbjct 435 LEKWPVGLFGHLLPRHLEIIYDINWFFLQDVAKKFPKDVDLLSRISIIEENSPERQIRMA 494
Query 182 NLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRWIYCSNR 241
LA++G VNGVA +HSEL+K +F +F +FY KF+NVTNG+TPRRW+ +N
Sbjct 495 FLAIVGSHKVNGVAELHSELIKTTIFKDFVKFYGPS----KFVNVTNGITPRRWLKQANP 550
Query 242 GLADLFSNWLG--SDSWLKELDMIAGLMNHIDNPSF 275
LA L S L ++ +L ++ + L ++++ F
Sbjct 551 SLAKLISETLNDPTEEYLLDMAKLTQLGKYVEDKEF 586
> CE24003
Length=882
Score = 278 bits (710), Expect = 1e-74, Method: Compositional matrix adjust.
Identities = 140/284 (49%), Positives = 192/284 (67%), Gaps = 16/284 (5%)
Query 2 LFDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFKK-V 60
F+ G Y+++V +R +E+I+ VLYPNDN GKELRLKQQYF AT+QD++RRFK +
Sbjct 291 FFNDGDYVQAVMDRNLSENITRVLYPNDNMFLGKELRLKQQYFLVAATLQDIIRRFKSSI 350
Query 61 PGR------DWKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYT 114
G +++ PDK+ QLNDTHP+I IPEL+R+L+DVEGL WD AWD+ + + YT
Sbjct 351 YGNREAVRVNFETFPDKVAIQLNDTHPSIGIPELIRLLIDVEGLTWDQAWDICIKTYAYT 410
Query 115 NHTVLPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGN 174
NHT+LPEALE+W L+ LLPRHL II EIN +F+N F D++++ RMSI EE +
Sbjct 411 NHTLLPEALERWPVSLMQNLLPRHLEIIYEINQKFMNTISQRFPGDFDRMRRMSIVEEAD 470
Query 175 ---EKRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVT 231
EKRI MA+L ++ +NGVAA+HS+L+K + F +F EFY D+F N TNG+T
Sbjct 471 QFGEKRINMAHLCIVASHAINGVAALHSDLLKSSTFRDFYEFYP-----DRFQNKTNGIT 525
Query 232 PRRWIYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSF 275
PRRW+ SN LADL +G +SW+ LD + L + ++ F
Sbjct 526 PRRWLLLSNPSLADLIVEKIG-ESWITNLDELQKLKEYANDAGF 568
> At3g46970
Length=841
Score = 258 bits (660), Expect = 8e-69, Method: Compositional matrix adjust.
Identities = 128/280 (45%), Positives = 177/280 (63%), Gaps = 10/280 (3%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFKKVP- 61
F+ G Y + + A+ I VLYP D T GK LRLKQQ+F C A++QD++ RF +
Sbjct 262 FNEGEYELAAQLHSRAQQICTVLYPGDATENGKLLRLKQQFFLCSASLQDIISRFHERST 321
Query 62 ---GRDWKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTNHTV 118
R W E P K+ Q+NDTHPT+AIPELMR+L+D GL WD AWD+T + YTNHTV
Sbjct 322 TEGSRKWSEFPSKVAVQMNDTHPTLAIPELMRLLMDDNGLGWDEAWDVTSKTVAYTNHTV 381
Query 119 LPEALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFGDDWNKIGRMSIYEEGNEKR- 177
LPEALEKWS L+ +LLPRH+ II EI+ RF+ R D +KI +SI + +K
Sbjct 382 LPEALEKWSQSLMWKLLPRHMEIIEEIDKRFVQTIRDTRVDLEDKISSLSILDNNPQKPV 441
Query 178 IRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRWIY 237
+RMANL V+ VNGVA +HS+++K LF ++ + +KF N TNG+TPRRW+
Sbjct 442 VRMANLCVVSSHTVNGVAQLHSDILKAELFADYVSIWP-----NKFQNKTNGITPRRWLR 496
Query 238 CSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSFRA 277
+ L+D+ + WL +D W+ +LD++ GL DN ++
Sbjct 497 FCSPELSDIITKWLKTDKWITDLDLLTGLRQFADNEELQS 536
> At3g29320
Length=962
Score = 178 bits (451), Expect = 1e-44, Method: Compositional matrix adjust.
