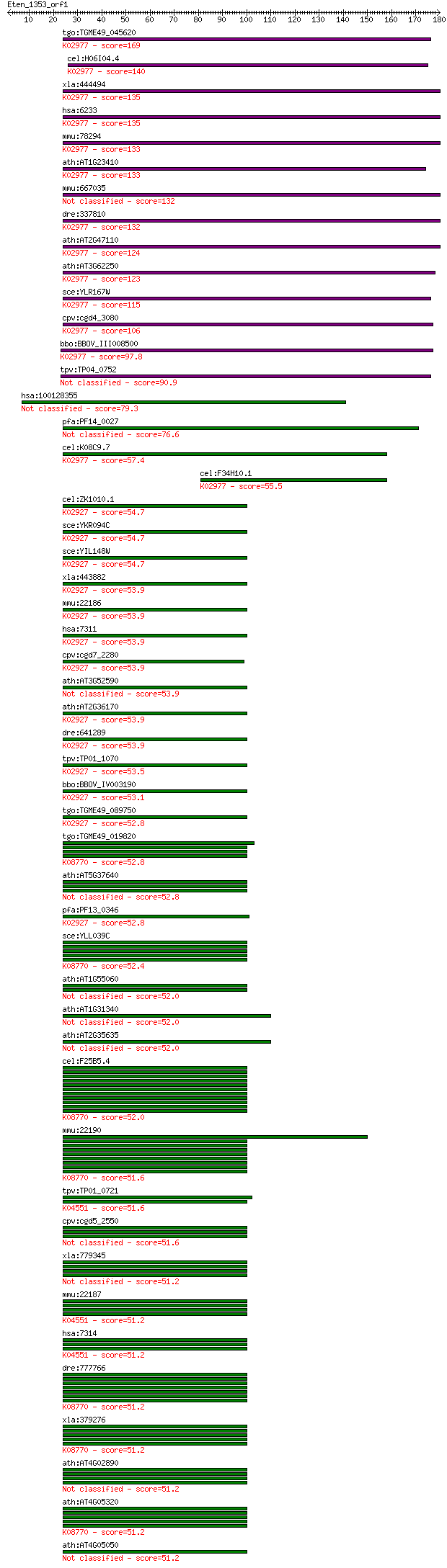
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_1353_orf1
Length=179
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_045620 ubiquitin / ribosomal protein S27a fusion pr... 169 5e-42
cel:H06I04.4 ubl-1; UBiquitin-Like family member (ubl-1); K029... 140 3e-33
xla:444494 rps27a, MGC81889; ribosomal protein S27a; K02977 sm... 135 1e-31
hsa:6233 RPS27A, CEP80, UBA80, UBCEP1, UBCEP80; ribosomal prot... 135 1e-31
mmu:78294 Rps27a, 0610006J14Rik, Uba52, Ubb, Ubc; ribosomal pr... 133 4e-31
ath:AT1G23410 ubiquitin extension protein, putative / 40S ribo... 133 4e-31
mmu:667035 Gm8430, EG667035; predicted pseudogene 8430 132 5e-31
dre:337810 rps27a, MGC66168, hm:zeh0386, ik:tdsubc_1f2, tdsubc... 132 6e-31
ath:AT2G47110 UBQ6; UBQ6; protein binding; K02977 small subuni... 124 1e-28
ath:AT3G62250 UBQ5; UBQ5 (ubiquitin 5); protein binding / stru... 123 4e-28
sce:YLR167W RPS31, RPS37, UBI3; Rps31p; K02977 small subunit r... 115 7e-26
cpv:cgd4_3080 ribosomal protein S27a, ubiquitin plus zincribbo... 106 4e-23
bbo:BBOV_III008500 17.m07744; ribosomal protein S27a family pr... 97.8 2e-20
tpv:TP04_0752 hypothetical protein 90.9 2e-18
hsa:100128355 similar to hCG1790904 79.3 6e-15
pfa:PF14_0027 40S ribosomal protein S31/UBI, putative 76.6 4e-14
cel:K08C9.7 hypothetical protein; K02977 small subunit ribosom... 57.4 2e-08
cel:F34H10.1 hypothetical protein; K02977 small subunit riboso... 55.5 1e-07
cel:ZK1010.1 ubq-2; UBiQuitin family member (ubq-2); K02927 la... 54.7 1e-07
sce:YKR094C RPL40B, CEP52B, UB12, UBI2; L40B; K02927 large sub... 54.7 2e-07
sce:YIL148W RPL40A, CEP52A, UB11, UBI1; L40A; K02927 large sub... 54.7 2e-07
xla:443882 uba52, MGC80109; ubiquitin A-52 residue ribosomal p... 53.9 3e-07
mmu:22186 Uba52, D8Ertd21e, Gm1863, MGC107645, Rps27a, Ubb, Ub... 53.9 3e-07
hsa:7311 UBA52, CEP52, HUBCEP52, MGC126879, MGC126881, MGC5712... 53.9 3e-07
cpv:cgd7_2280 60S ribosomal protein L40 ; K02927 large subunit... 53.9 3e-07
ath:AT3G52590 UBQ1; UBQ1 (UBIQUITIN EXTENSION PROTEIN 1); prot... 53.9 3e-07
ath:AT2G36170 ubiquitin extension protein 2 (UBQ2) / 60S ribos... 53.9 3e-07
dre:641289 uba52, MGC123344, wu:fa91f08, zgc:123344; ubiquitin... 53.9 3e-07
tpv:TP01_1070 ubiquitin/ribosomal fusion protein; K02927 large... 53.5 3e-07
bbo:BBOV_IV003190 21.m02806; ubiquitin / ribosomal protein CEP... 53.1 4e-07
