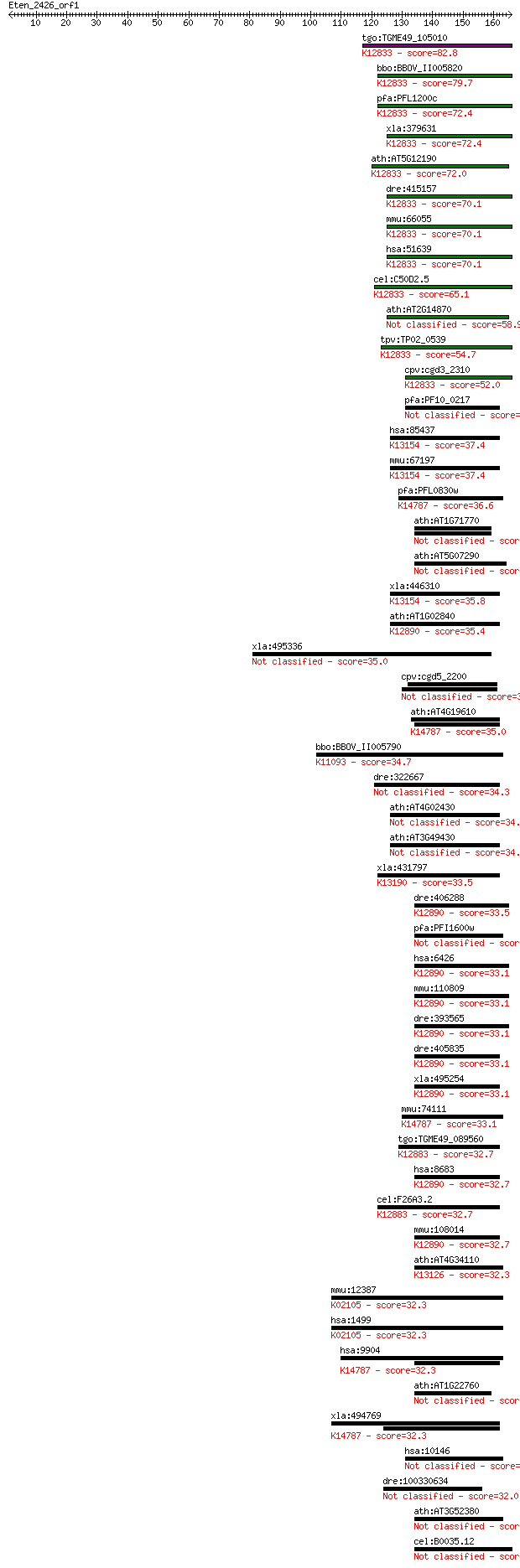
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_2426_orf1
Length=165
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_105010 RNA binding protein, putative ; K12833 pre-m... 82.8 5e-16
bbo:BBOV_II005820 18.m06483; pre-mRNA branch site protein p14;... 79.7 4e-15
pfa:PFL1200c splicing factor 3b subunit, putative; K12833 pre-... 72.4 6e-13
xla:379631 sf3b14, MGC68842; splicing factor 3B, 14 kDa subuni... 72.4 7e-13
ath:AT5G12190 RNA recognition motif (RRM)-containing protein; ... 72.0 8e-13
dre:415157 sf3b14, zgc:86708; splicing factor 3B; K12833 pre-m... 70.1 3e-12
mmu:66055 0610009D07Rik, 6030419K15Rik, AV001342, Sf3b14; RIKE... 70.1 3e-12
hsa:51639 SF3B14, HSPC175, Ht006, P14, SAP14, SF3B14a; splicin... 70.1 3e-12
cel:C50D2.5 hypothetical protein; K12833 pre-mRNA branch site ... 65.1 1e-10
ath:AT2G14870 RNA recognition motif (RRM)-containing protein 58.9 8e-09
tpv:TP02_0539 hypothetical protein; K12833 pre-mRNA branch sit... 54.7 1e-07
cpv:cgd3_2310 hypothetical protein ; K12833 pre-mRNA branch si... 52.0 9e-07
pfa:PF10_0217 pre-mRNA splicing factor, putative 39.3 0.006
hsa:85437 ZCRB1, MADP-1, MADP1, MGC26805, RBM36, ZCCHC19; zinc... 37.4 0.022
mmu:67197 Zcrb1, 2700088M22Rik, Madp-1; zinc finger CCHC-type ... 37.4 0.023
pfa:PFL0830w RNA binding protein, putative; K14787 multiple RN... 36.6 0.035
ath:AT1G71770 PAB5; PAB5 (POLY(A)-BINDING PROTEIN 5); RNA bind... 36.2 0.057
ath:AT5G07290 AML4; AML4 (ARABIDOPSIS MEI2-LIKE 4); RNA bindin... 35.8 0.066
xla:446310 zcrb1, MGC82154; zinc finger CCHC-type and RNA bind... 35.8 0.075
ath:AT1G02840 SR1; SR1; RNA binding / nucleic acid binding / n... 35.4 0.083
xla:495336 hypothetical LOC495336 35.0 0.11
cpv:cgd5_2200 T8M16_190-plant like protein with RRM domain 35.0 0.12
ath:AT4G19610 RNA binding / nucleic acid binding / nucleotide ... 35.0 0.12
bbo:BBOV_II005790 18.m06481; u1 snRNP; K11093 U1 small nuclear... 34.7 0.17
dre:322667 fb69f10; wu:fb69f10 34.3 0.17
ath:AT4G02430 pre-mRNA splicing factor, putative / SR1 protein... 34.3 0.18
ath:AT3G49430 SRp34a; SRp34a (Ser/Arg-rich protein 34a); RNA b... 34.3 0.22
xla:431797 rbm15, MGC83913; RNA binding motif protein 15; K131... 33.5 0.29
dre:406288 srsf1a, sfrs1a, sfrs1l, wu:fb52g10, wu:fb97g12, zgc... 33.5 0.33
pfa:PFI1600w mRNA processing protein, putative 33.1 0.38
hsa:6426 SRSF1, ASF, FLJ53078, MGC5228, SF2, SF2p33, SFRS1, SR... 33.1 0.39
