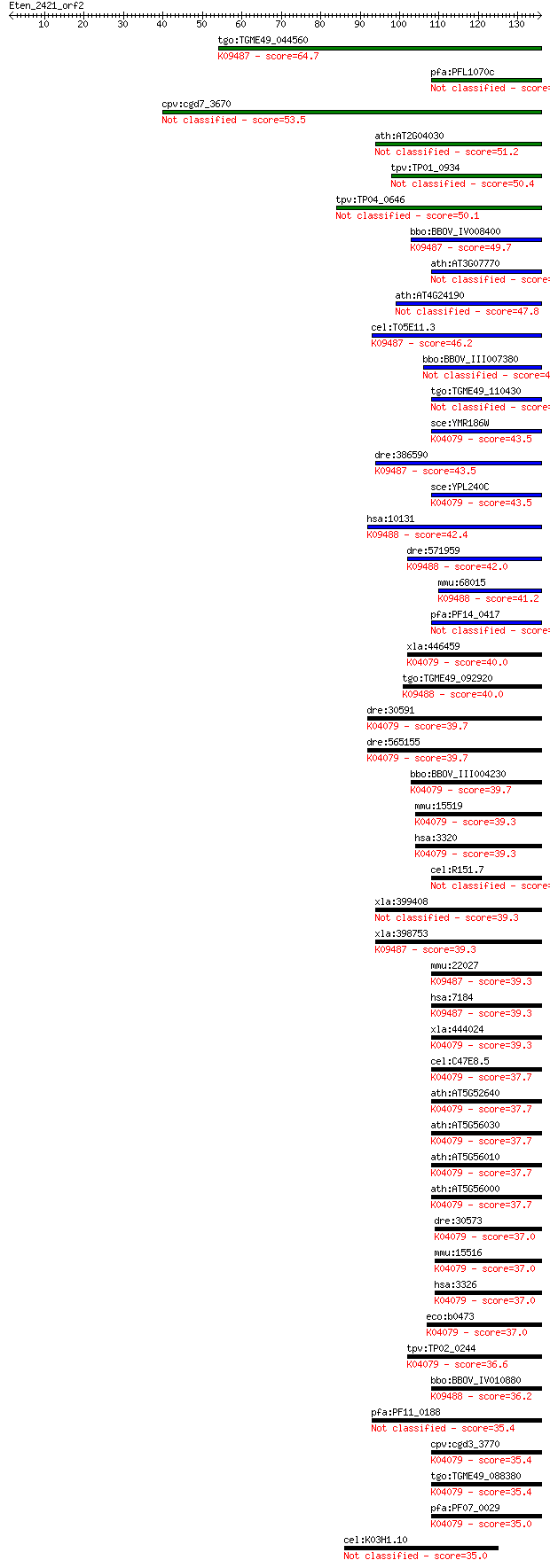
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_2421_orf2
Length=135
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_044560 heat shock protein 90, putative (EC:2.7.13.3... 64.7 8e-11
pfa:PFL1070c endoplasmin homolog precursor, putative 55.1 6e-08
cpv:cgd7_3670 heat shock protein 90 (Hsp90), signal peptide pl... 53.5 2e-07
ath:AT2G04030 CR88; CR88; ATP binding 51.2 8e-07
tpv:TP01_0934 heat shock protein 90 50.4
tpv:TP04_0646 heat shock protein 90 50.1
bbo:BBOV_IV008400 23.m06066; heat shock protein 90; K09487 hea... 49.7 3e-06
ath:AT3G07770 ATP binding 49.7 3e-06
ath:AT4G24190 SHD; SHD (SHEPHERD); ATP binding / unfolded prot... 47.8 1e-05
cel:T05E11.3 hypothetical protein; K09487 heat shock protein 9... 46.2 3e-05
bbo:BBOV_III007380 17.m07646; heat shock protein 90 45.4
tgo:TGME49_110430 heat shock protein 90, putative (EC:3.2.1.3 ... 45.1 7e-05
sce:YMR186W HSC82, HSP90; Cytoplasmic chaperone of the Hsp90 f... 43.5 2e-04
dre:386590 hsp90b1, GP96, GRP94, fb61d09, tra-1, tra1, wu:fb61... 43.5 2e-04
sce:YPL240C HSP82, HSP90; Hsp82p; K04079 molecular chaperone HtpG 43.5 2e-04
hsa:10131 TRAP1, HSP75, HSP90L; TNF receptor-associated protei... 42.4 4e-04
dre:571959 trap1, fc85a11, wu:fc85a11; TNF receptor-associated... 42.0 6e-04
mmu:68015 Trap1, 2410002K23Rik, HSP75; TNF receptor-associated... 41.2 9e-04
pfa:PF14_0417 HSP90 41.2 0.001
xla:446459 hsp90ab1, hsp90b, hsp90beta; heat shock protein 90k... 40.0 0.002
tgo:TGME49_092920 heat shock protein 90, putative ; K09488 TNF... 40.0 0.002
dre:30591 hsp90a.1, fb17b01, hsp90, hsp90a, hsp90alpha, wu:fb1... 39.7 0.002
dre:565155 hsp90a.2, cb820, hsp90a2; heat shock protein 90-alp... 39.7 0.002
bbo:BBOV_III004230 17.m07381; hsp90 protein; K04079 molecular ... 39.7 0.003
mmu:15519 Hsp90aa1, 86kDa, 89kDa, AL024080, AL024147, Hsp86-1,... 39.3 0.003
