bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
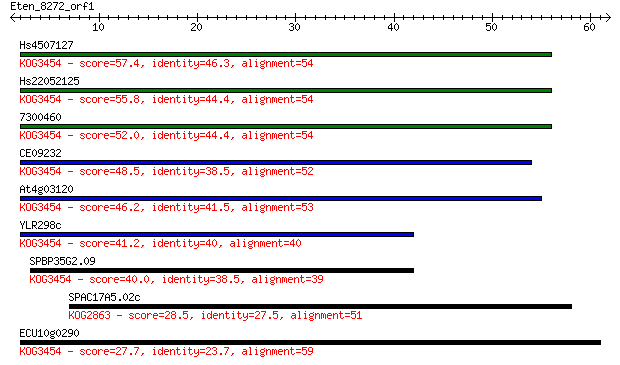
Query= Eten_8272_orf1
Length=61
Score E
Sequences producing significant alignments: (Bits) Value
Hs4507127 57.4 7e-09
Hs22052125 55.8 2e-08
7300460 52.0 3e-07
CE09232 48.5 3e-06
At4g03120 46.2 2e-05
YLR298c 41.2 5e-04
SPBP35G2.09 40.0 0.001
SPAC17A5.02c 28.5 3.3
ECU10g0290 27.7 4.7
> Hs4507127
Length=159
Score = 57.4 bits (137), Expect = 7e-09, Method: Compositional matrix adjust.
Identities = 25/54 (46%), Positives = 34/54 (62%), Gaps = 3/54 (5%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQAVA 55
YC+YCD YLTH SP+ R+ H SGR+H +Y+Q + E QA + ID+ A
Sbjct 5 YCDYCDTYLTHDSPSVRKTHCSGRKHKENVKDYYQKWMEE---QAQSLIDKTTA 55
> Hs22052125
Length=155
Score = 55.8 bits (133), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 24/54 (44%), Positives = 34/54 (62%), Gaps = 3/54 (5%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQAVA 55
YC+YC+ YLTH SP R+ H SGR+H +Y+Q ++E QA + ID+ A
Sbjct 5 YCDYCNTYLTHDSPPVRKTHCSGRKHKENVKDYYQKWMKE---QAQSLIDKTTA 55
> 7300460
Length=145
Score = 52.0 bits (123), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/54 (44%), Positives = 32/54 (59%), Gaps = 3/54 (5%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQAVA 55
YC+YCD YLTH SP+ R+ H +GR+H + Y+Q + E QA ID A
Sbjct 5 YCDYCDTYLTHDSPSVRKTHCTGRKHRDNVKFYYQKWMEE---QAQHLIDATTA 55
> CE09232
Length=142
Score = 48.5 bits (114), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 20/52 (38%), Positives = 31/52 (59%), Gaps = 3/52 (5%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQA 53
YC+YCD +LTH SP+ R+ H GR+H + ++Q + + QA +DQ
Sbjct 5 YCDYCDTFLTHDSPSVRKTHNGGRKHKDNVRMFYQKWMED---QAQKLVDQT 53
> At4g03120
Length=112
Score = 46.2 bits (108), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 22/53 (41%), Positives = 31/53 (58%), Gaps = 3/53 (5%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQAV 54
YC+YCD YLTH SP+ R+QH +G +H Y+Q + Q + IDQ +
Sbjct 5 YCDYCDTYLTHDSPSVRKQHNAGYKHKANVRIYYQQFEEQ---QTQSLIDQRI 54
> YLR298c
Length=231
Score = 41.2 bits (95), Expect = 5e-04, Method: Composition-based stats.
Identities = 16/40 (40%), Positives = 25/40 (62%), Gaps = 0/40 (0%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRE 41
YCEYC YLTH + + R+ H G+ H+ +Y++N R+
Sbjct 5 YCEYCHSYLTHDTLSVRKSHLVGKNHLRITADYYRNKARD 44
> SPBP35G2.09
Length=182
Score = 40.0 bits (92), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 15/39 (38%), Positives = 25/39 (64%), Gaps = 0/39 (0%)
Query 3 CEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRE 41
C+YC ++LTH S + R+ H +GR HI +Y+ + +E
Sbjct 6 CDYCQVWLTHDSQSVRKAHNAGRAHIQNVQDYYTKVAQE 44
> SPAC17A5.02c
Length=466
Score = 28.5 bits (62), Expect = 3.3, Method: Composition-based stats.
Identities = 14/51 (27%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query 7 DIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQAVAMR 57
DI+L+H P G QH + + K +F+N + + +PA + V ++
Sbjct 178 DIFLSHDWPRGIEQHGDVAKLLRHK-PFFRNEVERNDLGSPALEELLVELK 227
> ECU10g0290
Length=122
Score = 27.7 bits (60), Expect = 4.7, Method: Compositional matrix adjust.
Identities = 14/59 (23%), Positives = 26/59 (44%), Gaps = 4/59 (6%)
Query 2 YCEYCDIYLTHSSPAGRRQHASGRRHINQKVEYFQNLLRESTFQAPAFIDQAVAMRAIQ 60
+CE+C+ L + RR H G +H + Y+ + + A + A +R I+
Sbjct 5 FCEFCNKMLLNDRLRSRRMHCGGAKHGLMRKAYYMEMFEKEDIAA----EMASILRDIK 59
Lambda K H
0.326 0.135 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1178852562
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40