bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_4563_orf3
Length=85
Score E
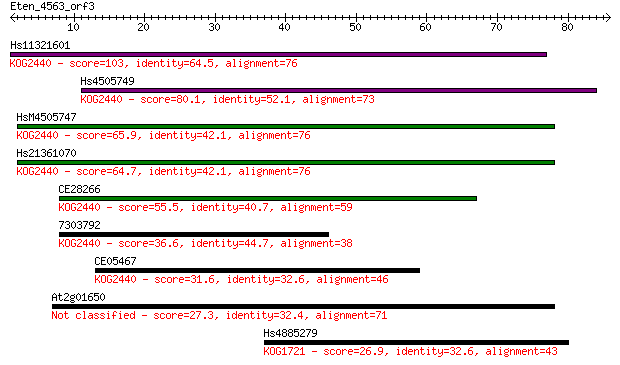
Sequences producing significant alignments: (Bits) Value
Hs11321601 103 8e-23
Hs4505749 80.1 9e-16
HsM4505747 65.9 2e-11
Hs21361070 64.7 4e-11
CE28266 55.5 2e-08
7303792 36.6 0.010
CE05467 31.6 0.34
At2g01650 27.3 7.7
Hs4885279 26.9 8.8
> Hs11321601
Length=784
Score = 103 bits (257), Expect = 8e-23, Method: Composition-based stats.
Identities = 49/76 (64%), Positives = 59/76 (77%), Gaps = 0/76 (0%)
Query 1 GKGKKFLSEASGCVLGISKRTLLFQPGGELKKETNLEPRIPKEQWGLKLGPLMKFLAKYR 60
G+GKKF ++ S CVLGISKR ++FQP ELKK+T+ E RIPKEQW LKL PLMK LAKY+
Sbjct 706 GRGKKFTTDDSICVLGISKRNVIFQPVAELKKQTDFEHRIPKEQWWLKLRPLMKILAKYK 765
Query 61 VSFGGSDASQREPVQP 76
S+ SD+ Q E VQP
Sbjct 766 ASYDVSDSGQLEHVQP 781
> Hs4505749
Length=780
Score = 80.1 bits (196), Expect = 9e-16, Method: Composition-based stats.
Identities = 38/73 (52%), Positives = 48/73 (65%), Gaps = 0/73 (0%)
Query 11 SGCVLGISKRTLLFQPGGELKKETNLEPRIPKEQWGLKLGPLMKFLAKYRVSFGGSDASQ 70
SGCVLG+ KR L+FQP ELK +T+ E RIPKEQW LKL P++K LAKY + SD +
Sbjct 707 SGCVLGMRKRALVFQPVAELKDQTDFEHRIPKEQWWLKLRPILKILAKYEIDLDTSDHAH 766
Query 71 REPVQPRGPEEPA 83
E + + E A
Sbjct 767 LEHITRKRSGEAA 779
> HsM4505747
Length=780
Score = 65.9 bits (159), Expect = 2e-11, Method: Composition-based stats.
Identities = 32/77 (41%), Positives = 48/77 (62%), Gaps = 1/77 (1%)
Query 2 KGKKFLSEA-SGCVLGISKRTLLFQPGGELKKETNLEPRIPKEQWGLKLGPLMKFLAKYR 60
KG+ F + S CV+G+ K+ + F P ELKK+T+ E R+P+EQW L L ++K LA+YR
Sbjct 696 KGRVFANAPDSACVIGLKKKAVAFSPVTELKKDTDFEHRMPREQWWLSLRLMLKMLAQYR 755
Query 61 VSFGGSDASQREPVQPR 77
+S + + E V R
Sbjct 756 ISMAAYVSGELEHVTRR 772
> Hs21361070
Length=827
Score = 64.7 bits (156), Expect = 4e-11, Method: Composition-based stats.
Identities = 32/77 (41%), Positives = 48/77 (62%), Gaps = 1/77 (1%)
Query 2 KGKKFLSEA-SGCVLGISKRTLLFQPGGELKKETNLEPRIPKEQWGLKLGPLMKFLAKYR 60
KG+ F + S CV+G+ K+ + F P ELKK+T+ E R+P+EQW L L ++K LA+YR
Sbjct 743 KGRVFANAPDSACVIGLKKKAVAFSPVTELKKDTDFEHRMPREQWWLSLRLMLKMLAQYR 802
Query 61 VSFGGSDASQREPVQPR 77
+S + + E V R
Sbjct 803 ISMAAYVSGELEHVTRR 819
> CE28266
Length=814
Score = 55.5 bits (132), Expect = 2e-08, Method: Composition-based stats.
Identities = 24/59 (40%), Positives = 38/59 (64%), Gaps = 0/59 (0%)
Query 8 SEASGCVLGISKRTLLFQPGGELKKETNLEPRIPKEQWGLKLGPLMKFLAKYRVSFGGS 66
S + +LG+ R ++F P +LKKET+ E R+P EQW + L PL++ LA++R + S
Sbjct 740 SAHTATLLGLKGRKVVFTPVQDLKKETDFEHRLPSEQWWMALRPLLRVLARHRSTVESS 798
> 7303792
Length=950
Score = 36.6 bits (83), Expect = 0.010, Method: Composition-based stats.
Identities = 17/38 (44%), Positives = 22/38 (57%), Gaps = 0/38 (0%)
Query 8 SEASGCVLGISKRTLLFQPGGELKKETNLEPRIPKEQW 45
S + +LGI R F P +L ETN + RIPK+QW
Sbjct 876 SPDTATLLGIVSRQYRFSPLVDLIAETNFDQRIPKKQW 913
> CE05467
Length=756
Score = 31.6 bits (70), Expect = 0.34, Method: Composition-based stats.
Identities = 15/46 (32%), Positives = 24/46 (52%), Gaps = 0/46 (0%)
Query 13 CVLGISKRTLLFQPGGELKKETNLEPRIPKEQWGLKLGPLMKFLAK 58
CV+G+ +L + P L K+ E +P W L L PL++ + K
Sbjct 702 CVIGLRGSSLRYVPVQGLGKKVCFEHGVPHNMWWLDLHPLVEAMTK 747
> At2g01650
Length=465
Score = 27.3 bits (59), Expect = 7.7, Method: Compositional matrix adjust.
Identities = 23/74 (31%), Positives = 35/74 (47%), Gaps = 5/74 (6%)
Query 7 LSEASGCVLGISKRTLLFQPGGELKKETN---LEPRIPKEQWGLKLGPLMKFLAKYRVSF 63
+ EA G V G LL G ELK+E + +P E+ + + ++ +L K +
Sbjct 212 IKEAIGDVAG--GVELLELVGFELKEENDEIWAVMDVPSEEQSILINKVVGYLEKRKTES 269
Query 64 GGSDASQREPVQPR 77
GS A EPV P+
Sbjct 270 SGSSAQVMEPVAPK 283
> Hs4885279
Length=1106
Score = 26.9 bits (58), Expect = 8.8, Method: Compositional matrix adjust.
Identities = 14/43 (32%), Positives = 19/43 (44%), Gaps = 0/43 (0%)
Query 37 EPRIPKEQWGLKLGPLMKFLAKYRVSFGGSDASQREPVQPRGP 79
EP + G L P M F + +GG + + EP RGP
Sbjct 706 EPEVGTSMVGSGLNPYMDFPPTDTLGYGGPEGAAAEPYGARGP 748
Lambda K H
0.315 0.138 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1194132014
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40