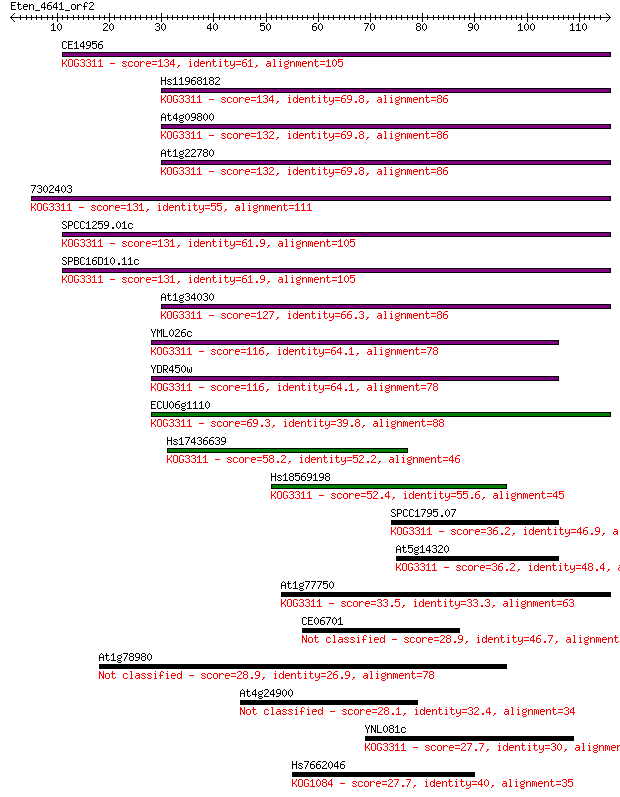
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_4641_orf2
Length=115
Score E
Sequences producing significant alignments: (Bits) Value
CE14956 134 5e-32
Hs11968182 134 5e-32
At4g09800 132 2e-31
At1g22780 132 2e-31
7302403 131 3e-31
SPCC1259.01c 131 4e-31
SPBC16D10.11c 131 4e-31
At1g34030 127 7e-30
YML026c 116 8e-27
YDR450w 116 8e-27
ECU06g1110 69.3 1e-12
Hs17436639 58.2 4e-09
Hs18569198 52.4 2e-07
SPCC1795.07 36.2 0.015
At5g14320 36.2 0.015
At1g77750 33.5 0.089
CE06701 28.9 2.6
At1g78980 28.9 2.7
At4g24900 28.1 4.5
YNL081c 27.7 4.8
Hs7662046 27.7 5.7
> CE14956
Length=154
Score = 134 bits (336), Expect = 5e-32, Method: Compositional matrix adjust.
Identities = 64/111 (57%), Positives = 80/111 (72%), Gaps = 6/111 (5%)
Query 11 AHCCCSPAA------AALLLLLLLLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSA 64
A CC A A L +IV I+ +P+Q+ IP+WFLNRQKD+KDGK+ + +
Sbjct 41 AFVCCRKADVDVNKRAGELTEEDFDKIVTIMQNPSQYKIPNWFLNRQKDIKDGKTGQLLS 100
Query 65 NNLDSVWREDFERLKKMRLHRGLRHYWGLKVRGQHTKTTGRHGRTVGVAKK 115
+D+ RED ER+KK+RLHRGLRHYWGL+VRGQHTKTTGR GRTVGV+KK
Sbjct 101 TAVDNKLREDLERMKKIRLHRGLRHYWGLRVRGQHTKTTGRKGRTVGVSKK 151
> Hs11968182
Length=152
Score = 134 bits (336), Expect = 5e-32, Method: Compositional matrix adjust.
Identities = 60/86 (69%), Positives = 71/86 (82%), Gaps = 0/86 (0%)
Query 30 QIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRH 89
+++ I+ +P Q+ IP WFLNRQKDVKDGK V AN LD+ RED ERLKK+R HRGLRH
Sbjct 66 RVITIMQNPRQYKIPDWFLNRQKDVKDGKYSQVLANGLDNKLREDLERLKKIRAHRGLRH 125
Query 90 YWGLKVRGQHTKTTGRHGRTVGVAKK 115
+WGL+VRGQHTKTTGR GRTVGV+KK
Sbjct 126 FWGLRVRGQHTKTTGRRGRTVGVSKK 151
> At4g09800
Length=152
Score = 132 bits (331), Expect = 2e-31, Method: Compositional matrix adjust.