Identities = 81/158 (51%), Positives = 112/158 (70%), Gaps = 2/158 (1%)
Query 3 FDVGRYLESVRERQNAESISAVLYPNDNTMEGKELRLKQQYFFCCATIQDVLRRFKKVPG 62
++ G++ E+ NAE I VLYP D + EGK LRLKQQY C A++QD++ RF+ G
Sbjct 328 YNSGKHTEAAEALFNAEKICFVLYPGDESTEGKALRLKQQYTLCSASLQDIVARFETRSG 387
Query 63 RD--WKELPDKIQCQLNDTHPTIAIPELMRILLDVEGLDWDTAWDLTRRCFNYTNHTVLP 120
+ W+E P+K+ Q+NDTHPT+ IPELMRIL+D++GL W+ AW +T+R YTNHTVLP
Sbjct 388 GNVNWEEFPEKVAVQMNDTHPTLCIPELMRILMDLKGLSWEDAWKITQRTVAYTNHTVLP 447
Query 121 EALEKWSADLISRLLPRHLLIINEINFRFLNEARSIFG 158
EALEKWS +L+ +LLPRH+ II +I+ + S +G
Sbjct 448 EALEKWSLELMEKLLPRHVEIIEKIDEELVRTIVSEYG 485
Score = 97.8 bits (242), Expect = 2e-20, Method: Compositional matrix adjust.
Identities = 47/102 (46%), Positives = 68/102 (66%), Gaps = 5/102 (4%)
Query 176 KRIRMANLAVIGGRHVNGVAAIHSELVKKNLFPEFDEFYSRQGVNDKFLNVTNGVTPRRW 235
K +RMANLAV+GG VNGVA IHSE+VK+++F +F + + +KF N TNGVTPRRW
Sbjct 560 KMVRMANLAVVGGHAVNGVAEIHSEIVKQDVFNDFVQLWP-----EKFQNKTNGVTPRRW 614
Query 236 IYCSNRGLADLFSNWLGSDSWLKELDMIAGLMNHIDNPSFRA 277
I N L+D+ +NW+G++ W+ + +A L DN ++
Sbjct 615 IRFCNPYLSDIITNWIGTEDWVLNTEKVAELRKFADNEDLQS 656
> 7290501
Length=246
Score = 32.3 bits (72), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 17/37 (45%), Positives = 20/37 (54%), Gaps = 1/37 (2%)
Query 97 GLDWDTAW-DLTRRCFNYTNHTVLPEALEKWSADLIS 132
GLDW W L R F+ +H + PE LEK LIS
Sbjct 76 GLDWQRWWLQLVARTFSCVDHGLAPEKLEKIGQRLIS 112
> At2g02580
Length=500
Score = 29.6 bits (65), Expect = 7.9, Method: Compositional matrix adjust.
Identities = 23/89 (25%), Positives = 42/89 (47%), Gaps = 15/89 (16%)
Query 144 EINFRFLNEARSIFGDDWNKIGRMSIYEEGNEKRIRMANLAVIGGRHVNGVAAIHSELVK 203
E+++ +L+ A S F D W ++ R+ + E + KR V+ + I E V+
Sbjct 106 ELSYNYLDIAFSPFDDYWKELRRICVQELFSAKR-------------VHSIQPIKEEEVR 152
Query 204 KNLFPEFDEFYSRQGVN--DKFLNVTNGV 230
K + + + VN +KFL++T V
Sbjct 153 KLIVSATESASQKSPVNLSEKFLDLTVSV 181
Lambda K H
0.323 0.139 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 6021067820
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40