hsa:3320 HSP90AA1, FLJ31884, HSP86, HSP89A, HSP90A, HSP90N, HS... 39.3 0.003
cel:R151.7 hypothetical protein 39.3 0.003
xla:399408 hsp90b1, ecgp, gp96, grp94, tra1; heat shock protei... 39.3 0.003
xla:398753 hypothetical protein MGC68448; K09487 heat shock pr... 39.3 0.003
mmu:22027 Hsp90b1, ERp99, GRP94, TA-3, Targ2, Tra-1, Tra1, end... 39.3 0.004
hsa:7184 HSP90B1, ECGP, GP96, GRP94, TRA1; heat shock protein ... 39.3 0.004
xla:444024 hsp90aa1.1, MGC82579, hsp86, hsp89, hsp90, hsp90a, ... 39.3 0.004
cel:C47E8.5 daf-21; abnormal DAuer Formation family member (da... 37.7 0.009
ath:AT5G52640 ATHSP90.1 (HEAT SHOCK PROTEIN 90.1); ATP binding... 37.7 0.010
ath:AT5G56030 HSP81-2 (HEAT SHOCK PROTEIN 81-2); ATP binding; ... 37.7 0.010
ath:AT5G56010 HSP81-3; ATP binding / unfolded protein binding;... 37.7 0.010
ath:AT5G56000 heat shock protein 81-4 (HSP81-4); K04079 molecu... 37.7 0.010
dre:30573 hsp90ab1, hsp90b, hsp90beta, wu:fa29f01, wu:fa91e11,... 37.0 0.015
mmu:15516 Hsp90ab1, 90kDa, AL022974, C81438, Hsp84, Hsp84-1, H... 37.0 0.015
hsa:3326 HSP90AB1, D6S182, FLJ26984, HSP90-BETA, HSP90B, HSPC2... 37.0 0.015
eco:b0473 htpG, ECK0467, JW0462; molecular chaperone HSP90 fam... 37.0 0.018
tpv:TP02_0244 heat shock protein 90; K04079 molecular chaperon... 36.6 0.024
bbo:BBOV_IV010880 23.m05763; heat shock protein 75; K09488 TNF... 36.2 0.029
pfa:PF11_0188 heat shock protein 90, putative 35.4 0.055
cpv:cgd3_3770 Hsp90 ; K04079 molecular chaperone HtpG 35.4 0.055
tgo:TGME49_088380 heat shock protein 90 ; K04079 molecular cha... 35.4 0.055
pfa:PF07_0029 heat shock protein 86; K04079 molecular chaperon... 35.0 0.062
cel:K03H1.10 hypothetical protein 35.0 0.070
> tgo:TGME49_044560 heat shock protein 90, putative (EC:2.7.13.3);
K09487 heat shock protein 90kDa beta
Length=847
Score = 64.7 bits (156), Expect = 8e-11, Method: Composition-based stats.