Identities = 60/86 (69%), Positives = 69/86 (80%), Gaps = 0/86 (0%)
Query 30 QIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRH 89
++ IVA+P QF IP WFLNRQKD KDGK V +N LD R+D ERLKK+R HRGLRH
Sbjct 66 NLMTIVANPRQFKIPDWFLNRQKDYKDGKYSQVVSNALDMKLRDDLERLKKIRNHRGLRH 125
Query 90 YWGLKVRGQHTKTTGRHGRTVGVAKK 115
YWGL+VRGQHTKTTGR G+TVGV+KK
Sbjct 126 YWGLRVRGQHTKTTGRRGKTVGVSKK 151
> At1g22780
Length=152
Score = 132 bits (331), Expect = 2e-31, Method: Compositional matrix adjust.
Identities = 60/86 (69%), Positives = 69/86 (80%), Gaps = 0/86 (0%)
Query 30 QIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRH 89
++ IVA+P QF IP WFLNRQKD KDGK V +N LD R+D ERLKK+R HRGLRH
Sbjct 66 NLMTIVANPRQFKIPDWFLNRQKDYKDGKYSQVVSNALDMKLRDDLERLKKIRNHRGLRH 125
Query 90 YWGLKVRGQHTKTTGRHGRTVGVAKK 115
YWGL+VRGQHTKTTGR G+TVGV+KK
Sbjct 126 YWGLRVRGQHTKTTGRRGKTVGVSKK 151
> 7302403
Length=152
Score = 131 bits (330), Expect = 3e-31, Method: Compositional matrix adjust.
Identities = 61/111 (54%), Positives = 78/111 (70%), Gaps = 10/111 (9%)
Query 5 SLTTSAAHCCCSPAAAALLLLLLLLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSA 64
LT A C ++V I+++P Q+ +P+WFLNRQKD+ DGK +++
Sbjct 51 DLTKRAGECTEEEVD----------KVVTIISNPLQYKVPNWFLNRQKDIIDGKYWQLTS 100
Query 65 NNLDSVWREDFERLKKMRLHRGLRHYWGLKVRGQHTKTTGRHGRTVGVAKK 115
+NLDS R+D ERLKK+R HRGLRHYWGL+VRGQHTKTTGR GRTVGV+KK
Sbjct 101 SNLDSKLRDDLERLKKIRSHRGLRHYWGLRVRGQHTKTTGRRGRTVGVSKK 151
> SPCC1259.01c
Length=152
Score = 131 bits (329), Expect = 4e-31, Method: Compositional matrix adjust.
Identities = 65/111 (58%), Positives = 78/111 (70%), Gaps = 6/111 (5%)
Query 11 AHCCCSPAA------AALLLLLLLLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSA 64
A+ C A A L L +IV I+ +P+QF IPSWFLNRQKD+ DGKS + A
Sbjct 41 ANIVCKKADIDMSKRAGELTTEELERIVTIIQNPSQFKIPSWFLNRQKDINDGKSFQLLA 100
Query 65 NNLDSVWREDFERLKKMRLHRGLRHYWGLKVRGQHTKTTGRHGRTVGVAKK 115
NN+DS RED ERLKK++ HRGLRH L+VRGQHTKTTGR G+TVGV+KK
Sbjct 101 NNVDSKLREDLERLKKIQTHRGLRHALDLRVRGQHTKTTGRRGKTVGVSKK 151
> SPBC16D10.11c
Length=152
Score = 131 bits (329), Expect = 4e-31, Method: Compositional matrix adjust.
Identities = 65/111 (58%), Positives = 78/111 (70%), Gaps = 6/111 (5%)
Query 11 AHCCCSPAA------AALLLLLLLLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSA 64
A+ C A A L L +IV I+ +P+QF IPSWFLNRQKD+ DGKS + A
Sbjct 41 ANIVCKKADIDMSKRAGELTTEELERIVTIIQNPSQFKIPSWFLNRQKDINDGKSFQLLA 100
Query 65 NNLDSVWREDFERLKKMRLHRGLRHYWGLKVRGQHTKTTGRHGRTVGVAKK 115
NN+DS RED ERLKK++ HRGLRH L+VRGQHTKTTGR G+TVGV+KK
Sbjct 101 NNVDSKLREDLERLKKIQTHRGLRHALDLRVRGQHTKTTGRRGKTVGVSKK 151
> At1g34030
Length=237
Score = 127 bits (318), Expect = 7e-30, Method: Compositional matrix adjust.