tgo:TGME49_089750 ubiquitin / ribosomal protein CEP52 fusion p... 52.8 6e-07
tgo:TGME49_019820 polyubiquitin, putative ; K08770 ubiquitin C 52.8 6e-07
ath:AT5G37640 UBQ9; UBQ9; protein binding 52.8 6e-07
pfa:PF13_0346 60S ribosomal protein L40/UBI, putative; K02927 ... 52.8 7e-07
sce:YLL039C UBI4, SCD2, UB14; Ubi4p; K08770 ubiquitin C 52.4 9e-07
ath:AT1G55060 UBQ12; UBQ12 (UBIQUITIN 12); protein binding 52.0 9e-07
ath:AT1G31340 RUB1; RUB1 (RELATED TO UBIQUITIN 1); protein bin... 52.0 1e-06
ath:AT2G35635 UBQ7; UBQ7; protein binding 52.0 1e-06
cel:F25B5.4 ubq-1; UBiQuitin family member (ubq-1); K08770 ubi... 52.0 1e-06
mmu:22190 Ubc, 2700054O04Rik, AI194771, Rps27a, TI-225, Uba52,... 51.6 1e-06
tpv:TP01_0721 ubiquitin; K04551 ubiquitin B 51.6 1e-06
cpv:cgd5_2550 polyubiquitin with 3 Ub domains 51.6 1e-06
xla:779345 hypothetical protein MGC154789 51.2 2e-06
mmu:22187 Ubb, AL033289, Rps27a, Uba52, Ubb2, Ubc; ubiquitin B... 51.2 2e-06
hsa:7314 UBB, FLJ25987, MGC8385, RPS27A, UBA52, UBC; ubiquitin... 51.2 2e-06
dre:777766 ubc, im:7042025, wu:fb37h06, zgc:153686; ubiquitin ... 51.2 2e-06
xla:379276 MGC53081; similar to ubiquitin C; K08770 ubiquitin C 51.2 2e-06
ath:AT4G02890 UBQ14; UBQ14; protein binding 51.2 2e-06
ath:AT4G05320 UBQ10; UBQ10 (POLYUBIQUITIN 10); protein binding... 51.2 2e-06
ath:AT4G05050 UBQ11; UBQ11 (UBIQUITIN 11); protein binding 51.2 2e-06
> tgo:TGME49_045620 ubiquitin / ribosomal protein S27a fusion
protein, putative ; K02977 small subunit ribosomal protein S27Ae
Length=154
Score = 169 bits (428), Expect = 5e-42, Method: Compositional matrix adjust.
Identities = 109/152 (71%), Positives = 129/152 (84%), Gaps = 0/152 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV S G++L +EV+ TST+ ++K+ L+AR G+ A QRL FG+ A++D TL E G
Sbjct 1 MQIFVLSVGGDTLALEVAPTSTIRDVKEQLQARQGIEADDQRLCFGLHALEDDETLGELG 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
VEEE TL+QSLELLGAGKKRK+KTYTKPKKQKHK KKVKLAVLKFYKVD N+KVTRLRKE
Sbjct 61 VEEECTLYQSLELLGAGKKRKKKTYTKPKKQKHKKKKVKLAVLKFYKVDGNDKVTRLRKE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNA 175
CP +TCG+GVFMA H+NR YCGRCG+TYILNA
Sbjct 121 CPRETCGAGVFMAAHKNRTYCGRCGLTYILNA 152
> cel:H06I04.4 ubl-1; UBiquitin-Like family member (ubl-1); K02977
small subunit ribosomal protein S27Ae
Length=163
Score = 140 bits (353), Expect = 3e-33, Method: Compositional matrix adjust.