Identities = 37/83 (44%), Positives = 47/83 (56%), Gaps = 1/83 (1%)
Query 54 SAAADPAGASDAAAAEPAAAPAAAAAAAEGAPQGAPQGPLAELAKLRADA-EGHQETHEY 112
SA+ P+ AA AA P A A P + A L +A + QE+H+Y
Sbjct 31 SASVSPSALWVAATETDAAEPLTAEEAPRSLPIDESEKAAAPLTAEEQEAVQKSQESHQY 90
Query 113 QAEVTRLMDIIVNSLYSQREVFL 135
Q EV+RLMDII+NSLY+QREVFL
Sbjct 91 QTEVSRLMDIIINSLYTQREVFL 113
> pfa:PFL1070c endoplasmin homolog precursor, putative
Length=821
Score = 55.1 bits (131), Expect = 6e-08, Method: Composition-based stats.
Identities = 23/28 (82%), Positives = 27/28 (96%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E+H+YQ EVTRLMDIIVNSLY+Q+EVFL
Sbjct 73 ESHQYQTEVTRLMDIIVNSLYTQKEVFL 100
> cpv:cgd7_3670 heat shock protein 90 (Hsp90), signal peptide
plus ER retention motif
Length=787
Score = 53.5 bits (127), Expect = 2e-07, Method: Composition-based stats.
Identities = 35/107 (32%), Positives = 53/107 (49%), Gaps = 11/107 (10%)
Query 40 VVPFCFLGLLLLRCSAAADPAGAS----------DAAAAEPAAAPAAAAAAAEGAPQGAP 89
+V FC + L + P+GA+ + + A + A +E
Sbjct 8 LVFFCLIINLCFGSESTGVPSGATFDESQLGDLNNIDLSSFGANGGSFADESEAVVDSIT 67
Query 90 QGPLAELAKL-RADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
PL EL+ A + E++E+Q EV+RLMDII+NSLYSQ++VFL
Sbjct 68 PAPLPELSNDDEAAIQKTSESYEFQTEVSRLMDIIINSLYSQKDVFL 114
> ath:AT2G04030 CR88; CR88; ATP binding
Length=780
Score = 51.2 bits (121), Expect = 8e-07, Method: Composition-based stats.
Identities = 24/42 (57%), Positives = 31/42 (73%), Gaps = 0/42 (0%)
Query 94 AELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
A +A+ EG E EYQAEV+RL+D+IV+SLYS +EVFL
Sbjct 63 AAVAEKETTEEGSGEKFEYQAEVSRLLDLIVHSLYSHKEVFL 104
> tpv:TP01_0934 heat shock protein 90
Length=1009
Score = 50.4 bits (119), Expect = 2e-06, Method: Composition-based stats.
Identities = 24/38 (63%), Positives = 30/38 (78%), Gaps = 1/38 (2%)
Query 98 KLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
KL D+ E +EYQAEVTRL+DIIVNSLYS +++FL
Sbjct 72 KLFKDS-AKSEKYEYQAEVTRLLDIIVNSLYSSKDIFL 108
> tpv:TP04_0646 heat shock protein 90
Length=913
Score = 50.1 bits (118), Expect = 2e-06, Method: Composition-based stats.
Identities = 25/52 (48%), Positives = 33/52 (63%), Gaps = 1/52 (1%)
Query 84 APQGAPQGPLAELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
A +G P P A G Q T+ +QAEV+R+MDIIVNSLY+ R++FL
Sbjct 108 AYEGDPSTPKAPQEPPEVSLSGEQ-TYPFQAEVSRVMDIIVNSLYTDRDIFL 158
> bbo:BBOV_IV008400 23.m06066; heat shock protein 90; K09487 heat
shock protein 90kDa beta
Length=795
Score = 49.7 bits (117), Expect = 3e-06, Method: Composition-based stats.
Identities = 21/33 (63%), Positives = 26/33 (78%), Gaps = 0/33 (0%)
Query 103 AEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
A H E+H YQA+ R+MDIIVNSLYS ++VFL
Sbjct 84 AAKHGESHTYQADFARVMDIIVNSLYSNKDVFL 116
> ath:AT3G07770 ATP binding
Length=799
Score = 49.7 bits (117), Expect = 3e-06, Method: Composition-based stats.
