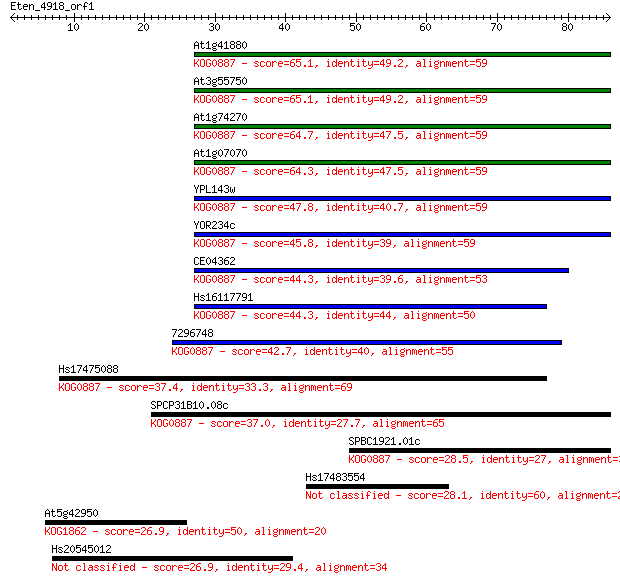
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_4918_orf1
Length=85
Score E
Sequences producing significant alignments: (Bits) Value
At1g41880 65.1 3e-11
At3g55750 65.1 3e-11
At1g74270 64.7 5e-11
At1g07070 64.3 6e-11
YPL143w 47.8 5e-06
YOR234c 45.8 2e-05
CE04362 44.3 5e-05
Hs16117791 44.3 6e-05
7296748 42.7 2e-04
Hs17475088 37.4 0.007
SPCP31B10.08c 37.0 0.009
SPBC1921.01c 28.5 3.4
Hs17483554 28.1 3.9
At5g42950 26.9 8.1
Hs20545012 26.9 9.1
> At1g41880
Length=111
Score = 65.1 bits (157), Expect = 3e-11, Method: Compositional matrix adjust.
Identities = 29/59 (49%), Positives = 42/59 (71%), Gaps = 0/59 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
L+ RG VLGYKRS+ NQ P L+ +E + T+++ +Y+GKR+AY Y+AKTK G +R
Sbjct 11 LYVRGTVLGYKRSKSNQYPNTSLIQIEGVNTQEEVNWYKGKRLAYIYKAKTKKNGSHYR 69
> At3g55750
Length=111
Score = 65.1 bits (157), Expect = 3e-11, Method: Compositional matrix adjust.
Identities = 29/59 (49%), Positives = 42/59 (71%), Gaps = 0/59 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
L+ RG VLGYKRS+ NQ P L+ +E + T+++ +Y+GKR+AY Y+AKTK G +R
Sbjct 11 LYVRGTVLGYKRSKSNQYPNTSLVQIEGVNTQEEVNWYKGKRLAYIYKAKTKKNGSHYR 69
> At1g74270
Length=112
Score = 64.7 bits (156), Expect = 5e-11, Method: Compositional matrix adjust.
Identities = 28/59 (47%), Positives = 42/59 (71%), Gaps = 0/59 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
L+ RG +LGYKRS+ NQ P L+ +E + T+++ +Y+GKR+AY Y+AKTK G +R
Sbjct 12 LYVRGTILGYKRSKSNQYPNTSLVQIEGVNTQEEVNWYKGKRMAYIYKAKTKKNGSHYR 70
> At1g07070
Length=112
Score = 64.3 bits (155), Expect = 6e-11, Method: Compositional matrix adjust.
Identities = 28/59 (47%), Positives = 41/59 (69%), Gaps = 0/59 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
L+ RG +LGYKRS+ NQ P L+ +E + T ++ +Y+GKR+AY Y+AKTK G +R
Sbjct 12 LYVRGTILGYKRSKSNQYPNTSLVQVEGVNTTEEVSWYKGKRMAYIYKAKTKKNGSHYR 70
> YPL143w
Length=107
Score = 47.8 bits (112), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 24/59 (40%), Positives = 34/59 (57%), Gaps = 0/59 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
L+ +G L Y+RS+ P L+ +E + T +D FY GKR+AY YRA + RG R
Sbjct 7 LYVKGKHLSYQRSKRVNNPNVSLIKIEGVATPQDAQFYLGKRIAYVYRASKEVRGSKIR 65
> YOR234c
Length=107
Score = 45.8 bits (107), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 23/59 (38%), Positives = 34/59 (57%), Gaps = 0/59 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
L+ +G L Y+RS+ P L+ +E + T ++ FY GKR+AY YRA + RG R
Sbjct 7 LYVKGKHLSYQRSKRVNNPNVSLIKIEGVATPQEAQFYLGKRIAYVYRASKEVRGSKIR 65
> CE04362
Length=124
Score = 44.3 bits (103), Expect = 5e-05, Method: Compositional matrix adjust.