Identities = 57/86 (66%), Positives = 67/86 (77%), Gaps = 0/86 (0%)
Query 30 QIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRH 89
++ IVA+P QF IP WFLNRQKD KDGK V +N LD R+D ERLKK+R HRGLRH
Sbjct 79 NLMTIVANPRQFKIPDWFLNRQKDYKDGKYSQVVSNALDMKLRDDLERLKKIRNHRGLRH 138
Query 90 YWGLKVRGQHTKTTGRHGRTVGVAKK 115
YWGL+VRGQHTKTTGR G+TV + +K
Sbjct 139 YWGLRVRGQHTKTTGRRGKTVALDQK 164
> YML026c
Length=146
Score = 116 bits (291), Expect = 8e-27, Method: Compositional matrix adjust.
Identities = 50/78 (64%), Positives = 63/78 (80%), Gaps = 0/78 (0%)
Query 28 LLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGL 87
L +IV I+ +PT + IP+WFLNRQ D+ DGK H ANN++S R+D ERLKK+R HRG+
Sbjct 66 LERIVQIMQNPTHYKIPAWFLNRQNDITDGKDYHTLANNVESKLRDDLERLKKIRAHRGI 125
Query 88 RHYWGLKVRGQHTKTTGR 105
RH+WGL+VRGQHTKTTGR
Sbjct 126 RHFWGLRVRGQHTKTTGR 143
> YDR450w
Length=146
Score = 116 bits (291), Expect = 8e-27, Method: Compositional matrix adjust.
Identities = 50/78 (64%), Positives = 63/78 (80%), Gaps = 0/78 (0%)
Query 28 LLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGL 87
L +IV I+ +PT + IP+WFLNRQ D+ DGK H ANN++S R+D ERLKK+R HRG+
Sbjct 66 LERIVQIMQNPTHYKIPAWFLNRQNDITDGKDYHTLANNVESKLRDDLERLKKIRAHRGI 125
Query 88 RHYWGLKVRGQHTKTTGR 105
RH+WGL+VRGQHTKTTGR
Sbjct 126 RHFWGLRVRGQHTKTTGR 143
> ECU06g1110
Length=153
Score = 69.3 bits (168), Expect = 1e-12, Method: Compositional matrix adjust.
Identities = 35/88 (39%), Positives = 51/88 (57%), Gaps = 0/88 (0%)
Query 28 LLQIVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGL 87
+ +I + P IP ++N Q+D+ DG + H+ LD+ R ER KK + R
Sbjct 65 MKRISDAILDPASVGIPESYMNHQRDIIDGTTSHLIGTRLDADLRMMIERGKKNKRIRAY 124
Query 88 RHYWGLKVRGQHTKTTGRHGRTVGVAKK 115
R GLKVRGQ TK+ GR GR++GV++K
Sbjct 125 RLDVGLKVRGQRTKSNGRRGRSMGVSRK 152
> Hs17436639
Length=112
Score = 58.2 bits (139), Expect = 4e-09, Method: Compositional matrix adjust.