Identities = 22/28 (78%), Positives = 25/28 (89%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E EYQAEV+RLMD+IVNSLYS +EVFL
Sbjct 95 EKFEYQAEVSRLMDLIVNSLYSNKEVFL 122
> ath:AT4G24190 SHD; SHD (SHEPHERD); ATP binding / unfolded protein
binding
Length=823
Score = 47.8 bits (112), Expect = 1e-05, Method: Composition-based stats.
Identities = 22/37 (59%), Positives = 30/37 (81%), Gaps = 4/37 (10%)
Query 99 LRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
LR++AE E+QAEV+RLMDII+NSLYS +++FL
Sbjct 72 LRSNAE----KFEFQAEVSRLMDIIINSLYSNKDIFL 104
> cel:T05E11.3 hypothetical protein; K09487 heat shock protein
90kDa beta
Length=760
Score = 46.2 bits (108), Expect = 3e-05, Method: Composition-based stats.
Identities = 20/43 (46%), Positives = 32/43 (74%), Gaps = 4/43 (9%)
Query 93 LAELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
++++ +LR+ AE HE+QAEV R+M +I+NSLY +E+FL
Sbjct 51 VSQIKELRSKAE----KHEFQAEVNRMMKLIINSLYRNKEIFL 89
> bbo:BBOV_III007380 17.m07646; heat shock protein 90
Length=795
Score = 45.4 bits (106), Expect = 4e-05, Method: Composition-based stats.
Identities = 17/30 (56%), Positives = 28/30 (93%), Gaps = 0/30 (0%)
Query 106 HQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
+++T+ +QAEV+R+MDIIVNSLY+ +++FL
Sbjct 119 NEQTYPFQAEVSRVMDIIVNSLYTDKDIFL 148
> tgo:TGME49_110430 heat shock protein 90, putative (EC:3.2.1.3
2.7.13.3)
Length=1100
Score = 45.1 bits (105), Expect = 7e-05, Method: Compositional matrix adjust.
Identities = 18/28 (64%), Positives = 23/28 (82%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E +QAEV R++DIIVNSLY+ R+VFL
Sbjct 246 EIFPFQAEVKRVLDIIVNSLYTDRDVFL 273
> sce:YMR186W HSC82, HSP90; Cytoplasmic chaperone of the Hsp90
family, redundant in function and nearly identical with Hsp82p,
and together they are essential; expressed constitutively
at 10-fold higher basal levels than HSP82 and induced 2-3
fold by heat shock; K04079 molecular chaperone HtpG
Length=705
Score = 43.5 bits (101), Expect = 2e-04, Method: Composition-based stats.
Identities = 16/28 (57%), Positives = 25/28 (89%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET E+QAE+T+LM +I+N++YS +E+FL
Sbjct 4 ETFEFQAEITQLMSLIINTVYSNKEIFL 31
> dre:386590 hsp90b1, GP96, GRP94, fb61d09, tra-1, tra1, wu:fb61d09,
wu:fq25g01; heat shock protein 90, beta (grp94), member
1; K09487 heat shock protein 90kDa beta
Length=793
Score = 43.5 bits (101), Expect = 2e-04, Method: Composition-based stats.
Identities = 19/42 (45%), Positives = 29/42 (69%), Gaps = 4/42 (9%)
Query 94 AELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
++L ++R AE H +QAEV R+M +I+NSLY +E+FL
Sbjct 64 SQLKEIRDKAE----KHAFQAEVNRMMKLIINSLYKNKEIFL 101
> sce:YPL240C HSP82, HSP90; Hsp82p; K04079 molecular chaperone
HtpG
Length=709
Score = 43.5 bits (101), Expect = 2e-04, Method: Composition-based stats.
Identities = 16/28 (57%), Positives = 25/28 (89%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET E+QAE+T+LM +I+N++YS +E+FL
Sbjct 4 ETFEFQAEITQLMSLIINTVYSNKEIFL 31
> hsa:10131 TRAP1, HSP75, HSP90L; TNF receptor-associated protein
1; K09488 TNF receptor-associated protein 1
Length=704
Score = 42.4 bits (98), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 18/44 (40%), Positives = 27/44 (61%), Gaps = 0/44 (0%)
Query 92 PLAELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
PL + +G HE+QAE +L+DI+ SLYS++EVF+
Sbjct 70 PLHSIISSTESVQGSTSKHEFQAETKKLLDIVARSLYSEKEVFI 113
> dre:571959 trap1, fc85a11, wu:fc85a11; TNF receptor-associated
protein 1; K09488 TNF receptor-associated protein 1
Length=719
Score = 42.0 bits (97), Expect = 6e-04, Method: Compositional matrix adjust.