Identities = 77/149 (51%), Positives = 106/149 (71%), Gaps = 5/149 (3%)
Query 26 IFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAGVE 85
+FV++ +L +EV+A VL +KQ +EA G+ A QRL + ++D+ + G++
Sbjct 2 VFVKTLH-RTLFLEVAANEDVLSIKQKIEAAEGIPAEEQRLCYAGRQLEDS----DCGID 56
Query 86 EEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKECP 145
EAT++ +LELLG KKRK+K YT PKK K K KKVKLAVLK+YK+D+N K+TRLRKEC
Sbjct 57 SEATIYVNLELLGGAKKRKKKVYTTPKKNKRKPKKVKLAVLKYYKIDENGKITRLRKECQ 116
Query 146 SKTCGSGVFMAQHENRNYCGRCGVTYILN 174
+CG GVFMAQH NR+YCGRC T +++
Sbjct 117 QPSCGGGVFMAQHANRHYCGRCHDTLVVD 145
> xla:444494 rps27a, MGC81889; ribosomal protein S27a; K02977
small subunit ribosomal protein S27Ae
Length=156
Score = 135 bits (339), Expect = 1e-31, Method: Compositional matrix adjust.
Identities = 76/156 (48%), Positives = 109/156 (69%), Gaps = 0/156 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+K+YT PKK KHK KKVKLAVLK+YKVD+N K++RLR+E
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKSYTTPKKNKHKRKKVKLAVLKYYKVDENGKISRLRRE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGGDQ 179
CPS CG+GVFMA H +R+YCG+C +TY N D+
Sbjct 121 CPSDECGAGVFMASHFDRHYCGKCCLTYCFNKPEDK 156
> hsa:6233 RPS27A, CEP80, UBA80, UBCEP1, UBCEP80; ribosomal protein
S27a; K02977 small subunit ribosomal protein S27Ae
Length=156
Score = 135 bits (339), Expect = 1e-31, Method: Compositional matrix adjust.
Identities = 76/156 (48%), Positives = 109/156 (69%), Gaps = 0/156 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+K+YT PKK KHK KKVKLAVLK+YKVD+N K++RLR+E
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKSYTTPKKNKHKRKKVKLAVLKYYKVDENGKISRLRRE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGGDQ 179
CPS CG+GVFMA H +R+YCG+C +TY N D+
Sbjct 121 CPSDECGAGVFMASHFDRHYCGKCCLTYCFNKPEDK 156
> mmu:78294 Rps27a, 0610006J14Rik, Uba52, Ubb, Ubc; ribosomal
protein S27A; K02977 small subunit ribosomal protein S27Ae
Length=156
Score = 133 bits (334), Expect = 4e-31, Method: Compositional matrix adjust.
Identities = 75/156 (48%), Positives = 108/156 (69%), Gaps = 0/156 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+K+YT PKK KHK KKVKLAVLK+YKVD+N K++RLR+E
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKSYTTPKKNKHKRKKVKLAVLKYYKVDENGKISRLRRE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGGDQ 179
CPS CG+GVFM H +R+YCG+C +TY N D+
Sbjct 121 CPSDECGAGVFMGSHFDRHYCGKCCLTYCFNKPEDK 156
> ath:AT1G23410 ubiquitin extension protein, putative / 40S ribosomal
protein S27A (RPS27aA); K02977 small subunit ribosomal
protein S27Ae
Length=156
Score = 133 bits (334), Expect = 4e-31, Method: Compositional matrix adjust.
Identities = 75/150 (50%), Positives = 106/150 (70%), Gaps = 0/150 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+KTYTKPKK KH +KKVKLAVL+FYKVD + KV RL+KE
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKTYTKPKKIKHTHKKVKLAVLQFYKVDGSGKVQRLKKE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYIL 173
CPS +CG G FMA H +R+YCG+CG TY+
Sbjct 121 CPSVSCGPGTFMASHFDRHYCGKCGTTYVF 150
> mmu:667035 Gm8430, EG667035; predicted pseudogene 8430
Length=156
Score = 132 bits (333), Expect = 5e-31, Method: Compositional matrix adjust.