mmu:110809 Srsf1, 1110054N12Rik, 5730507C05Rik, 6330415C05Rik,... 33.1 0.41
dre:393565 srsf1b, MGC65898, sfrs1, sfrs1b, wu:fb80g05, zgc:11... 33.1 0.42
dre:405835 MGC77449; zgc:77449; K12890 splicing factor, argini... 33.1 0.46
xla:495254 srsf9, sfrs9, srp30c; serine/arginine-rich splicing... 33.1 0.47
mmu:74111 Rbm19, 1200009A02Rik, AV302714, KIAA0682, NPO; RNA b... 33.1 0.48
tgo:TGME49_089560 nuclear cap-binding protein, putative ; K128... 32.7 0.49
hsa:8683 SRSF9, SFRS9, SRp30c; serine/arginine-rich splicing f... 32.7 0.50
cel:F26A3.2 ncbp-2; Nuclear Cap-Binding Protein family member ... 32.7 0.61
mmu:108014 Srsf9, 25kDa, 2610029M16Rik, SRp30c, Sfrs9; serine/... 32.7 0.61
ath:AT4G34110 PAB2; PAB2 (POLY(A) BINDING 2); RNA binding / tr... 32.3 0.66
mmu:12387 Ctnnb1, Bfc, Catnb, Mesc; catenin (cadherin associat... 32.3 0.69
hsa:1499 CTNNB1, CTNNB, DKFZp686D02253, FLJ25606, FLJ37923; ca... 32.3 0.70
hsa:9904 RBM19, DKFZp586F1023, KIAA0682; RNA binding motif pro... 32.3 0.72
ath:AT1G22760 PAB3; PAB3 (POLY(A) BINDING PROTEIN 3); RNA bind... 32.3 0.73
xla:494769 rbm19; RNA binding motif protein 19; K14787 multipl... 32.3 0.76
hsa:10146 G3BP1, G3BP, HDH-VIII, MGC111040; GTPase activating ... 32.0 0.98
dre:100330634 RNA binding motif protein 12-like 32.0 1.0
ath:AT3G52380 CP33; CP33; RNA binding 32.0 1.1
cel:B0035.12 hypothetical protein 31.6 1.1
> tgo:TGME49_105010 RNA binding protein, putative ; K12833 pre-mRNA
branch site protein p14
Length=156
Score = 82.8 bits (203), Expect = 5e-16, Method: Compositional matrix adjust.
Identities = 32/49 (65%), Positives = 44/49 (89%), Gaps = 0/49 (0%)
Query 117 LAGRGRPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
++ RGRP ++AP++SRI+YVRNLPFKI +ELYD+FGKYG++RQIR+G
Sbjct 37 VSTRGRPTKIAPDMSRIIYVRNLPFKITDDELYDIFGKYGAVRQIRKGN 85
> bbo:BBOV_II005820 18.m06483; pre-mRNA branch site protein p14;
K12833 pre-mRNA branch site protein p14
Length=122
Score = 79.7 bits (195), Expect = 4e-15, Method: Compositional matrix adjust.
Identities = 33/44 (75%), Positives = 40/44 (90%), Gaps = 0/44 (0%)
Query 122 RPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
R RL+PEVSRILY+RNLP+KI+ EELYD+FGKYGS+RQIR+G
Sbjct 10 RTMRLSPEVSRILYLRNLPYKISAEELYDIFGKYGSVRQIRKGN 53
> pfa:PFL1200c splicing factor 3b subunit, putative; K12833 pre-mRNA
branch site protein p14
Length=106
Score = 72.4 bits (176), Expect = 6e-13, Method: Compositional matrix adjust.