Identities = 16/34 (47%), Positives = 25/34 (73%), Gaps = 0/34 (0%)
Query 102 DAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
+ +G HE+QAE +L+DI+ SLYS++EVF+
Sbjct 95 NVQGSFSKHEFQAETKKLLDIVARSLYSEKEVFI 128
> mmu:68015 Trap1, 2410002K23Rik, HSP75; TNF receptor-associated
protein 1; K09488 TNF receptor-associated protein 1
Length=706
Score = 41.2 bits (95), Expect = 9e-04, Method: Compositional matrix adjust.
Identities = 15/26 (57%), Positives = 22/26 (84%), Gaps = 0/26 (0%)
Query 110 HEYQAEVTRLMDIIVNSLYSQREVFL 135
HE+QAE +L+DI+ SLYS++EVF+
Sbjct 90 HEFQAETKKLLDIVARSLYSEKEVFI 115
> pfa:PF14_0417 HSP90
Length=927
Score = 41.2 bits (95), Expect = 0.001, Method: Composition-based stats.
Identities = 16/28 (57%), Positives = 24/28 (85%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E + ++AEV ++MDIIVNSLY+ ++VFL
Sbjct 100 EKYNFKAEVNKVMDIIVNSLYTDKDVFL 127
> xla:446459 hsp90ab1, hsp90b, hsp90beta; heat shock protein 90kDa
alpha (cytosolic), class B member 1; K04079 molecular chaperone
HtpG
Length=722
Score = 40.0 bits (92), Expect = 0.002, Method: Composition-based stats.
Identities = 15/34 (44%), Positives = 24/34 (70%), Gaps = 0/34 (0%)
Query 102 DAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
+ E ET +QAE+ +LM +I+N+ YS +E+FL
Sbjct 7 NGEEEVETFAFQAEIAQLMSLIINTFYSNKEIFL 40
> tgo:TGME49_092920 heat shock protein 90, putative ; K09488 TNF
receptor-associated protein 1
Length=861
Score = 40.0 bits (92), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 16/35 (45%), Positives = 26/35 (74%), Gaps = 2/35 (5%)
Query 101 ADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
AD+EG E H ++AE +L+ I+ +SLY+ +EVF+
Sbjct 154 ADSEG--EVHTFKAETKKLLHIVTHSLYTDKEVFV 186
> dre:30591 hsp90a.1, fb17b01, hsp90, hsp90a, hsp90alpha, wu:fb17b01,
zgc:86652; heat shock protein 90-alpha 1; K04079 molecular
chaperone HtpG
Length=726
Score = 39.7 bits (91), Expect = 0.002, Method: Composition-based stats.
Identities = 18/44 (40%), Positives = 28/44 (63%), Gaps = 2/44 (4%)
Query 92 PLAELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
P A ++ D E ET +QAE+ +LM +I+N+ YS +E+FL
Sbjct 2 PEAHEQQMMEDEE--VETFAFQAEIAQLMSLIINTFYSNKEIFL 43
> dre:565155 hsp90a.2, cb820, hsp90a2; heat shock protein 90-alpha
2; K04079 molecular chaperone HtpG
Length=734
Score = 39.7 bits (91), Expect = 0.002, Method: Composition-based stats.
Identities = 18/44 (40%), Positives = 28/44 (63%), Gaps = 2/44 (4%)
Query 92 PLAELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
P A ++ D E ET +QAE+ +LM +I+N+ YS +E+FL
Sbjct 2 PEAHEQQMMEDEE--VETFAFQAEIAQLMSLIINTFYSNKEIFL 43
> bbo:BBOV_III004230 17.m07381; hsp90 protein; K04079 molecular
chaperone HtpG
Length=712
Score = 39.7 bits (91), Expect = 0.003, Method: Composition-based stats.
