bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
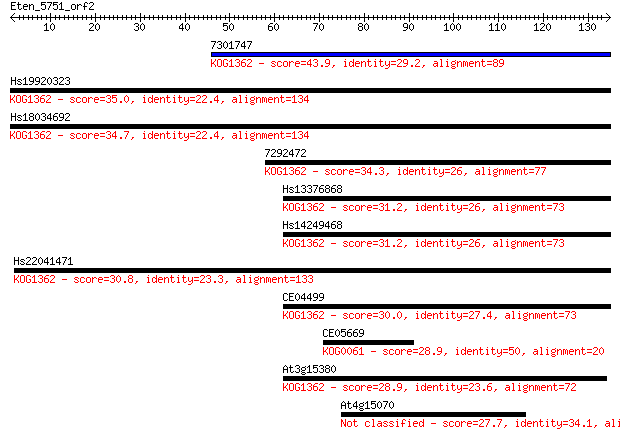
Query= Eten_5751_orf2
Length=134
Score E
Sequences producing significant alignments: (Bits) Value
7301747 43.9 9e-05
Hs19920323 35.0 0.039
Hs18034692 34.7 0.047
7292472 34.3 0.063
Hs13376868 31.2 0.54
Hs14249468 31.2 0.56
Hs22041471 30.8 0.68
CE04499 30.0 1.1
CE05669 28.9 2.6
At3g15380 28.9 3.1
At4g15070 27.7 6.5
> 7301747
Length=796
Score = 43.9 bits (102), Expect = 9e-05, Method: Composition-based stats.
Identities = 26/89 (29%), Positives = 46/89 (51%), Gaps = 10/89 (11%)
Query 46 EFVWDVRFVFFGALWWFALFWVIEAMMAFAHFVISYSATVWYFSPPEGADERDVGWFPPL 105
E W + + FG LW + F + AF++ V++ + WY++ +RDV +F
Sbjct 533 EIKWAIFYNVFGFLW-LSFF-----ISAFSYMVLASTFARWYWT----FKKRDVPYFTLT 582
Query 106 VAMGLGSIHHMGSFAVGGLVMGATRPIRL 134
A +++H+G+ A G L++ R IRL
Sbjct 583 RAFFQTAVYHLGTVAFGSLILAIVRLIRL 611
> Hs19920323
Length=654
Score = 35.0 bits (79), Expect = 0.039, Method: Composition-based stats.
Identities = 30/140 (21%), Positives = 61/140 (43%), Gaps = 31/140 (22%)
Query 1 AATIVWVWLWLWSYVYMASAGSAGATEIPMGLEQNGDIGILPLHREFVWDVRFVFFGAL- 59
A + WV+ W+ + +++ + GS P+ EQ V F G L
Sbjct 344 ALVLFWVY-WIMTLLFLGTTGS------PVQNEQGF--------------VEFKISGPLQ 382
Query 60 --WWF---ALFWVIEAMMAFAHFVISYSATVWYFSPPEGADERDVGWFPPLVAMGLGSIH 114
WW+ L W+ E ++A ++ + +YF+ D+R++ + P L ++ +
Sbjct 383 YMWWYHVVGLIWISEFILACQQMTVAGAVVTYYFT----RDKRNLPFTPILASVNRLIRY 438
Query 115 HMGSFAVGGLVMGATRPIRL 134
H+G+ A G ++ + R+
Sbjct 439 HLGTVAKGSFIITLVKIPRM 458
> Hs18034692
Length=657
Score = 34.7 bits (78), Expect = 0.047, Method: Composition-based stats.
Identities = 30/140 (21%), Positives = 61/140 (43%), Gaps = 31/140 (22%)
Query 1 AATIVWVWLWLWSYVYMASAGSAGATEIPMGLEQNGDIGILPLHREFVWDVRFVFFGAL- 59
A + WV+ W+ + +++ + GS P+ EQ V F G L
Sbjct 344 ALVLFWVY-WIMTLLFLGTTGS------PVQNEQGF--------------VEFKISGPLQ 382
Query 60 --WWF---ALFWVIEAMMAFAHFVISYSATVWYFSPPEGADERDVGWFPPLVAMGLGSIH 114
WW+ L W+ E ++A ++ + +YF+ D+R++ + P L ++ +
Sbjct 383 YMWWYHVVGLIWISEFILACQQMTVAGAVVTYYFT----RDKRNLPFTPILASVNRLIRY 438
Query 115 HMGSFAVGGLVMGATRPIRL 134
H+G+ A G ++ + R+
Sbjct 439 HLGTVAKGSFIITLVKIPRM 458
> 7292472
Length=691
Score = 34.3 bits (77), Expect = 0.063, Method: Composition-based stats.
Identities = 20/80 (25%), Positives = 34/80 (42%), Gaps = 11/80 (13%)
Query 58 ALWWF---ALFWVIEAMMAFAHFVISYSATVWYFSPPEGADERDVGWFPPLVAMGLGSIH 114
+++W L W +E + A F ++ + WYF P P A+G +
Sbjct 408 SMFWIYVVGLIWTVEFIFACQQFALAAAVAFWYFQKPTST--------PTFYAIGKLVKY 459
Query 115 HMGSFAVGGLVMGATRPIRL 134
H+G+ A G V+ + RL
Sbjct 460 HLGTVAKGSFVITIFKIPRL 479
> Hs13376868
Length=712
Score = 31.2 bits (69), Expect = 0.54, Method: Composition-based stats.