Identities = 31/44 (70%), Positives = 38/44 (86%), Gaps = 0/44 (0%)
Query 122 RPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
R RL EVSRILYVRNLP+KI+ +ELYD+FGKYG++RQIR+G
Sbjct 4 RNIRLPAEVSRILYVRNLPYKISADELYDIFGKYGTVRQIRKGN 47
> xla:379631 sf3b14, MGC68842; splicing factor 3B, 14 kDa subunit;
K12833 pre-mRNA branch site protein p14
Length=125
Score = 72.4 bits (176), Expect = 7e-13, Method: Compositional matrix adjust.
Identities = 31/41 (75%), Positives = 36/41 (87%), Gaps = 0/41 (0%)
Query 125 RLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
RL PEV+RILY+RNLP+KI GEE+YD+FGKYG IRQIR G
Sbjct 12 RLPPEVNRILYIRNLPYKITGEEMYDIFGKYGPIRQIRVGN 52
> ath:AT5G12190 RNA recognition motif (RRM)-containing protein;
K12833 pre-mRNA branch site protein p14
Length=124
Score = 72.0 bits (175), Expect = 8e-13, Method: Compositional matrix adjust.
Identities = 31/45 (68%), Positives = 36/45 (80%), Gaps = 0/45 (0%)
Query 120 RGRPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRG 164
R RL PEV+R+LYVRNLPF I EE+YD+FGKYG+IRQIR G
Sbjct 7 RKSNTRLPPEVNRVLYVRNLPFNITSEEMYDIFGKYGAIRQIRIG 51
> dre:415157 sf3b14, zgc:86708; splicing factor 3B; K12833 pre-mRNA
branch site protein p14
Length=125
Score = 70.1 bits (170), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 30/41 (73%), Positives = 35/41 (85%), Gaps = 0/41 (0%)
Query 125 RLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
RL PEV+RILY+RNLP+KI EE+YD+FGKYG IRQIR G
Sbjct 12 RLPPEVNRILYIRNLPYKITAEEMYDIFGKYGPIRQIRVGN 52
> mmu:66055 0610009D07Rik, 6030419K15Rik, AV001342, Sf3b14; RIKEN
cDNA 0610009D07 gene; K12833 pre-mRNA branch site protein
p14
Length=125
Score = 70.1 bits (170), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 30/41 (73%), Positives = 35/41 (85%), Gaps = 0/41 (0%)
Query 125 RLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
RL PEV+RILY+RNLP+KI EE+YD+FGKYG IRQIR G
Sbjct 12 RLPPEVNRILYIRNLPYKITAEEMYDIFGKYGPIRQIRVGN 52
> hsa:51639 SF3B14, HSPC175, Ht006, P14, SAP14, SF3B14a; splicing
factor 3B, 14 kDa subunit; K12833 pre-mRNA branch site protein
p14
Length=125
Score = 70.1 bits (170), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 30/41 (73%), Positives = 35/41 (85%), Gaps = 0/41 (0%)
Query 125 RLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
RL PEV+RILY+RNLP+KI EE+YD+FGKYG IRQIR G
Sbjct 12 RLPPEVNRILYIRNLPYKITAEEMYDIFGKYGPIRQIRVGN 52
> cel:C50D2.5 hypothetical protein; K12833 pre-mRNA branch site
protein p14
Length=138
Score = 65.1 bits (157), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 26/45 (57%), Positives = 37/45 (82%), Gaps = 0/45 (0%)
Query 121 GRPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
R +L PEV+RILY++NLP+KI EE+Y++FGK+G++RQIR G
Sbjct 8 NRGAKLPPEVNRILYIKNLPYKITTEEMYEIFGKFGAVRQIRVGN 52
> ath:AT2G14870 RNA recognition motif (RRM)-containing protein
Length=101
Score = 58.9 bits (141), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 24/40 (60%), Positives = 32/40 (80%), Gaps = 0/40 (0%)
Query 125 RLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIRRG 164
RL PEV+R+LY+ NLPF I E+ YD+FG+Y +IRQ+R G
Sbjct 13 RLPPEVTRLLYICNLPFSITSEDTYDLFGRYSTIRQVRIG 52
> tpv:TP02_0539 hypothetical protein; K12833 pre-mRNA branch site
protein p14
Length=134
Score = 54.7 bits (130), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 26/53 (49%), Positives = 32/53 (60%), Gaps = 10/53 (18%)
Query 123 PQRLAPEVSRILYVR----------NLPFKINGEELYDVFGKYGSIRQIRRGT 165
P EV R + +R NLP+KI EELYD+FGKYGS+RQIR+G
Sbjct 13 PTTFGLEVQRAIQLRHGGSSKTSNLNLPYKITSEELYDIFGKYGSVRQIRKGN 65
> cpv:cgd3_2310 hypothetical protein ; K12833 pre-mRNA branch
site protein p14
Length=86
Score = 52.0 bits (123), Expect = 9e-07, Method: Compositional matrix adjust.