Identities = 75/156 (48%), Positives = 108/156 (69%), Gaps = 0/156 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIGNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+K+YT PKK KHK KKVKLAVLK+YKVD+N K++RLR+E
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKSYTTPKKNKHKRKKVKLAVLKYYKVDENGKISRLRRE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGGDQ 179
CPS CG+GVFM H +R+YCG+C +TY N D+
Sbjct 121 CPSDECGAGVFMGSHFDRHYCGKCCLTYCFNKPEDK 156
> dre:337810 rps27a, MGC66168, hm:zeh0386, ik:tdsubc_1f2, tdsubc_1f2,
xx:tdsubc_1f2, zgc:66168; ribosomal protein S27a; K02977
small subunit ribosomal protein S27Ae
Length=156
Score = 132 bits (332), Expect = 6e-31, Method: Compositional matrix adjust.
Identities = 75/156 (48%), Positives = 108/156 (69%), Gaps = 0/156 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+K+YT PKK KHK KKVKLAVLK+YKVD+N K+ RLR+E
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKSYTTPKKNKHKRKKVKLAVLKYYKVDENGKIHRLRRE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGGDQ 179
CP+ CG+GVFMA H +R+YCG+C +TY N D+
Sbjct 121 CPADECGAGVFMASHFDRHYCGKCCLTYCFNKPEDK 156
> ath:AT2G47110 UBQ6; UBQ6; protein binding; K02977 small subunit
ribosomal protein S27Ae
Length=157
Score = 124 bits (312), Expect = 1e-28, Method: Compositional matrix adjust.
Identities = 79/156 (50%), Positives = 111/156 (71%), Gaps = 0/156 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+KTYTKPKK KHK+KKVKLAVL+FYKVD + KV RLRKE
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKTYTKPKKIKHKHKKVKLAVLQFYKVDGSGKVQRLRKE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGGDQ 179
CP+ TCG+G FMA H +R+YCG+CG+TY+ G Q
Sbjct 121 CPNATCGAGTFMASHFDRHYCGKCGLTYVYQKEGAQ 156
> ath:AT3G62250 UBQ5; UBQ5 (ubiquitin 5); protein binding / structural
constituent of ribosome; K02977 small subunit ribosomal
protein S27Ae
Length=157
Score = 123 bits (308), Expect = 4e-28, Method: Compositional matrix adjust.
Identities = 77/154 (50%), Positives = 110/154 (71%), Gaps = 0/154 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G KKRK+KTYTKPKK KHK+KKVKLAVL+FYK+D + KV RLRKE
Sbjct 61 IQKESTLHLVLRLRGGAKKRKKKTYTKPKKIKHKHKKVKLAVLQFYKIDGSGKVQRLRKE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAGG 177
CP+ TCG+G FMA H +R+YCG+CG+TY+ G
Sbjct 121 CPNATCGAGTFMASHFDRHYCGKCGLTYVYQKEG 154
> sce:YLR167W RPS31, RPS37, UBI3; Rps31p; K02977 small subunit
ribosomal protein S27Ae
Length=152
Score = 115 bits (289), Expect = 7e-26, Method: Compositional matrix adjust.
Identities = 73/152 (48%), Positives = 106/152 (69%), Gaps = 0/152 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+++E+TL L L G GKKRK+K YT PKK KHK+KKVKLAVL +YKVD KVT+LR+E
Sbjct 61 IQKESTLHLVLRLRGGGKKRKKKVYTTPKKIKHKHKKVKLAVLSYYKVDAEGKVTKLRRE 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNA 175
C + TCG+GVF+A H++R YCG+C Y +NA
Sbjct 121 CSNPTCGAGVFLANHKDRLYCGKCHSVYKVNA 152
> cpv:cgd4_3080 ribosomal protein S27a, ubiquitin plus zincribbon,
UB13p ; K02977 small subunit ribosomal protein S27Ae
Length=159
Score = 106 bits (265), Expect = 4e-23, Method: Compositional matrix adjust.
Identities = 78/153 (50%), Positives = 107/153 (69%), Gaps = 2/153 (1%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIF R G + +EV T +V EL+ + G++ Q + +G +D+ TLE+AG
Sbjct 4 MQIFFRYGLGNTRSLEVDPTMSVKELRHIISEFSGISIDSQCISYGFGILDEFETLEQAG 63
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+ + +TL+ S +LG G K+K+K +TKPKK KHK KKVKLAVLK+YKVD +KV +LR+E
Sbjct 64 ISDYSTLYVSEAMLG-GAKKKKKNFTKPKKIKHKKKKVKLAVLKYYKVD-GDKVVKLRRE 121
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVTYILNAG 176
CP++TCG+GVFMAQH NR CGRCG+TY + G
Sbjct 122 CPAETCGAGVFMAQHFNRTSCGRCGLTYFPSKG 154
> bbo:BBOV_III008500 17.m07744; ribosomal protein S27a family
protein; K02977 small subunit ribosomal protein S27Ae
Length=156
Score = 97.8 bits (242), Expect = 2e-20, Method: Compositional matrix adjust.
