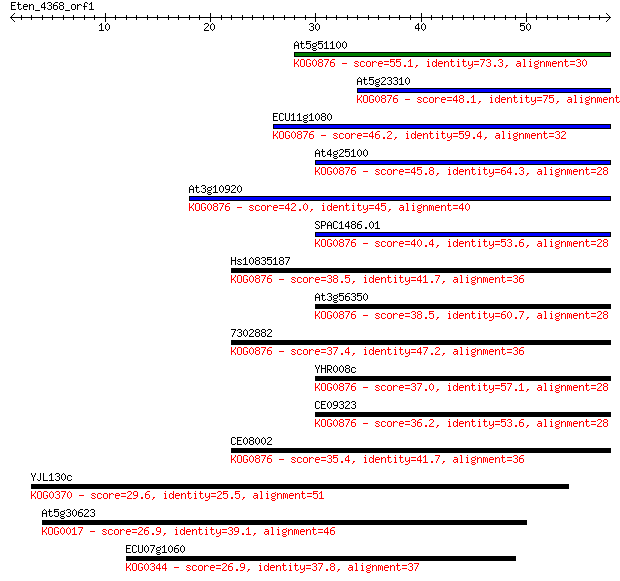
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_4368_orf1
Length=57
Score E
Sequences producing significant alignments: (Bits) Value
At5g51100 55.1 3e-08
At5g23310 48.1 3e-06
ECU11g1080 46.2 2e-05
At4g25100 45.8 2e-05
At3g10920 42.0 3e-04
SPAC1486.01 40.4 0.001
Hs10835187 38.5 0.003
At3g56350 38.5 0.003
7302882 37.4 0.007
YHR008c 37.0 0.008
CE09323 36.2 0.014
CE08002 35.4 0.024
YJL130c 29.6 1.3
At5g30623 26.9 8.1
ECU07g1060 26.9 9.7
> At5g51100
Length=305
Score = 55.1 bits (131), Expect = 3e-08, Method: Compositional matrix adjust.
Identities = 22/30 (73%), Positives = 28/30 (93%), Gaps = 0/30 (0%)
Query 28 FELPPLPYPMDALEPYISKETLEYHYGKHH 57
FEL P PYP+DALEP++S+ETL+YH+GKHH
Sbjct 53 FELKPPPYPLDALEPHMSRETLDYHWGKHH 82
> At5g23310
Length=263
Score = 48.1 bits (113), Expect = 3e-06, Method: Composition-based stats.
Identities = 18/24 (75%), Positives = 22/24 (91%), Gaps = 0/24 (0%)
Query 34 PYPMDALEPYISKETLEYHYGKHH 57
PYP+DALEPY+S+ TLE H+GKHH
Sbjct 56 PYPLDALEPYMSRRTLEVHWGKHH 79
> ECU11g1080
Length=233
Score = 46.2 bits (108), Expect = 2e-05, Method: Composition-based stats.
Identities = 19/32 (59%), Positives = 23/32 (71%), Gaps = 0/32 (0%)
Query 26 MPFELPPLPYPMDALEPYISKETLEYHYGKHH 57
M F+LP L YP DALEP I +ET+ H+ KHH
Sbjct 8 MEFKLPELDYPYDALEPIIDEETMRTHHSKHH 39
> At4g25100
Length=212
Score = 45.8 bits (107), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 18/28 (64%), Positives = 25/28 (89%), Gaps = 0/28 (0%)
Query 30 LPPLPYPMDALEPYISKETLEYHYGKHH 57
L P P+ +DALEP++SK+TLE+H+GKHH
Sbjct 13 LKPPPFALDALEPHMSKQTLEFHWGKHH 40
> At3g10920
Length=231
Score = 42.0 bits (97), Expect = 3e-04, Method: Composition-based stats.
Identities = 18/40 (45%), Positives = 23/40 (57%), Gaps = 0/40 (0%)
Query 18 KFLELSAKMPFELPPLPYPMDALEPYISKETLEYHYGKHH 57
+ L + F LP LPY ALEP IS E ++ H+ KHH
Sbjct 21 RLLRIRGIQTFTLPDLPYDYGALEPAISGEIMQIHHQKHH 60
> SPAC1486.01
Length=218
Score = 40.4 bits (93), Expect = 0.001, Method: Composition-based stats.