Identities = 20/35 (57%), Positives = 29/35 (82%), Gaps = 0/35 (0%)
Query 131 SRILYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
S I+Y+R LP+ I+ +LYD+FG++G+IRQIRRG
Sbjct 15 SSIIYLRQLPYDISSTDLYDIFGRHGTIRQIRRGV 49
> pfa:PF10_0217 pre-mRNA splicing factor, putative
Length=538
Score = 39.3 bits (90), Expect = 0.006, Method: Composition-based stats.
Identities = 17/31 (54%), Positives = 22/31 (70%), Gaps = 0/31 (0%)
Query 131 SRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
S +YV NLP + EE+YD+FGKYG I+ I
Sbjct 11 SSCIYVGNLPGNVIEEEVYDLFGKYGRIKYI 41
> hsa:85437 ZCRB1, MADP-1, MADP1, MGC26805, RBM36, ZCCHC19; zinc
finger CCHC-type and RNA binding motif 1; K13154 U11/U12
small nuclear ribonucleoprotein 31 kDa protein
Length=217
Score = 37.4 bits (85), Expect = 0.022, Method: Compositional matrix adjust.
Identities = 16/36 (44%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query 126 LAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
LAP S + YV NLPF + +LY +F KYG + ++
Sbjct 5 LAPSKSTV-YVSNLPFSLTNNDLYRIFSKYGKVVKV 39
> mmu:67197 Zcrb1, 2700088M22Rik, Madp-1; zinc finger CCHC-type
and RNA binding motif 1; K13154 U11/U12 small nuclear ribonucleoprotein
31 kDa protein
Length=217
Score = 37.4 bits (85), Expect = 0.023, Method: Compositional matrix adjust.
Identities = 16/36 (44%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query 126 LAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
LAP S + YV NLPF + +LY +F KYG + ++
Sbjct 5 LAPSKSTV-YVSNLPFSLTNNDLYRIFSKYGKVVKV 39
> pfa:PFL0830w RNA binding protein, putative; K14787 multiple
RNA-binding domain-containing protein 1
Length=1119
Score = 36.6 bits (83), Expect = 0.035, Method: Composition-based stats.
Identities = 13/34 (38%), Positives = 25/34 (73%), Gaps = 0/34 (0%)
Query 129 EVSRILYVRNLPFKINGEELYDVFGKYGSIRQIR 162
+V++ L V+NL F++N EEL +F +G+++ +R
Sbjct 980 QVTKKLVVKNLAFQVNKEELRKLFSAFGNVKSVR 1013
> ath:AT1G71770 PAB5; PAB5 (POLY(A)-BINDING PROTEIN 5); RNA binding
/ poly(A) binding / translation initiation factor
Length=668
Score = 36.2 bits (82), Expect = 0.057, Method: Compositional matrix adjust.
Identities = 14/25 (56%), Positives = 18/25 (72%), Gaps = 0/25 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSI 158
+YV+NLP +I +EL FGKYG I
Sbjct 227 VYVKNLPKEITDDELKKTFGKYGDI 251
Score = 30.4 bits (67), Expect = 2.8, Method: Compositional matrix adjust.
Identities = 10/25 (40%), Positives = 19/25 (76%), Gaps = 0/25 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSI 158
LY++NL +N E+L ++F +YG++
Sbjct 330 LYLKNLDDSVNDEKLKEMFSEYGNV 354
> ath:AT5G07290 AML4; AML4 (ARABIDOPSIS MEI2-LIKE 4); RNA binding
/ nucleic acid binding / nucleotide binding
Length=907
Score = 35.8 bits (81), Expect = 0.066, Method: Composition-based stats.
Identities = 15/30 (50%), Positives = 21/30 (70%), Gaps = 0/30 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQIRR 163
L+V NL I+ EEL+ +F YG IR++RR
Sbjct 297 LWVNNLDSSISNEELHGIFSSYGEIREVRR 326
> xla:446310 zcrb1, MGC82154; zinc finger CCHC-type and RNA binding
motif 1; K13154 U11/U12 small nuclear ribonucleoprotein
31 kDa protein
Length=218
Score = 35.8 bits (81), Expect = 0.075, Method: Compositional matrix adjust.
Identities = 15/36 (41%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query 126 LAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
LAP S + YV NLPF + +L+ +F KYG + ++
Sbjct 5 LAPSKSTV-YVSNLPFSLTNNDLHRIFSKYGKVVKV 39
> ath:AT1G02840 SR1; SR1; RNA binding / nucleic acid binding /
nucleotide binding; K12890 splicing factor, arginine/serine-rich
1/9
Length=303
Score = 35.4 bits (80), Expect = 0.083, Method: Compositional matrix adjust.