Identities = 19/75 (25%), Positives = 35/75 (46%), Gaps = 7/75 (9%)
Query 62 FALFWVIEAMMAFAHFVIS--YSATVWYFSPPEGADERDVGWFPPLVAMGLGSIHHMGSF 119
LFW + ++A V++ +++ W F P+ D+ FP + A +H GS
Sbjct 450 LGLFWTLNWVLALGQCVLAGAFASFYWAFHKPQ-----DIPTFPLISAFIRTLRYHTGSL 504
Query 120 AVGGLVMGATRPIRL 134
A G L++ + R+
Sbjct 505 AFGALILTLVQIARV 519
> Hs14249468
Length=710
Score = 31.2 bits (69), Expect = 0.56, Method: Composition-based stats.
Identities = 19/75 (25%), Positives = 35/75 (46%), Gaps = 7/75 (9%)
Query 62 FALFWVIEAMMAFAHFVIS--YSATVWYFSPPEGADERDVGWFPPLVAMGLGSIHHMGSF 119
LFW + ++A V++ +++ W F P+ D+ FP + A +H GS
Sbjct 448 LGLFWTLNWVLALGQCVLAGAFASFYWAFHKPQ-----DIPTFPLISAFIRTLRYHTGSL 502
Query 120 AVGGLVMGATRPIRL 134
A G L++ + R+
Sbjct 503 AFGALILTLVQIARV 517
> Hs22041471
Length=605
Score = 30.8 bits (68), Expect = 0.68, Method: Composition-based stats.
Identities = 31/139 (22%), Positives = 57/139 (41%), Gaps = 29/139 (20%)
Query 2 ATIVWVWLWLWSYVYMASAGSAGATEIPMGLEQNGDIGILPLHR-EFVWDVRFVFFGALW 60
A +++ W+ LW V + S G+AGA ++ G G + PL ++W +
Sbjct 290 AILIFFWV-LWVAVLL-SLGTAGAAQVMEG----GQVEYKPLSGIRYMWSYHLI------ 337
Query 61 WFALFWVIEAMMAFAHFVISYSATVWYFS-----PPEGADERDVGWFPPLVAMGLGSIHH 115
L W E ++A I+ + YF+ PP+ P L ++ + +H
Sbjct 338 --GLIWTSEFILACQQMTIAGAVVTCYFNRSKNDPPD---------HPILSSLSILFFYH 386
Query 116 MGSFAVGGLVMGATRPIRL 134
G+ G ++ R R+
Sbjct 387 QGTVVKGSFLISVVRIPRI 405
> CE04499
Length=771
Score = 30.0 bits (66), Expect = 1.1, Method: Composition-based stats.
Identities = 20/74 (27%), Positives = 38/74 (51%), Gaps = 5/74 (6%)
Query 62 FALFWVIEAMMAFAHFVISYSATVWYFSPPEGADER-DVGWFPPLVAMGLGSIHHMGSFA 120
FA FW+ + A ++ + +Y++ D+R DV FP + A+ +++GS A
Sbjct 515 FAFFWLSCFVTALGDIALAGAFASYYWA----RDKRHDVPTFPVIRALNRAIRYNLGSIA 570
Query 121 VGGLVMGATRPIRL 134
G L++ + IR+
Sbjct 571 FGSLIIAIVKIIRV 584
> CE05669
Length=695
Score = 28.9 bits (63), Expect = 2.6, Method: Composition-based stats.
Identities = 10/20 (50%), Positives = 13/20 (65%), Gaps = 0/20 (0%)
Query 71 MMAFAHFVISYSATVWYFSP 90
+M F F+I+Y A WYF P
Sbjct 585 LMVFGGFMITYEAIPWYFLP 604
> At3g15380
Length=722
Score = 28.9 bits (63), Expect = 3.1, Method: Compositional matrix adjust.
Identities = 17/72 (23%), Positives = 37/72 (51%), Gaps = 2/72 (2%)
Query 62 FALFWVIEAMMAFAHFVISYSATVWYFSPPEGADERDVGWFPPLVAMGLGSIHHMGSFAV 121
F +W + +A + VI+ S +Y++ E + E + + P +M + +++GS A+
Sbjct 442 FGCYWATQFFIASSATVIAGSVASYYWAQGEASPE--IPFLPVFASMKRLARYNLGSVAL 499
Query 122 GGLVMGATRPIR 133
G L++ +R
Sbjct 500 GSLIVSFVESVR 511
> At4g15070
Length=889
Score = 27.7 bits (60), Expect = 6.5, Method: Composition-based stats.
Identities = 14/41 (34%), Positives = 18/41 (43%), Gaps = 0/41 (0%)
Query 75 AHFVISYSATVWYFSPPEGADERDVGWFPPLVAMGLGSIHH 115
AH + + VW EG E D+ P V + GSI H
Sbjct 607 AHSKCATQSNVWDGKELEGEPEEDINEIEPFVTLRDGSIRH 647
Lambda K H
0.329 0.142 0.510
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1393086308
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40