bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_6487_orf1
Length=124
Score E
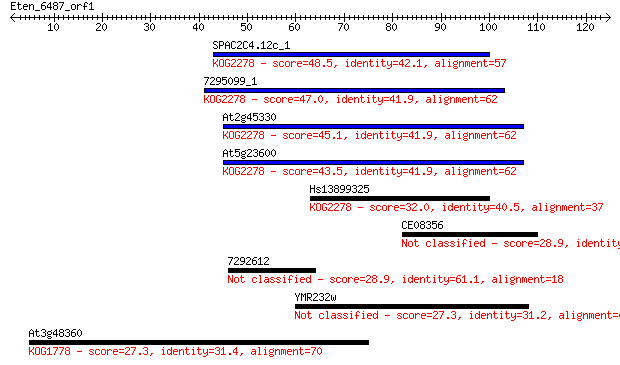
Sequences producing significant alignments: (Bits) Value
SPAC2C4.12c_1 48.5 3e-06
7295099_1 47.0 9e-06
At2g45330 45.1 3e-05
At5g23600 43.5 9e-05
Hs13899325 32.0 0.26
CE08356 28.9 2.2
7292612 28.9 2.2
YMR232w 27.3 6.2
At3g48360 27.3 7.1
> SPAC2C4.12c_1
Length=225
Score = 48.5 bits (114), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 24/57 (42%), Positives = 35/57 (61%), Gaps = 0/57 (0%)
Query 43 KKLNFILRHGASLFRLRIREDGFVRVRDLLALPCMRSVTWNDLKSAVEGNFKRRYEL 99
K L+ +LRH A L+IREDG++ V +L LP R + L S V+GN K+R+ +
Sbjct 39 KALSKVLRHTAKANGLQIREDGYIEVDSILKLPQFRGMGMELLLSIVKGNDKKRFTM 95
> 7295099_1
Length=245
Score = 47.0 bits (110), Expect = 9e-06, Method: Compositional matrix adjust.
Identities = 26/62 (41%), Positives = 34/62 (54%), Gaps = 0/62 (0%)
Query 41 LIKKLNFILRHGASLFRLRIREDGFVRVRDLLALPCMRSVTWNDLKSAVEGNFKRRYELV 100
L KKL+++LRHGA + IR DGFV V DL P T LK + K+RY L
Sbjct 11 LSKKLSWLLRHGAKTEGITIRADGFVSVPDLQKHPRYLCFTLEKLKEIAAADAKQRYTLR 70
Query 101 YD 102
++
Sbjct 71 WN 72
> At2g45330
Length=257
Score = 45.1 bits (105), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 26/69 (37%), Positives = 41/69 (59%), Gaps = 7/69 (10%)
Query 45 LNFILRHGASLFRLRIREDGFVRVRDLLALPC-------MRSVTWNDLKSAVEGNFKRRY 97
L ILRH A+ RL +R DGFV+V DLL L ++S T ++++ AV + K+R+
Sbjct 59 LTRILRHMATELRLNMRGDGFVKVEDLLNLNLKTSANIQLKSHTIDEIREAVRRDNKQRF 118
Query 98 ELVYDKTEV 106
L+ + E+
Sbjct 119 SLIDENGEL 127
> At5g23600
Length=212
Score = 43.5 bits (101), Expect = 9e-05, Method: Compositional matrix adjust.
Identities = 26/69 (37%), Positives = 40/69 (57%), Gaps = 7/69 (10%)
Query 45 LNFILRHGASLFRLRIREDGFVRVRDLLALPC-------MRSVTWNDLKSAVEGNFKRRY 97
L ILRH A+ RL +R DGFV+V DLL L + S T ++++ AV + K+R+
Sbjct 14 LTRILRHMATELRLNMRGDGFVKVEDLLNLNLKTCANIQLNSHTIDEIREAVTRDNKKRF 73
Query 98 ELVYDKTEV 106
L+ + E+
Sbjct 74 SLIDEDGEL 82
> Hs13899325
Length=204
Score = 32.0 bits (71), Expect = 0.26, Method: Compositional matrix adjust.
Identities = 15/37 (40%), Positives = 22/37 (59%), Gaps = 0/37 (0%)
Query 63 DGFVRVRDLLALPCMRSVTWNDLKSAVEGNFKRRYEL 99
DGFV + LL LP R + D++ V+ N K+R+ L
Sbjct 4 DGFVPLGTLLQLPQFRGFSAEDVQRVVDTNRKQRFAL 40
> CE08356
Length=577
Score = 28.9 bits (63), Expect = 2.2, Method: Composition-based stats.
Identities = 13/28 (46%), Positives = 14/28 (50%), Gaps = 0/28 (0%)
Query 82 WNDLKSAVEGNFKRRYELVYDKTEVETC 109
WN L + E N KRR YD EV C
Sbjct 318 WNGLPHSSEWNPKRRARFFYDDLEVVDC 345
> 7292612
Length=487
Score = 28.9 bits (63), Expect = 2.2, Method: Composition-based stats.
Identities = 11/18 (61%), Positives = 13/18 (72%), Gaps = 0/18 (0%)
Query 46 NFILRHGASLFRLRIRED 63
NF LRHG SLF ++ED
Sbjct 255 NFCLRHGTSLFYTSVKED 272
> YMR232w
Length=677
Score = 27.3 bits (59), Expect = 6.2, Method: Composition-based stats.
Identities = 15/63 (23%), Positives = 26/63 (41%), Gaps = 15/63 (23%)
Query 60 IREDGFVRVRDLLALPCMRSVTWNDLKSAVEGNF---------------KRRYELVYDKT 104
++E G +++ D+L P R W D +E + +R+Y L +K
Sbjct 266 LKESGCIKLEDILKSPMKRLTQWIDTLETLESCYEDILSPELGLKLSPTRRKYSLFSNKL 325
Query 105 EVE 107
E E
Sbjct 326 ETE 328
> At3g48360
Length=367
Score = 27.3 bits (59), Expect = 7.1, Method: Compositional matrix adjust.
Identities = 22/70 (31%), Positives = 30/70 (42%), Gaps = 7/70 (10%)
Query 5 GCTAPGGRAAADEDSSSSSNPDSSCCCAFSARGGYILIKKLNFILRHGASLFRLRIREDG 64
GCT G D + S + S C AFS G L ++RH A R R + G
Sbjct 237 GCTLVGPSNVVDNNKKSMTAEKSEPCKAFSTCYG------LQLLIRHFAVCKR-RNNDKG 289
Query 65 FVRVRDLLAL 74
+R + +L L
Sbjct 290 CLRCKRMLQL 299
Lambda K H
0.321 0.133 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1187579072
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40