Identities = 13/33 (39%), Positives = 25/33 (75%), Gaps = 0/33 (0%)
Query 103 AEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
A QET+ + A++++L+ +I+N+ YS +E+FL
Sbjct 2 ATAQQETYAFNADISQLLSLIINAFYSNKEIFL 34
> mmu:15519 Hsp90aa1, 86kDa, 89kDa, AL024080, AL024147, Hsp86-1,
Hsp89, Hsp90, Hspca, hsp4; heat shock protein 90, alpha (cytosolic),
class A member 1; K04079 molecular chaperone HtpG
Length=733
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 15/32 (46%), Positives = 23/32 (71%), Gaps = 0/32 (0%)
Query 104 EGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E ET +QAE+ +LM +I+N+ YS +E+FL
Sbjct 14 EEEVETFAFQAEIAQLMSLIINTFYSNKEIFL 45
> hsa:3320 HSP90AA1, FLJ31884, HSP86, HSP89A, HSP90A, HSP90N,
HSPC1, HSPCA, HSPCAL1, HSPCAL4, HSPN, Hsp89, Hsp90, LAP2; heat
shock protein 90kDa alpha (cytosolic), class A member 1;
K04079 molecular chaperone HtpG
Length=854
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 15/32 (46%), Positives = 23/32 (71%), Gaps = 0/32 (0%)
Query 104 EGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E ET +QAE+ +LM +I+N+ YS +E+FL
Sbjct 136 EEEVETFAFQAEIAQLMSLIINTFYSNKEIFL 167
> cel:R151.7 hypothetical protein
Length=479
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 16/28 (57%), Positives = 20/28 (71%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
+ HE+QAE LMDI+ SLYS EVF+
Sbjct 43 QRHEFQAETRNLMDIVAKSLYSHSEVFV 70
> xla:399408 hsp90b1, ecgp, gp96, grp94, tra1; heat shock protein
90kDa beta (Grp94), member 1
Length=804
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 17/42 (40%), Positives = 28/42 (66%), Gaps = 4/42 (9%)
Query 94 AELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
A++ ++R +E +QAEV R+M +I+NSLY +E+FL
Sbjct 64 AQIKEIREKSE----KFAFQAEVNRMMKLIINSLYKNKEIFL 101
> xla:398753 hypothetical protein MGC68448; K09487 heat shock
protein 90kDa beta
Length=805
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 17/42 (40%), Positives = 28/42 (66%), Gaps = 4/42 (9%)
Query 94 AELAKLRADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
A++ ++R +E +QAEV R+M +I+NSLY +E+FL
Sbjct 64 AQIKEIREKSE----KFAFQAEVNRMMKLIINSLYKNKEIFL 101
> mmu:22027 Hsp90b1, ERp99, GRP94, TA-3, Targ2, Tra-1, Tra1, endoplasmin,
gp96; heat shock protein 90, beta (Grp94), member
1; K09487 heat shock protein 90kDa beta
Length=802
Score = 39.3 bits (90), Expect = 0.004, Method: Composition-based stats.
Identities = 15/28 (53%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E +QAEV R+M +I+NSLY +E+FL
Sbjct 74 EKFAFQAEVNRMMKLIINSLYKNKEIFL 101
> hsa:7184 HSP90B1, ECGP, GP96, GRP94, TRA1; heat shock protein
90kDa beta (Grp94), member 1; K09487 heat shock protein 90kDa
beta
Length=803
Score = 39.3 bits (90), Expect = 0.004, Method: Composition-based stats.
Identities = 15/28 (53%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
E +QAEV R+M +I+NSLY +E+FL
Sbjct 74 EKFAFQAEVNRMMKLIINSLYKNKEIFL 101
> xla:444024 hsp90aa1.1, MGC82579, hsp86, hsp89, hsp90, hsp90a,
hspc1, hspca, hspn, lap2; heat shock protein 90kDa alpha (cytosolic),
class A member 1, gene 1; K04079 molecular chaperone
HtpG
Length=729
Score = 39.3 bits (90), Expect = 0.004, Method: Composition-based stats.
Identities = 14/28 (50%), Positives = 22/28 (78%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET +QAE+ +LM +I+N+ YS +E+FL
Sbjct 19 ETFAFQAEIAQLMSLIINTFYSNKEIFL 46
> cel:C47E8.5 daf-21; abnormal DAuer Formation family member (daf-21);
K04079 molecular chaperone HtpG
Length=702
Score = 37.7 bits (86), Expect = 0.009, Method: Composition-based stats.