Identities = 69/155 (44%), Positives = 101/155 (65%), Gaps = 2/155 (1%)
Query 23 IMQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEA 82
+ Q+F++ + G + +++ + TV ++K+ + G+ Q L+ G+ DD +E+
Sbjct 3 VAQVFIKLFNGNTAVIDLFGSETVADIKRQVAELYGLEGCSQSLWAGLVEADDESLIEDF 62
Query 83 GVEEEAT-LFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLR 141
++E L Q +++ G GKKRK+K +T PKK KHK KKVKL VLK+YKV+ +KV RL
Sbjct 63 ASDDEIIELVQHIDVCGGGKKRKKKQHTTPKKIKHKKKKVKLNVLKYYKVE-GDKVVRLL 121
Query 142 KECPSKTCGSGVFMAQHENRNYCGRCGVTYILNAG 176
KEC S TCG GVFMAQH +R+YCGRCG TY+ A
Sbjct 122 KECDSSTCGRGVFMAQHHDRDYCGRCGTTYLRTAA 156
> tpv:TP04_0752 hypothetical protein
Length=162
Score = 90.9 bits (224), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 72/154 (46%), Positives = 95/154 (61%), Gaps = 2/154 (1%)
Query 23 IMQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEA 82
I Q+ V + G + T+ ++K +L + + L++G++ +DD L E
Sbjct 8 IGQVLVNLFNGRTAVFGSDEYQTIGQIKNYLLGQYTLENINHSLWYGLNELDDELELSEL 67
Query 83 -GVEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLR 141
G E +L LELLG GKKRK+K YT PKK KHK KKVKLAVLK+YKV D ++V RL
Sbjct 68 LGSESTISLDHHLELLGGGKKRKKKQYTTPKKVKHKKKKVKLAVLKYYKV-DGDQVVRLL 126
Query 142 KECPSKTCGSGVFMAQHENRNYCGRCGVTYILNA 175
K+CP + CG GVFMA H NR YCGRC +TY+ A
Sbjct 127 KDCPGENCGRGVFMAAHNNRTYCGRCQLTYLSTA 160
> hsa:100128355 similar to hCG1790904
Length=334
Score = 79.3 bits (194), Expect = 6e-15, Method: Compositional matrix adjust.
Identities = 54/134 (40%), Positives = 81/134 (60%), Gaps = 3/134 (2%)
Query 7 EWRRKGSDICASPWYVIMQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRL 66
+WR GS + S +QIFV++ G+++ +EV + + +K ++ + G+ QRL
Sbjct 54 DWR--GSPVPQSAAATKLQIFVKTLRGKTITLEVEPSDAIENVKAKIQDKEGIPPDQQRL 111
Query 67 FFGVSAVDDACTLEEAGVEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVL 126
F ++D TL + +++E TL L L G KKRK K+Y PKK KHK KKVKL VL
Sbjct 112 IFAGKQLEDGRTLSDCNIQKEPTLHLLLRLCGVAKKRK-KSYPTPKKNKHKRKKVKLDVL 170
Query 127 KFYKVDDNNKVTRL 140
K YKVD+N+K++ L
Sbjct 171 KCYKVDENDKISAL 184
> pfa:PF14_0027 40S ribosomal protein S31/UBI, putative
Length=149
Score = 76.6 bits (187), Expect = 4e-14, Method: Compositional matrix adjust.