Identities = 16/36 (44%), Positives = 22/36 (61%), Gaps = 0/36 (0%)
Query 126 LAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
++ SR +YV NLP I E+ D+F KYG + QI
Sbjct 1 MSSRSSRTVYVGNLPGDIREREVEDLFSKYGPVVQI 36
> xla:495336 hypothetical LOC495336
Length=711
Score = 35.0 bits (79), Expect = 0.11, Method: Compositional matrix adjust.
Identities = 22/79 (27%), Positives = 40/79 (50%), Gaps = 1/79 (1%)
Query 81 FAGCSLSGVPRECCGAAAAAAAAAAAKSPAMQPAPLLAGRGRPQRLAPEVSRI-LYVRNL 139
A +++ + + G + A+ A K + L + + +R + VS + LYV+NL
Sbjct 233 HAEAAVAAMHGKEIGGRSLYASRAQRKEERQEELKLRLEKQKAERRSKYVSNVNLYVKNL 292
Query 140 PFKINGEELYDVFGKYGSI 158
+I+ E L ++F KYG I
Sbjct 293 DDEIDDERLKEIFSKYGPI 311
> cpv:cgd5_2200 T8M16_190-plant like protein with RRM domain
Length=600
Score = 35.0 bits (79), Expect = 0.12, Method: Composition-based stats.
Identities = 12/29 (41%), Positives = 20/29 (68%), Gaps = 0/29 (0%)
Query 132 RILYVRNLPFKINGEELYDVFGKYGSIRQ 160
R LY++NLPF N E + +VF ++G + +
Sbjct 97 RRLYIKNLPFSANTESVIEVFSQFGDVEE 125
Score = 33.5 bits (75), Expect = 0.35, Method: Composition-based stats.
Identities = 13/31 (41%), Positives = 21/31 (67%), Gaps = 0/31 (0%)
Query 130 VSRILYVRNLPFKINGEELYDVFGKYGSIRQ 160
+ R L+VRNL F+ N + L +V G+YG + +
Sbjct 183 IRRKLFVRNLGFETNEDSLSEVMGQYGELEE 213
> ath:AT4G19610 RNA binding / nucleic acid binding / nucleotide
binding; K14787 multiple RNA-binding domain-containing protein
1
Length=816
Score = 35.0 bits (79), Expect = 0.12, Method: Composition-based stats.
Identities = 15/29 (51%), Positives = 21/29 (72%), Gaps = 0/29 (0%)
Query 133 ILYVRNLPFKINGEELYDVFGKYGSIRQI 161
IL V+NLPF +EL +FGK+GS+ +I
Sbjct 489 ILLVKNLPFASTEKELAQMFGKFGSLDKI 517
Score = 31.6 bits (70), Expect = 1.4, Method: Composition-based stats.
Identities = 12/28 (42%), Positives = 18/28 (64%), Gaps = 0/28 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI 161
L+VRNLP+ EEL + F +G I ++
Sbjct 296 LFVRNLPYTATEEELMEHFSTFGKISEV 323
> bbo:BBOV_II005790 18.m06481; u1 snRNP; K11093 U1 small nuclear
ribonucleoprotein 70kDa
Length=247
Score = 34.7 bits (78), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 20/61 (32%), Positives = 32/61 (52%), Gaps = 2/61 (3%)
Query 102 AAAAAKSPAMQPAPLLAGRGRPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
A A P PA L G P L + + L+V N+P+++ ++L+ F YG +R+I
Sbjct 79 AERDAYDPKYDPA-LARGATNPW-LTHDPYKTLFVSNIPYEVTEKQLWKEFDVYGRVRRI 136
Query 162 R 162
R
Sbjct 137 R 137
> dre:322667 fb69f10; wu:fb69f10
Length=3136
Score = 34.3 bits (77), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 15/41 (36%), Positives = 23/41 (56%), Gaps = 0/41 (0%)
Query 121 GRPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
GR P+ +R L++ NL N ++L DVF ++G I I
Sbjct 430 GRIDEFHPKATRTLFIGNLEKTTNYQQLLDVFQRFGEIVDI 470
> ath:AT4G02430 pre-mRNA splicing factor, putative / SR1 protein,
putative
Length=278
Score = 34.3 bits (77), Expect = 0.18, Method: Compositional matrix adjust.
Identities = 16/36 (44%), Positives = 22/36 (61%), Gaps = 0/36 (0%)
Query 126 LAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
++ SR +YV NLP I E+ D+F KYG + QI
Sbjct 1 MSSRSSRTIYVGNLPGDIREREVEDLFSKYGPVVQI 36
> ath:AT3G49430 SRp34a; SRp34a (Ser/Arg-rich protein 34a); RNA
binding / nucleic acid binding / nucleotide binding
Length=297
Score = 34.3 bits (77), Expect = 0.22, Method: Compositional matrix adjust.