Identities = 13/28 (46%), Positives = 22/28 (78%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET +QAE+ +LM +I+N+ YS +E++L
Sbjct 6 ETFAFQAEIAQLMSLIINTFYSNKEIYL 33
> ath:AT5G52640 ATHSP90.1 (HEAT SHOCK PROTEIN 90.1); ATP binding
/ unfolded protein binding; K04079 molecular chaperone HtpG
Length=705
Score = 37.7 bits (86), Expect = 0.010, Method: Composition-based stats.
Identities = 13/28 (46%), Positives = 22/28 (78%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET +QAE+ +L+ +I+N+ YS +E+FL
Sbjct 10 ETFAFQAEINQLLSLIINTFYSNKEIFL 37
> ath:AT5G56030 HSP81-2 (HEAT SHOCK PROTEIN 81-2); ATP binding;
K04079 molecular chaperone HtpG
Length=699
Score = 37.7 bits (86), Expect = 0.010, Method: Composition-based stats.
Identities = 13/28 (46%), Positives = 22/28 (78%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET +QAE+ +L+ +I+N+ YS +E+FL
Sbjct 5 ETFAFQAEINQLLSLIINTFYSNKEIFL 32
> ath:AT5G56010 HSP81-3; ATP binding / unfolded protein binding;
K04079 molecular chaperone HtpG
Length=699
Score = 37.7 bits (86), Expect = 0.010, Method: Composition-based stats.
Identities = 13/28 (46%), Positives = 22/28 (78%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET +QAE+ +L+ +I+N+ YS +E+FL
Sbjct 5 ETFAFQAEINQLLSLIINTFYSNKEIFL 32
> ath:AT5G56000 heat shock protein 81-4 (HSP81-4); K04079 molecular
chaperone HtpG
Length=699
Score = 37.7 bits (86), Expect = 0.010, Method: Composition-based stats.
Identities = 13/28 (46%), Positives = 22/28 (78%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET +QAE+ +L+ +I+N+ YS +E+FL
Sbjct 5 ETFAFQAEINQLLSLIINTFYSNKEIFL 32
> dre:30573 hsp90ab1, hsp90b, hsp90beta, wu:fa29f01, wu:fa91e11,
wu:fd59e11, wu:gcd22h07; heat shock protein 90, alpha (cytosolic),
class B member 1; K04079 molecular chaperone HtpG
Length=725
Score = 37.0 bits (84), Expect = 0.015, Method: Composition-based stats.
Identities = 13/27 (48%), Positives = 21/27 (77%), Gaps = 0/27 (0%)
Query 109 THEYQAEVTRLMDIIVNSLYSQREVFL 135
T +QAE+ +LM +I+N+ YS +E+FL
Sbjct 13 TFAFQAEIAQLMSLIINTFYSNKEIFL 39
> mmu:15516 Hsp90ab1, 90kDa, AL022974, C81438, Hsp84, Hsp84-1,
Hsp90, Hspcb, MGC115780; heat shock protein 90 alpha (cytosolic),
class B member 1; K04079 molecular chaperone HtpG
Length=724
Score = 37.0 bits (84), Expect = 0.015, Method: Composition-based stats.
Identities = 13/27 (48%), Positives = 21/27 (77%), Gaps = 0/27 (0%)
Query 109 THEYQAEVTRLMDIIVNSLYSQREVFL 135
T +QAE+ +LM +I+N+ YS +E+FL
Sbjct 14 TFAFQAEIAQLMSLIINTFYSNKEIFL 40
> hsa:3326 HSP90AB1, D6S182, FLJ26984, HSP90-BETA, HSP90B, HSPC2,
HSPCB; heat shock protein 90kDa alpha (cytosolic), class
B member 1; K04079 molecular chaperone HtpG
Length=724
Score = 37.0 bits (84), Expect = 0.015, Method: Composition-based stats.