Identities = 63/147 (42%), Positives = 89/147 (60%), Gaps = 2/147 (1%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
M+I + ESL +E S + + +K+ + G+ LQ+L+ ++D LE
Sbjct 1 MKILINIPYDESLCLESSNINNIKNVKEQIFELKGIPYELQKLYKNGRHLEDEELLEIDK 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
+ TL + LLG KK+K+K Y KPKK+KHK KKVKLAVLKFYKV D+ KV RL+++
Sbjct 61 SDYAYTLNLNFGLLGGAKKKKKKVYKKPKKEKHKKKKVKLAVLKFYKVGDDGKVFRLKRQ 120
Query 144 CPSKTCGSGVFMAQHENRNYCGRCGVT 170
C + C G MA H +R+YCGRC +T
Sbjct 121 CDN--CAPGTLMASHFDRDYCGRCHLT 145
> cel:K08C9.7 hypothetical protein; K02977 small subunit ribosomal
protein S27Ae
Length=154
Score = 57.4 bits (137), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 40/134 (29%), Positives = 62/134 (46%), Gaps = 12/134 (8%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
+Q+FVR +P +S V +L + QRL + D G
Sbjct 20 LQMFVRL-----MPKNQRKSSIVQKLISPFKDGSTCKEEEQRLCYAAGKGD-------CG 67
Query 84 VEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRLRKE 143
++ AT++ + ELLG KKRK+K Y PKK +K K++K+D+N K++RL
Sbjct 68 IKAAATIYVNCELLGGAKKRKKKVYIIPKKNNISQRKSSSPSPKYFKIDENGKISRLPTG 127
Query 144 CPSKTCGSGVFMAQ 157
P+ + FM Q
Sbjct 128 IPTVSIQREAFMTQ 141
> cel:F34H10.1 hypothetical protein; K02977 small subunit ribosomal
protein S27Ae
Length=142
Score = 55.5 bits (132), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 28/77 (36%), Positives = 44/77 (57%), Gaps = 0/77 (0%)
Query 81 EAGVEEEATLFQSLELLGAGKKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTRL 140
+ G++ AT++ + ELLG KKRK+K YT PKK +K K++K+D+N K++RL
Sbjct 53 DCGIKAAATIYVNCELLGGAKKRKKKVYTIPKKNNISQRKSSSPSPKYFKIDENGKISRL 112
Query 141 RKECPSKTCGSGVFMAQ 157
+ + FM Q
Sbjct 113 PTGILTASIQREAFMTQ 129
> cel:ZK1010.1 ubq-2; UBiQuitin family member (ubq-2); K02927
large subunit ribosomal protein L40e
Length=128
Score = 54.7 bits (130), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> sce:YKR094C RPL40B, CEP52B, UB12, UBI2; L40B; K02927 large subunit
ribosomal protein L40e
Length=128
Score = 54.7 bits (130), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> sce:YIL148W RPL40A, CEP52A, UB11, UBI1; L40A; K02927 large subunit
ribosomal protein L40e
Length=128
Score = 54.7 bits (130), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> xla:443882 uba52, MGC80109; ubiquitin A-52 residue ribosomal
protein fusion product 1; K02927 large subunit ribosomal protein
L40e
Length=128
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> mmu:22186 Uba52, D8Ertd21e, Gm1863, MGC107645, Rps27a, Ubb,
Ubc; ubiquitin A-52 residue ribosomal protein fusion product
1; K02927 large subunit ribosomal protein L40e
Length=128
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> hsa:7311 UBA52, CEP52, HUBCEP52, MGC126879, MGC126881, MGC57125,
RPL40; ubiquitin A-52 residue ribosomal protein fusion
product 1; K02927 large subunit ribosomal protein L40e
Length=128
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> cpv:cgd7_2280 60S ribosomal protein L40 ; K02927 large subunit
ribosomal protein L40e
Length=132
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/75 (32%), Positives = 44/75 (58%), Gaps = 0/75 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 5 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 64
Query 84 VEEEATLFQSLELLG 98
+++E+TL L L G
Sbjct 65 IQKESTLHLVLRLRG 79
> ath:AT3G52590 UBQ1; UBQ1 (UBIQUITIN EXTENSION PROTEIN 1); protein
binding / structural constituent of ribosome
Length=128
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> ath:AT2G36170 ubiquitin extension protein 2 (UBQ2) / 60S ribosomal
protein L40 (RPL40A); K02927 large subunit ribosomal
protein L40e
Length=128
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> dre:641289 uba52, MGC123344, wu:fa91f08, zgc:123344; ubiquitin
A-52 residue ribosomal protein fusion product 1; K02927 large
subunit ribosomal protein L40e
Length=128
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> tpv:TP01_1070 ubiquitin/ribosomal fusion protein; K02927 large
subunit ribosomal protein L40e
Length=131
Score = 53.5 bits (127), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> bbo:BBOV_IV003190 21.m02806; ubiquitin / ribosomal protein CEP52;
K02927 large subunit ribosomal protein L40e
Length=131
Score = 53.1 bits (126), Expect = 4e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> tgo:TGME49_089750 ubiquitin / ribosomal protein CEP52 fusion
protein, putative ; K02927 large subunit ribosomal protein
L40e
Length=129
Score = 52.8 bits (125), Expect = 6e-07, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ ++V + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLDVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> tgo:TGME49_019820 polyubiquitin, putative ; K08770 ubiquitin
C
Length=307
Score = 52.8 bits (125), Expect = 6e-07, Method: Compositional matrix adjust.