Identities = 16/36 (44%), Positives = 21/36 (58%), Gaps = 0/36 (0%)
Query 126 LAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
++ SR +YV NLP I E+ D+F KYG I I
Sbjct 1 MSGRFSRSIYVGNLPGDIREHEIEDIFYKYGRIVDI 36
> xla:431797 rbm15, MGC83913; RNA binding motif protein 15; K13190
RNA-binding protein 15
Length=830
Score = 33.5 bits (75), Expect = 0.29, Method: Compositional matrix adjust.
Identities = 16/44 (36%), Positives = 27/44 (61%), Gaps = 4/44 (9%)
Query 122 RPQRLAPEV----SRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
R + LA E+ +R L+V NL + E+Y VFG++G+I ++
Sbjct 285 RDEDLALEIDARANRTLFVGNLDVIVKETEIYRVFGRFGTITEV 328
> dre:406288 srsf1a, sfrs1a, sfrs1l, wu:fb52g10, wu:fb97g12, zgc:66146;
serine/arginine-rich splicing factor 1a; K12890 splicing
factor, arginine/serine-rich 1/9
Length=245
Score = 33.5 bits (75), Expect = 0.33, Method: Compositional matrix adjust.
Identities = 18/35 (51%), Positives = 23/35 (65%), Gaps = 4/35 (11%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI----RRG 164
+YV NLP I +++ DVF KYG+IR I RRG
Sbjct 17 IYVGNLPPDIRTKDVEDVFYKYGAIRDIDLKNRRG 51
> pfa:PFI1600w mRNA processing protein, putative
Length=761
Score = 33.1 bits (74), Expect = 0.38, Method: Composition-based stats.
Identities = 12/29 (41%), Positives = 19/29 (65%), Gaps = 0/29 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQIR 162
L++ N+PF I ELY++ K G +R +R
Sbjct 10 LWIGNIPFDITENELYEILCKVGVVRNVR 38
> hsa:6426 SRSF1, ASF, FLJ53078, MGC5228, SF2, SF2p33, SFRS1,
SRp30a; serine/arginine-rich splicing factor 1; K12890 splicing
factor, arginine/serine-rich 1/9
Length=201
Score = 33.1 bits (74), Expect = 0.39, Method: Compositional matrix adjust.
Identities = 18/35 (51%), Positives = 23/35 (65%), Gaps = 4/35 (11%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI----RRG 164
+YV NLP I +++ DVF KYG+IR I RRG
Sbjct 18 IYVGNLPPDIRTKDIEDVFYKYGAIRDIDLKNRRG 52
> mmu:110809 Srsf1, 1110054N12Rik, 5730507C05Rik, 6330415C05Rik,
AI482334, AW491331, Asf, Sf2, Sfrs1; serine/arginine-rich
splicing factor 1; K12890 splicing factor, arginine/serine-rich
1/9
Length=201
Score = 33.1 bits (74), Expect = 0.41, Method: Compositional matrix adjust.
Identities = 18/35 (51%), Positives = 23/35 (65%), Gaps = 4/35 (11%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI----RRG 164
+YV NLP I +++ DVF KYG+IR I RRG
Sbjct 18 IYVGNLPPDIRTKDIEDVFYKYGAIRDIDLKNRRG 52
> dre:393565 srsf1b, MGC65898, sfrs1, sfrs1b, wu:fb80g05, zgc:111894,
zgc:65898, zgc:76897; serine/arginine-rich splicing
factor 1b; K12890 splicing factor, arginine/serine-rich 1/9
Length=245
Score = 33.1 bits (74), Expect = 0.42, Method: Compositional matrix adjust.
Identities = 18/35 (51%), Positives = 23/35 (65%), Gaps = 4/35 (11%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI----RRG 164
+YV NLP I +++ DVF KYG+IR I RRG
Sbjct 17 IYVGNLPPDIRTKDVEDVFYKYGAIRDIDLKNRRG 51
> dre:405835 MGC77449; zgc:77449; K12890 splicing factor, arginine/serine-rich
1/9
Length=244
Score = 33.1 bits (74), Expect = 0.46, Method: Compositional matrix adjust.
Identities = 13/28 (46%), Positives = 18/28 (64%), Gaps = 0/28 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI 161
+YV NLP + ++ D+F KYG IR I
Sbjct 6 IYVGNLPMDVQERDIEDLFFKYGKIRDI 33
> xla:495254 srsf9, sfrs9, srp30c; serine/arginine-rich splicing
factor 9; K12890 splicing factor, arginine/serine-rich 1/9
Length=230
Score = 33.1 bits (74), Expect = 0.47, Method: Compositional matrix adjust.