Identities = 24/46 (52%), Positives = 31/46 (67%), Gaps = 0/46 (0%)
Query 31 IVAIVASPTQFSIPSWFLNRQKDVKDGKSRHVSANNLDSVWREDFE 76
++ ++ +P Q+ IP WFLNRQKDVKDGK AN LD+ ED E
Sbjct 67 VITMMQNPHQYKIPGWFLNRQKDVKDGKYSQALANGLDNKLHEDLE 112
> Hs18569198
Length=139
Score = 52.4 bits (124), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 25/45 (55%), Positives = 31/45 (68%), Gaps = 0/45 (0%)
Query 51 QKDVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRHYWGLKV 95
QKD K G V AN+LD+ RED E+LKK+ RGL H+WGL+V
Sbjct 95 QKDGKAGNYSQVLANDLDNKLREDLEQLKKIPACRGLHHFWGLRV 139
> SPCC1795.07
Length=151
Score = 36.2 bits (82), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 15/32 (46%), Positives = 21/32 (65%), Gaps = 0/32 (0%)
Query 74 DFERLKKMRLHRGLRHYWGLKVRGQHTKTTGR 105
D RL +R +RG+RH GL V GQ+T+T +
Sbjct 76 DISRLVNIRCYRGMRHVNGLPVNGQNTRTNAK 107
> At5g14320
Length=197
Score = 36.2 bits (82), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 15/31 (48%), Positives = 21/31 (67%), Gaps = 0/31 (0%)
Query 75 FERLKKMRLHRGLRHYWGLKVRGQHTKTTGR 105
+RLK+++ +RG+RH GL RGQ TK R
Sbjct 150 IKRLKEIQCYRGVRHIQGLPCRGQRTKNNCR 180
> At1g77750
Length=154
Score = 33.5 bits (75), Expect = 0.089, Method: Compositional matrix adjust.
Identities = 21/65 (32%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Query 53 DVKDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRHYWGLKVRGQHTKTTGR--HGRTV 110
D+++ +H + L + +RL ++ +RG RH GL RGQ T T R G+ V
Sbjct 84 DLREEVGQHQHGDELRRRVGSEIQRLVEVDCYRGSRHRHGLPCRGQRTSTNARTKKGKAV 143
Query 111 GVAKK 115
+A K
Sbjct 144 AIAGK 148
> CE06701
Length=5170
Score = 28.9 bits (63), Expect = 2.6, Method: Composition-based stats.
Identities = 14/30 (46%), Positives = 19/30 (63%), Gaps = 0/30 (0%)
Query 57 GKSRHVSANNLDSVWREDFERLKKMRLHRG 86
GK RH S++NL ++FERL+K RG
Sbjct 4840 GKYRHQSSDNLSLTSLQEFERLEKEVGARG 4869
> At1g78980
Length=693
Score = 28.9 bits (63), Expect = 2.7, Method: Composition-based stats.
Identities = 21/82 (25%), Positives = 42/82 (51%), Gaps = 5/82 (6%)
Query 18 AAAALLLLLLLLQIVAIVASPTQFSIPSWFLNRQKDVKDGK----SRHVSANNLDSVWRE 73
A A L +L+L++ ++A+V S + S+ F++ K + H SA L +
Sbjct 277 AGACLGVLVLIIVLIALV-SKKKSSLSPHFIDEDNSHHTPKFKSLTSHGSAQELRVDFGN 335
Query 74 DFERLKKMRLHRGLRHYWGLKV 95
D++ ++ + + GL+HY +V
Sbjct 336 DYKGIRHISTYLGLKHYVSSRV 357
> At4g24900
Length=423
Score = 28.1 bits (61), Expect = 4.5, Method: Composition-based stats.
Identities = 11/34 (32%), Positives = 21/34 (61%), Gaps = 0/34 (0%)
Query 45 SWFLNRQKDVKDGKSRHVSANNLDSVWREDFERL 78
+W R+ +++ KS HV+ +N+D W +F R+
Sbjct 339 AWAERRKIEIEMEKSGHVTKSNIDPDWLPNFGRV 372
> YNL081c
Length=143
Score = 27.7 bits (60), Expect = 4.8, Method: Compositional matrix adjust.
Identities = 12/40 (30%), Positives = 23/40 (57%), Gaps = 0/40 (0%)
Query 69 SVWREDFERLKKMRLHRGLRHYWGLKVRGQHTKTTGRHGR 108
++ +++ +K+ + G+RH L VRGQHT+ + R
Sbjct 73 AIVKDNIALKRKIGSYSGMRHTLHLPVRGQHTRNNAKTAR 112
> Hs7662046
Length=2715
Score = 27.7 bits (60), Expect = 5.7, Method: Composition-based stats.
Identities = 14/35 (40%), Positives = 21/35 (60%), Gaps = 2/35 (5%)
Query 55 KDGKSRHVSANNLDSVWREDFERLKKMRLHRGLRH 89
+DG S V A +L+ WR E++++ R H LRH
Sbjct 2421 EDGFS--VEAESLEGAWRTLIEKVQEARGHARLRH 2453
Lambda K H
0.324 0.133 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1178471386
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40