Identities = 24/79 (30%), Positives = 46/79 (58%), Gaps = 0/79 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ ++V + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLDVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGAGKK 102
+++E+TL L L G G +
Sbjct 289 IQKESTLHLVLRLRGGGSR 307
Score = 50.4 bits (119), Expect = 4e-06, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ ++V + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLDVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 50.4 bits (119), Expect = 4e-06, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ ++V + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLDVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 50.4 bits (119), Expect = 4e-06, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ ++V + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLDVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
> ath:AT5G37640 UBQ9; UBQ9; protein binding
Length=322
Score = 52.8 bits (125), Expect = 6e-07, Method: Compositional matrix adjust.
Identities = 26/76 (34%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + GV QRL F +DD TL +
Sbjct 79 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGVPPDQQRLIFAGKQLDDGRTLADYN 138
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 139 IQKESTLHLVLRLRGG 154
Score = 47.0 bits (110), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 43/76 (56%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFVR+ +++ +EV ++ T +K ++ + G+ QRL F ++D TL +
Sbjct 155 MQIFVRTLTRKTIALEVESSDTTDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 214
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 215 IQKESTLHLVLRLCGG 230
Score = 38.9 bits (89), Expect = 0.010, Method: Compositional matrix adjust.
Identities = 19/76 (25%), Positives = 41/76 (53%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQI ++ +++ ++V ++ T+ +K ++ G+ QRL F +DD TL +
Sbjct 3 MQIHAKTLTEKTITIDVVSSDTINNVKAKIQDIEGIPLDQQRLIFSGKLLDDGRTLADYS 62
Query 84 VEEEATLFQSLELLGA 99
+++++ L +L L G
Sbjct 63 IQKDSILHLALRLRGG 78
> pfa:PF13_0346 60S ribosomal protein L40/UBI, putative; K02927
large subunit ribosomal protein L40e
Length=128
Score = 52.8 bits (125), Expect = 7e-07, Method: Compositional matrix adjust.
Identities = 23/77 (29%), Positives = 44/77 (57%), Gaps = 0/77 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ ++V + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLDVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGAG 100
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGGA 77
> sce:YLL039C UBI4, SCD2, UB14; Ubi4p; K08770 ubiquitin C
Length=381
Score = 52.4 bits (124), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 52.4 bits (124), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 52.4 bits (124), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 52.4 bits (124), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
Score = 52.4 bits (124), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 305 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 364
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 365 IQKESTLHLVLRLRGG 380
> ath:AT1G55060 UBQ12; UBQ12 (UBIQUITIN 12); protein binding
Length=230
Score = 52.0 bits (123), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ LK ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVESSDTIDNLKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 49.7 bits (117), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G++ QRL F +D TL +
Sbjct 153 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGISPDQQRLIFAGKQHEDGRTLADYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
> ath:AT1G31340 RUB1; RUB1 (RELATED TO UBIQUITIN 1); protein binding
Length=156
Score = 52.0 bits (123), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 29/86 (33%), Positives = 50/86 (58%), Gaps = 1/86 (1%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYT 109
+++E+TL L L G G K KT T
Sbjct 61 IQKESTLHLVLRLRG-GTMIKVKTLT 85
> ath:AT2G35635 UBQ7; UBQ7; protein binding
Length=154
Score = 52.0 bits (123), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 29/86 (33%), Positives = 50/86 (58%), Gaps = 1/86 (1%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGAGKKRKRKTYT 109
+++E+TL L L G G K KT T
Sbjct 61 IQKESTLHLVLRLRG-GTMIKVKTLT 85
> cel:F25B5.4 ubq-1; UBiQuitin family member (ubq-1); K08770 ubiquitin
C
Length=838
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 305 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 364
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 365 IQKESTLHLVLRLRGG 380
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 457 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 516
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 517 IQKESTLHLVLRLRGG 532
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 533 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 592
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 593 IQKESTLHLVLRLRGG 608
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 609 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 668
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 669 IQKESTLHLVLRLRGG 684
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 685 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 744
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 745 IQKESTLHLVLRLRGG 760
Score = 52.0 bits (123), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 761 MQIFVKTLTGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 820
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 821 IQKESTLHLVLRLRGG 836
Score = 51.6 bits (122), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV A+ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 381 MQIFVKTLIGKTITLEVEASDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 440
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 441 IQKESTLHLVLRLRGG 456
> mmu:22190 Ubc, 2700054O04Rik, AI194771, Rps27a, TI-225, Uba52,
Ubb; ubiquitin C; K08770 ubiquitin C
Length=734
Score = 51.6 bits (122), Expect = 1e-06, Method: Composition-based stats.