Identities = 13/27 (48%), Positives = 21/27 (77%), Gaps = 0/27 (0%)
Query 109 THEYQAEVTRLMDIIVNSLYSQREVFL 135
T +QAE+ +LM +I+N+ YS +E+FL
Sbjct 14 TFAFQAEIAQLMSLIINTFYSNKEIFL 40
> eco:b0473 htpG, ECK0467, JW0462; molecular chaperone HSP90 family;
K04079 molecular chaperone HtpG
Length=624
Score = 37.0 bits (84), Expect = 0.018, Method: Composition-based stats.
Identities = 14/29 (48%), Positives = 24/29 (82%), Gaps = 0/29 (0%)
Query 107 QETHEYQAEVTRLMDIIVNSLYSQREVFL 135
QET +Q+EV +L+ ++++SLYS +E+FL
Sbjct 4 QETRGFQSEVKQLLHLMIHSLYSNKEIFL 32
> tpv:TP02_0244 heat shock protein 90; K04079 molecular chaperone
HtpG
Length=721
Score = 36.6 bits (83), Expect = 0.024, Method: Composition-based stats.
Identities = 12/34 (35%), Positives = 24/34 (70%), Gaps = 0/34 (0%)
Query 102 DAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
D QE + + A++++L+ +I+N+ YS +E+FL
Sbjct 5 DETPDQEVYAFNADISQLLSLIINAFYSNKEIFL 38
> bbo:BBOV_IV010880 23.m05763; heat shock protein 75; K09488 TNF
receptor-associated protein 1
Length=623
Score = 36.2 bits (82), Expect = 0.029, Method: Composition-based stats.
Identities = 12/28 (42%), Positives = 24/28 (85%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
+T+E++AE+ +L+ I+ +SLY+ +EVF+
Sbjct 16 DTYEFKAEIQKLLQIVAHSLYTDKEVFV 43
> pfa:PF11_0188 heat shock protein 90, putative
Length=930
Score = 35.4 bits (80), Expect = 0.055, Method: Composition-based stats.
Identities = 14/44 (31%), Positives = 30/44 (68%), Gaps = 1/44 (2%)
Query 93 LAELAKL-RADAEGHQETHEYQAEVTRLMDIIVNSLYSQREVFL 135
+ E++K+ + + E +E++AE +L+ I+ +SLY+ +EVF+
Sbjct 55 ICEISKMNKRNYSSECENYEFKAETKKLLQIVAHSLYTDKEVFI 98
> cpv:cgd3_3770 Hsp90 ; K04079 molecular chaperone HtpG
Length=711
Score = 35.4 bits (80), Expect = 0.055, Method: Composition-based stats.
Identities = 12/28 (42%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET + A++ +LM +I+N+ YS +E+FL
Sbjct 15 ETFAFNADIQQLMSLIINTFYSNKEIFL 42
> tgo:TGME49_088380 heat shock protein 90 ; K04079 molecular chaperone
HtpG
Length=708
Score = 35.4 bits (80), Expect = 0.055, Method: Composition-based stats.
Identities = 12/28 (42%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET + A++ +LM +I+N+ YS +E+FL
Sbjct 5 ETFAFNADIQQLMSLIINTFYSNKEIFL 32
> pfa:PF07_0029 heat shock protein 86; K04079 molecular chaperone
HtpG
Length=745
Score = 35.0 bits (79), Expect = 0.062, Method: Composition-based stats.
Identities = 12/28 (42%), Positives = 21/28 (75%), Gaps = 0/28 (0%)
Query 108 ETHEYQAEVTRLMDIIVNSLYSQREVFL 135
ET + A++ +LM +I+N+ YS +E+FL
Sbjct 4 ETFAFNADIRQLMSLIINTFYSNKEIFL 31
> cel:K03H1.10 hypothetical protein
Length=322
Score = 35.0 bits (79), Expect = 0.070, Method: Compositional matrix adjust.
Identities = 20/46 (43%), Positives = 23/46 (50%), Gaps = 7/46 (15%)
Query 86 QGAPQGPLAELAKLRADAEGHQETHEYQAEVTR-------LMDIIV 124
QGA Q PL E A++R D GHQ + E R LMDI V
Sbjct 250 QGAAQSPLDEFARMRIDEGGHQLRTNQETETNRQNQSQQPLMDINV 295
Lambda K H
0.313 0.122 0.339
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2296762580
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40