Identities = 14/28 (50%), Positives = 19/28 (67%), Gaps = 0/28 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI 161
+YV NLP I +EL D+F +YG IR +
Sbjct 18 IYVGNLPSDIREKELEDLFDRYGRIRTV 45
> mmu:74111 Rbm19, 1200009A02Rik, AV302714, KIAA0682, NPO; RNA
binding motif protein 19; K14787 multiple RNA-binding domain-containing
protein 1
Length=952
Score = 33.1 bits (74), Expect = 0.48, Method: Compositional matrix adjust.
Identities = 13/33 (39%), Positives = 23/33 (69%), Gaps = 1/33 (3%)
Query 130 VSRILYVRNLPFKINGEELYDVFGKYGSIRQIR 162
S+IL VRN+PF+ N E+ ++F +G ++ +R
Sbjct 823 TSKIL-VRNIPFQANQREIRELFSTFGELKTVR 854
> tgo:TGME49_089560 nuclear cap-binding protein, putative ; K12883
nuclear cap-binding protein subunit 2
Length=282
Score = 32.7 bits (73), Expect = 0.49, Method: Compositional matrix adjust.
Identities = 16/35 (45%), Positives = 24/35 (68%), Gaps = 2/35 (5%)
Query 129 EVSR--ILYVRNLPFKINGEELYDVFGKYGSIRQI 161
+VSR ++YV NL F +ELY+VF + G IR++
Sbjct 29 KVSRSCVVYVGNLNFSTTEDELYEVFSQAGLIRRV 63
> hsa:8683 SRSF9, SFRS9, SRp30c; serine/arginine-rich splicing
factor 9; K12890 splicing factor, arginine/serine-rich 1/9
Length=221
Score = 32.7 bits (73), Expect = 0.50, Method: Compositional matrix adjust.
Identities = 14/28 (50%), Positives = 20/28 (71%), Gaps = 0/28 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI 161
+YV NLP + ++L D+F KYG IR+I
Sbjct 16 IYVGNLPTDVREKDLEDLFYKYGRIREI 43
> cel:F26A3.2 ncbp-2; Nuclear Cap-Binding Protein family member
(ncbp-2); K12883 nuclear cap-binding protein subunit 2
Length=154
Score = 32.7 bits (73), Expect = 0.61, Method: Compositional matrix adjust.
Identities = 14/40 (35%), Positives = 24/40 (60%), Gaps = 0/40 (0%)
Query 122 RPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
R Q A S LYV NL + +++Y++FG+ G +R++
Sbjct 27 RDQETALRTSCTLYVGNLSYYTKEDQVYELFGRAGDVRRV 66
> mmu:108014 Srsf9, 25kDa, 2610029M16Rik, SRp30c, Sfrs9; serine/arginine-rich
splicing factor 9; K12890 splicing factor, arginine/serine-rich
1/9
Length=222
Score = 32.7 bits (73), Expect = 0.61, Method: Compositional matrix adjust.
Identities = 14/28 (50%), Positives = 20/28 (71%), Gaps = 0/28 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI 161
+YV NLP + ++L D+F KYG IR+I
Sbjct 17 IYVGNLPSDVREKDLEDLFYKYGRIREI 44
> ath:AT4G34110 PAB2; PAB2 (POLY(A) BINDING 2); RNA binding /
translation initiation factor; K13126 polyadenylate-binding
protein
Length=629
Score = 32.3 bits (72), Expect = 0.66, Method: Compositional matrix adjust.
Identities = 11/29 (37%), Positives = 19/29 (65%), Gaps = 0/29 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQIR 162
LYV +L F + +L+D FG+ G++ +R
Sbjct 38 LYVGDLDFNVTDSQLFDAFGQMGTVVTVR 66
> mmu:12387 Ctnnb1, Bfc, Catnb, Mesc; catenin (cadherin associated
protein), beta 1; K02105 catenin (cadherin-associated protein),
beta 1
Length=781
Score = 32.3 bits (72), Expect = 0.69, Method: Compositional matrix adjust.
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 2/58 (3%)
Query 107 KSPAMQPAPLLAGRGRPQRLAPEVSRILYVRNLPF--KINGEELYDVFGKYGSIRQIR 162
S A AP L+G+G P+ + S++LY F E++ D+ G+Y R R
Sbjct 36 HSGATTTAPSLSGKGNPEEEDVDTSQVLYEWEQGFSQSFTQEQVADIDGQYAMTRAQR 93
> hsa:1499 CTNNB1, CTNNB, DKFZp686D02253, FLJ25606, FLJ37923;
catenin (cadherin-associated protein), beta 1, 88kDa; K02105
catenin (cadherin-associated protein), beta 1
Length=781
Score = 32.3 bits (72), Expect = 0.70, Method: Compositional matrix adjust.