Identities = 37/130 (28%), Positives = 62/130 (47%), Gaps = 11/130 (8%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 609 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 668
Query 84 VEEEATLFQSLELLGAG----KKRKRKTYTKPKKQKHKNKKVKLAVLKFYKVDDNNKVTR 139
+++E+TL L L G K KT T + KKVK + D +T
Sbjct 669 IQKESTLHLVLRLRGGMQIFVKTLTGKTITLDVEPSVTTKKVK-------QEDRRTFLTT 721
Query 140 LRKECPSKTC 149
+ K+ P C
Sbjct 722 VSKKSPPCAC 731
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 305 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 364
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 365 IQKESTLHLVLRLRGG 380
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 381 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 440
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 441 IQKESTLHLVLRLRGG 456
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 457 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 516
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 517 IQKESTLHLVLRLRGG 532
Score = 50.1 bits (118), Expect = 4e-06, Method: Composition-based stats.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 533 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 592
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 593 IQKESTLHLVLRLRGG 608
> tpv:TP01_0721 ubiquitin; K04551 ubiquitin B
Length=155
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 25/78 (32%), Positives = 45/78 (57%), Gaps = 0/78 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGAGK 101
+++E+TL L L G K
Sbjct 137 IQKESTLHLVLRLRGGYK 154
Score = 51.6 bits (122), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
> cpv:cgd5_2550 polyubiquitin with 3 Ub domains
Length=241
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 13 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 72
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 73 IQKESTLHLVLRLRGG 88
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 89 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 148
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 149 IQKESTLHLVLRLRGG 164
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 165 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 224
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 225 IQKESTLHLVLRLRGG 240
> xla:779345 hypothetical protein MGC154789
Length=305
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
> mmu:22187 Ubb, AL033289, Rps27a, Uba52, Ubb2, Ubc; ubiquitin
B; K04551 ubiquitin B
Length=305
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
> hsa:7314 UBB, FLJ25987, MGC8385, RPS27A, UBA52, UBC; ubiquitin
B; K04551 ubiquitin B
Length=229
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
> dre:777766 ubc, im:7042025, wu:fb37h06, zgc:153686; ubiquitin
C; K08770 ubiquitin C
Length=533
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 305 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 364
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 365 IQKESTLHLVLRLRGG 380
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 381 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 440
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 441 IQKESTLHLVLRLRGG 456
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 457 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 516
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 517 IQKESTLHLVLRLRGG 532
> xla:379276 MGC53081; similar to ubiquitin C; K08770 ubiquitin
C
Length=380
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 44/76 (57%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV + T+ +K ++ + G+ QRL F ++D TL +
Sbjct 305 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 364
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 365 IQKESTLHLVLRLRGG 380
> ath:AT4G02890 UBQ14; UBQ14; protein binding
Length=305
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
> ath:AT4G05320 UBQ10; UBQ10 (POLYUBIQUITIN 10); protein binding;
K08770 ubiquitin C
Length=381
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 77 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 136
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 137 IQKESTLHLVLRLRGG 152
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 153 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 212
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 213 IQKESTLHLVLRLRGG 228
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 229 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 288
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 289 IQKESTLHLVLRLRGG 304
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 305 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 364
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 365 IQKESTLHLVLRLRGG 380
> ath:AT4G05050 UBQ11; UBQ11 (UBIQUITIN 11); protein binding
Length=229
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 24/76 (31%), Positives = 45/76 (59%), Gaps = 0/76 (0%)
Query 24 MQIFVRSYAGESLPVEVSATSTVLELKQHLEARLGVAASLQRLFFGVSAVDDACTLEEAG 83
MQIFV++ G+++ +EV ++ T+ +K ++ + G+ QRL F ++D TL +
Sbjct 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60
Query 84 VEEEATLFQSLELLGA 99
+++E+TL L L G
Sbjct 61 IQKESTLHLVLRLRGG 76
Lambda K H
0.316 0.131 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4795148792
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40