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 0/53 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKN 79
L+ + I G+KR Q+ LL LE + ++D FY GKRV Y Y+A K
Sbjct 18 LYVKAIFTGFKRGLRTQSEHTSLLKLEGVFNKEDAGFYAGKRVVYLYKAHNKT 70
> Hs16117791
Length=110
Score = 44.3 bits (103), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 22/50 (44%), Positives = 28/50 (56%), Gaps = 0/50 (0%)
Query 27 LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAK 76
L+ + I GYKR NQ LL +E + R + FY GKR AY Y+AK
Sbjct 5 LWSKAIFAGYKRGLRNQREHTALLKIEGVYARDETEFYLGKRCAYVYKAK 54
> 7296748
Length=157
Score = 42.7 bits (99), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 22/56 (39%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query 24 RYG-LFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTK 78
R+G LF + + GYKR NQ +L +E + ++ FY GKR Y Y+A+TK
Sbjct 46 RHGRLFAKAVFTGYKRGLRNQHENQAILKIEGARRKEHGSFYVGKRCVYVYKAETK 101
> Hs17475088
Length=138
Score = 37.4 bits (85), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 23/72 (31%), Positives = 36/72 (50%), Gaps = 3/72 (4%)
Query 8 RQMRP---KRGGQQRAPNSRYGLFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFY 64
R+M+P ++ G + G + I GYK+ NQ LL +E++ + + F
Sbjct 11 RKMKPVYRQQDGTSKRQICLEGCGPKAIFAGYKQGLWNQREHTALLKIEDVYAQDETEFC 70
Query 65 RGKRVAYFYRAK 76
GKR AY Y+AK
Sbjct 71 LGKRSAYLYKAK 82
> SPCP31B10.08c
Length=108
Score = 37.0 bits (84), Expect = 0.009, Method: Compositional matrix adjust.
Identities = 18/65 (27%), Positives = 33/65 (50%), Gaps = 0/65 (0%)
Query 21 PNSRYGLFFRGIVLGYKRSRVNQTPPPPLLHLENLKTRKDPPFYRGKRVAYFYRAKTKNR 80
P + L+ + L ++RS+ P ++ +E ++++ FY GKRV Y Y++ R
Sbjct 2 PAQGHRLYVKAKHLSFQRSKHVIHPGTSIVKIEGCDSKEEAQFYLGKRVCYVYKSSKAVR 61
Query 81 GKIFR 85
G R
Sbjct 62 GSKIR 66
> SPBC1921.01c
Length=79
Score = 28.5 bits (62), Expect = 3.4, Method: Compositional matrix adjust.
Identities = 10/37 (27%), Positives = 21/37 (56%), Gaps = 0/37 (0%)
Query 49 LLHLENLKTRKDPPFYRGKRVAYFYRAKTKNRGKIFR 85
++ +E ++++ FY GKR+ + Y++ RG R
Sbjct 1 MVKIEGCDSKEEAQFYLGKRICFVYKSNKPVRGSKIR 37
> Hs17483554
Length=336
Score = 28.1 bits (61), Expect = 3.9, Method: Composition-based stats.
Identities = 12/20 (60%), Positives = 13/20 (65%), Gaps = 0/20 (0%)
Query 43 QTPPPPLLHLENLKTRKDPP 62
QTPP PLL LEN +T P
Sbjct 278 QTPPSPLLELENKQTTHSLP 297
> At5g42950
Length=1714
Score = 26.9 bits (58), Expect = 8.1, Method: Composition-based stats.
Identities = 10/20 (50%), Positives = 13/20 (65%), Gaps = 0/20 (0%)
Query 6 RWRQMRPKRGGQQRAPNSRY 25
RW + PK G Q+R PN R+
Sbjct 134 RWDNVAPKFGEQRRGPNDRW 153
> Hs20545012
Length=277
Score = 26.9 bits (58), Expect = 9.1, Method: Compositional matrix adjust.
Identities = 10/34 (29%), Positives = 21/34 (61%), Gaps = 0/34 (0%)
Query 7 WRQMRPKRGGQQRAPNSRYGLFFRGIVLGYKRSR 40
W+Q+RP+R + P +++G F + +G K+ +
Sbjct 163 WKQVRPERTKEDVLPKAKFGSDFMLLGIGEKKMK 196
Lambda K H
0.326 0.142 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1194132014
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40