Identities = 15/28 (53%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 30 LPPLPYPMDALEPYISKETLEYHYGKHH 57
LPPLPY +ALEP +S+ ++ H+ KHH
Sbjct 28 LPPLPYAYNALEPALSETIMKLHHDKHH 55
> Hs10835187
Length=222
Score = 38.5 bits (88), Expect = 0.003, Method: Composition-based stats.
Identities = 15/36 (41%), Positives = 23/36 (63%), Gaps = 0/36 (0%)
Query 22 LSAKMPFELPPLPYPMDALEPYISKETLEYHYGKHH 57
L ++ LP LPY ALEP+I+ + ++ H+ KHH
Sbjct 20 LGSRQKHSLPDLPYDYGALEPHINAQIMQLHHSKHH 55
> At3g56350
Length=241
Score = 38.5 bits (88), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 17/28 (60%), Positives = 20/28 (71%), Gaps = 0/28 (0%)
Query 30 LPPLPYPMDALEPYISKETLEYHYGKHH 57
LP LPY DALEP IS+E + H+ KHH
Sbjct 38 LPDLPYAYDALEPAISEEIMRLHHQKHH 65
> 7302882
Length=217
Score = 37.4 bits (85), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 17/36 (47%), Positives = 22/36 (61%), Gaps = 0/36 (0%)
Query 22 LSAKMPFELPPLPYPMDALEPYISKETLEYHYGKHH 57
L+ + LP LPY ALEP I +E +E H+ KHH
Sbjct 13 LAVRGKHTLPKLPYDYAALEPIICREIMELHHQKHH 48
> YHR008c
Length=233
Score = 37.0 bits (84), Expect = 0.008, Method: Composition-based stats.
Identities = 16/28 (57%), Positives = 18/28 (64%), Gaps = 0/28 (0%)
Query 30 LPPLPYPMDALEPYISKETLEYHYGKHH 57
LP L + ALEPYIS + E HY KHH
Sbjct 30 LPDLKWDFGALEPYISGQINELHYTKHH 57
> CE09323
Length=221
Score = 36.2 bits (82), Expect = 0.014, Method: Composition-based stats.
Identities = 15/28 (53%), Positives = 18/28 (64%), Gaps = 0/28 (0%)
Query 30 LPPLPYPMDALEPYISKETLEYHYGKHH 57
LP LPY LEP IS E ++ H+ KHH
Sbjct 28 LPDLPYDYADLEPVISHEIMQLHHQKHH 55
> CE08002
Length=218
Score = 35.4 bits (80), Expect = 0.024, Method: Composition-based stats.
Identities = 15/36 (41%), Positives = 21/36 (58%), Gaps = 0/36 (0%)
Query 22 LSAKMPFELPPLPYPMDALEPYISKETLEYHYGKHH 57
L+ + LP LP+ LEP IS E ++ H+ KHH
Sbjct 20 LAVRSKHTLPDLPFDYADLEPVISHEIMQLHHQKHH 55
> YJL130c
Length=2214
Score = 29.6 bits (65), Expect = 1.3, Method: Composition-based stats.
Identities = 13/52 (25%), Positives = 29/52 (55%), Gaps = 1/52 (1%)
Query 3 RTNSAFEFLKHTLLLKFLELSAKMPFELPPLPYPMDAL-EPYISKETLEYHY 53
R + +F F+ + + +EL+ K LP PYP++ L + Y++ + ++ +
Sbjct 1265 RASRSFPFISKVVGVNLIELATKAIMGLPLTPYPVEKLPDDYVAVKVPQFSF 1316
> At5g30623
Length=302
Score = 26.9 bits (58), Expect = 8.1, Method: Composition-based stats.
Identities = 18/47 (38%), Positives = 27/47 (57%), Gaps = 4/47 (8%)
Query 4 TNSAFEFLKHTLLLKFLELSAKMPFELP-PLPYPMDALEPYISKETL 49
T + EF+ +L+F S+ MP++LP LP PM EP I ++ L
Sbjct 219 TQTEDEFIDPAGVLRFGSSSSAMPYDLPVTLPIPM---EPPIFQQYL 262
> ECU07g1060
Length=416
Score = 26.9 bits (58), Expect = 9.7, Method: Composition-based stats.
Identities = 14/37 (37%), Positives = 20/37 (54%), Gaps = 2/37 (5%)
Query 12 KHTLLLKFLELSAKMPFELPPLPYPMDALEPYISKET 48
+H LL F +L K+PF+ P L + PY K+T
Sbjct 29 RHPLLCSFEDLKTKVPFDSPLL--TQKVVIPYFLKKT 63
Lambda K H
0.320 0.139 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1191047922
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40