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 2/58 (3%)
Query 107 KSPAMQPAPLLAGRGRPQRLAPEVSRILYVRNLPF--KINGEELYDVFGKYGSIRQIR 162
S A AP L+G+G P+ + S++LY F E++ D+ G+Y R R
Sbjct 36 HSGATTTAPSLSGKGNPEEEDVDTSQVLYEWEQGFSQSFTQEQVADIDGQYAMTRAQR 93
> hsa:9904 RBM19, DKFZp586F1023, KIAA0682; RNA binding motif protein
19; K14787 multiple RNA-binding domain-containing protein
1
Length=960
Score = 32.3 bits (72), Expect = 0.72, Method: Composition-based stats.
Identities = 18/53 (33%), Positives = 32/53 (60%), Gaps = 2/53 (3%)
Query 110 AMQPAPLLAGRGRPQRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQIR 162
A +PA LA + + R S+IL VRN+PF+ + E+ ++F +G ++ +R
Sbjct 812 ATKPAVTLARKKQVPR-KQTTSKIL-VRNIPFQAHSREIRELFSTFGELKTVR 862
Score = 30.4 bits (67), Expect = 3.1, Method: Composition-based stats.
Identities = 12/28 (42%), Positives = 19/28 (67%), Gaps = 0/28 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQI 161
L+VRNLP+ E+L +F KYG + ++
Sbjct 404 LFVRNLPYTSTEEDLEKLFSKYGPLSEL 431
> ath:AT1G22760 PAB3; PAB3 (POLY(A) BINDING PROTEIN 3); RNA binding
/ translation initiation factor
Length=660
Score = 32.3 bits (72), Expect = 0.73, Method: Compositional matrix adjust.
Identities = 13/25 (52%), Positives = 18/25 (72%), Gaps = 0/25 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSI 158
+YV+NLP +I +EL FGK+G I
Sbjct 231 VYVKNLPKEIGEDELRKTFGKFGVI 255
> xla:494769 rbm19; RNA binding motif protein 19; K14787 multiple
RNA-binding domain-containing protein 1
Length=920
Score = 32.3 bits (72), Expect = 0.76, Method: Composition-based stats.
Identities = 23/64 (35%), Positives = 32/64 (50%), Gaps = 13/64 (20%)
Query 107 KSPAMQPAPLLAGRGRP--QRLAPEV-------SRILYVRNLPFKINGEELYDVFGKYGS 157
KSP++Q RP QR E+ S L+VRNLP+ ++L +F KYG
Sbjct 356 KSPSVQSK----AESRPWEQRDKQELQQEDLSESGRLFVRNLPYSCTEDDLDKLFSKYGP 411
Query 158 IRQI 161
I +I
Sbjct 412 ISEI 415
Score = 28.9 bits (63), Expect = 7.2, Method: Composition-based stats.
Identities = 11/38 (28%), Positives = 21/38 (55%), Gaps = 0/38 (0%)
Query 124 QRLAPEVSRILYVRNLPFKINGEELYDVFGKYGSIRQI 161
Q P ++ V+NLP EL ++FG++G + ++
Sbjct 562 QAAGPRSKTVILVKNLPAGTKPAELQELFGRFGDLGRV 599
> hsa:10146 G3BP1, G3BP, HDH-VIII, MGC111040; GTPase activating
protein (SH3 domain) binding protein 1 (EC:3.6.4.13 3.6.4.12)
Length=466
Score = 32.0 bits (71), Expect = 0.98, Method: Compositional matrix adjust.
Identities = 12/32 (37%), Positives = 21/32 (65%), Gaps = 0/32 (0%)
Query 131 SRILYVRNLPFKINGEELYDVFGKYGSIRQIR 162
S L++ NLP +++ EL D F YG++ ++R
Sbjct 339 SHQLFIGNLPHEVDKSELKDFFQSYGNVVELR 370
> dre:100330634 RNA binding motif protein 12-like
Length=675
Score = 32.0 bits (71), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 16/33 (48%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Query 124 QRLAPEVS-RILYVRNLPFKINGEELYDVFGKY 155
QR P+ R LYVRNLPF + E+ D F Y
Sbjct 382 QRNTPDSQLRCLYVRNLPFDVRKVEIMDFFHGY 414
> ath:AT3G52380 CP33; CP33; RNA binding
Length=329
Score = 32.0 bits (71), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 12/29 (41%), Positives = 19/29 (65%), Gaps = 0/29 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQIR 162
LYV NLP+ I EL +FG+ G++ ++
Sbjct 118 LYVGNLPYTITSSELSQIFGEAGTVVDVQ 146
> cel:B0035.12 hypothetical protein
Length=836
Score = 31.6 bits (70), Expect = 1.1, Method: Composition-based stats.
Identities = 12/32 (37%), Positives = 22/32 (68%), Gaps = 0/32 (0%)
Query 134 LYVRNLPFKINGEELYDVFGKYGSIRQIRRGT 165
++VRN+ F+ +EL +F K+G++ +RR T
Sbjct 685 VFVRNVHFQATDDELKALFSKFGTVTSVRRVT 716
Lambda K H
0.323 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3962792044
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40